+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30035 | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | 3.6A Yeast Vo state3 prime | |||||||||||||||||||||||||||
 Map data Map data | sharpen | |||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||
 Keywords Keywords | V-ATPase / Vo sub-complex / rotary motor / transport protein | |||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcell wall mannoprotein biosynthetic process / ATPase-coupled ion transmembrane transporter activity / protein localization to vacuolar membrane / cellular response to alkaline pH / Insulin receptor recycling / Transferrin endocytosis and recycling / polyphosphate metabolic process / ROS and RNS production in phagocytes / Golgi lumen acidification / Amino acids regulate mTORC1 ...cell wall mannoprotein biosynthetic process / ATPase-coupled ion transmembrane transporter activity / protein localization to vacuolar membrane / cellular response to alkaline pH / Insulin receptor recycling / Transferrin endocytosis and recycling / polyphosphate metabolic process / ROS and RNS production in phagocytes / Golgi lumen acidification / Amino acids regulate mTORC1 / P-type proton-exporting transporter activity / vacuolar transport / vacuolar proton-transporting V-type ATPase, V0 domain / endosomal lumen acidification / vacuole organization / protein targeting to vacuole / proton-transporting V-type ATPase complex / fungal-type vacuole / vacuolar proton-transporting V-type ATPase complex / cellular hyperosmotic response / vacuolar acidification / fungal-type vacuole membrane / phosphatidylinositol-3,5-bisphosphate binding / proton transmembrane transporter activity / proton-transporting ATPase activity, rotational mechanism / intracellular copper ion homeostasis / RNA endonuclease activity / Neutrophil degranulation / proton transmembrane transport / cell periphery / transmembrane transport / endocytosis / ATPase binding / protein-containing complex assembly / intracellular iron ion homeostasis / membrane raft / Golgi membrane / endoplasmic reticulum membrane / membrane Similarity search - Function | |||||||||||||||||||||||||||
| Biological species |   | |||||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||||||||||||||||||||
 Authors Authors | Roh SH / Shekhar M | |||||||||||||||||||||||||||
| Funding support |  United States, United States,  Korea, Republic Of, Korea, Republic Of,  France, 8 items France, 8 items
| |||||||||||||||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2020 Journal: Sci Adv / Year: 2020Title: Cryo-EM and MD infer water-mediated proton transport and autoinhibition mechanisms of V complex. Authors: Soung-Hun Roh / Mrinal Shekhar / Grigore Pintilie / Christophe Chipot / Stephan Wilkens / Abhishek Singharoy / Wah Chiu /    Abstract: Rotary vacuolar adenosine triphosphatases (V-ATPases) drive transmembrane proton transport through a V proton channel subcomplex. Despite recent high-resolution structures of several rotary ATPases, ...Rotary vacuolar adenosine triphosphatases (V-ATPases) drive transmembrane proton transport through a V proton channel subcomplex. Despite recent high-resolution structures of several rotary ATPases, the dynamic mechanism of proton pumping remains elusive. Here, we determined a 2.7-Å cryo-electron microscopy (cryo-EM) structure of yeast V proton channel in nanodisc that reveals the location of ordered water molecules along the proton path, details of specific protein-lipid interactions, and the architecture of the membrane scaffold protein. Moreover, we uncover a state of V that shows the -ring rotated by ~14°. Molecular dynamics simulations demonstrate that the two rotary states are in thermal equilibrium and depict how the protonation state of essential glutamic acid residues couples water-mediated proton transfer with -ring rotation. Our cryo-EM models and simulations also rationalize a mechanism for inhibition of passive proton transport as observed for free V that is generated as a result of V-ATPase regulation by reversible disassembly in vivo. | |||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30035.map.gz emd_30035.map.gz | 3.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30035-v30.xml emd-30035-v30.xml emd-30035.xml emd-30035.xml | 25.2 KB 25.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30035_fsc.xml emd_30035_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_30035.png emd_30035.png | 109.7 KB | ||
