[English] 日本語
 Yorodumi
Yorodumi- EMDB-30064: Clam-shaped joint of double-headed nucleocapsid of Sendai Virus -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30064 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Clam-shaped joint of double-headed nucleocapsid of Sendai Virus | |||||||||
 Map data Map data | 3D reconstruction of SeV NCWT clam-shaped structure | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | Paramyxovirus nucleocapsid protein / Paramyxovirus nucleocapsid protein / helical viral capsid / viral nucleocapsid / host cell cytoplasm / ribonucleoprotein complex / structural molecule activity / RNA binding / Nucleoprotein Function and homology information Function and homology information | |||||||||
| Biological species |  Murine respirovirus Murine respirovirus | |||||||||
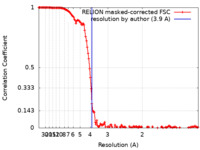
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Shen Q / Shan H / Zhang N | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2021 Journal: Commun Biol / Year: 2021Title: Structure and assembly of double-headed Sendai virus nucleocapsids. Authors: Na Zhang / Hong Shan / Mingdong Liu / Tianhao Li / Rui Luo / Liuyan Yang / Lei Qi / Xiaofeng Chu / Xin Su / Rui Wang / Yunhui Liu / Wenzhi Sun / Qing-Tao Shen /  Abstract: Paramyxoviruses, including the mumps virus, measles virus, Nipah virus and Sendai virus (SeV), have non-segmented single-stranded negative-sense RNA genomes which are encapsidated by nucleoproteins ...Paramyxoviruses, including the mumps virus, measles virus, Nipah virus and Sendai virus (SeV), have non-segmented single-stranded negative-sense RNA genomes which are encapsidated by nucleoproteins into helical nucleocapsids. Here, we reported a double-headed SeV nucleocapsid assembled in a tail-to-tail manner, and resolved its helical stems and clam-shaped joint at the respective resolutions of 2.9 and 3.9 Å, via cryo-electron microscopy. Our structures offer important insights into the mechanism of the helical polymerization, in particular via an unnoticed exchange of a N-terminal hole formed by three loops of nucleoproteins, and unveil the clam-shaped joint in a hyper-closed state for nucleocapsid dimerization. Direct visualization of the loop from the disordered C-terminal tail provides structural evidence that C-terminal tail is correlated to the curvature of nucleocapsid and links nucleocapsid condensation and genome replication and transcription with different assembly forms. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30064.map.gz emd_30064.map.gz | 152.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30064-v30.xml emd-30064-v30.xml emd-30064.xml emd-30064.xml | 8.8 KB 8.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30064_fsc.xml emd_30064_fsc.xml | 14.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_30064.png emd_30064.png | 240.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30064 http://ftp.pdbj.org/pub/emdb/structures/EMD-30064 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30064 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30064 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30064.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30064.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D reconstruction of SeV NCWT clam-shaped structure | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.65 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : clam-shaped joint of double-headed nucleocapsid of sendai virus
| Entire | Name: clam-shaped joint of double-headed nucleocapsid of sendai virus |
|---|---|
| Components |
|
-Supramolecule #1: clam-shaped joint of double-headed nucleocapsid of sendai virus
| Supramolecule | Name: clam-shaped joint of double-headed nucleocapsid of sendai virus type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Murine respirovirus Murine respirovirus |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: OTHER | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 289 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)