| Filedesc metadata |  emd-30035.cif.gz emd-30035.cif.gz | 7.4 KB | ||
| Others |  emd_30035_additional_1.map.gz emd_30035_additional_1.map.gz | 23.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30035 http://ftp.pdbj.org/pub/emdb/structures/EMD-30035 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30035 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30035 | HTTPS FTP |
-Related structure data
| Related structure data |  6m0sMC  6m0rC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30035.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30035.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpen | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: unfil
| File | emd_30035_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unfil | ||||||||||||
| Projections & Slices |
| ||||||||||||
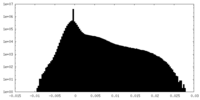
| Density Histograms |
- Sample components
Sample components
-Entire : V-type proton ATPase Vo sub-complex
| Entire | Name: V-type proton ATPase Vo sub-complex |
|---|---|
| Components |
|
-Supramolecule #1: V-type proton ATPase Vo sub-complex
| Supramolecule | Name: V-type proton ATPase Vo sub-complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 400 kDa/nm |
-Macromolecule #1: V-type proton ATPase subunit a, vacuolar isoform
| Macromolecule | Name: V-type proton ATPase subunit a, vacuolar isoform / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 94.289039 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EKEEAIFRSA EMALVQFYIP QEISRDSAYT LGQLGLVQFR DLNSKVRAFQ RTFVNEIRRL DNVERQYRYF YSLLKKHDIK LYEGDTDKY LDGSGELYVP PSGSVIDDYV RNASYLEERL IQMEDATDQI EVQKNDLEQY RFILQSGDEF FLKGDNTDST S YMDEDMID ...String: EKEEAIFRSA EMALVQFYIP QEISRDSAYT LGQLGLVQFR DLNSKVRAFQ RTFVNEIRRL DNVERQYRYF YSLLKKHDIK LYEGDTDKY LDGSGELYVP PSGSVIDDYV RNASYLEERL IQMEDATDQI EVQKNDLEQY RFILQSGDEF FLKGDNTDST S YMDEDMID ANGENIAAAI GASVNYVTGV IARDKVATLE QILWRVLRGN LFFKTVEIEQ PVYDVKTREY KHKNAFIVFS HG DLIIKRI RKIAESLDAN LYDVDSSNEG RSQQLAKVNK NLSDLYTVLK TTSTTLESEL YAIAKELDSW FQDVTREKAI FEI LNKSNY DTNRKILIAE GWIPRDELAT LQARLGEMIA RLGIDVPSII QVLDTNHTPP TFHRTNKFTA GFQSICDCYG IAQY REINA GLPTIVTFPF MFAIMFGDMG HGFLMTLAAL SLVLNEKKIN KMKRGEIFDM AFTGRYIILL MGVFSMYTGF LYNDI FSKT MTIFKSGWKW PDHWKKGESI TATSVGTYPI GLDWAWHGTE NALLFSNSYK MKLSILMGFI HMTYSYFFSL ANHLYF NSM IDIIGNFIPG LLFMQGIFGY LSVCIVYKWA VDWVKDGKPA PGLLNMLINM FLSPGTIDDE LYPHQAKVQV FLLLMAL VC IPWLLLVKPL HFKFTHKKKS HEPLPSTEAD ASSEDLEAQQ LISAMDADDA EEEEVGSGSH GEDFGDIMIH QVIHTIEF C LNCVSHTASY LRLWALSLAH AQLSSVLWTM TIQIAFGFRG FVGVFMTVAL FAMWFALTCA VLVLMEGTSA MLHSLRLHW VESMSKFFVG EGLPYEPFAF EYKDM UniProtKB: V-type proton ATPase subunit a, vacuolar isoform |
-Macromolecule #2: V-type proton ATPase subunit e
| Macromolecule | Name: V-type proton ATPase subunit e / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 8.186875 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSSFYTVVGV FIVVSAMSVL FWIMAPKNNQ AVWRSTVILT LAMMFLMWAI TFLCQLHPLV APRRSDLRPE F UniProtKB: V-type proton ATPase subunit e |
-Macromolecule #3: Uncharacterized protein YPR170W-B
| Macromolecule | Name: Uncharacterized protein YPR170W-B / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 7.487627 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: TGKAWCCTVL SAFGVVILSV IAHLFNTNHE SFVGSINDPE DGPAVAHTVY LAALVYLVFF VFCGFQVYL UniProtKB: V-type proton ATPase subunit f |
-Macromolecule #4: V-type proton ATPase subunit d
| Macromolecule | Name: V-type proton ATPase subunit d / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 39.822484 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEGVYFNIDN GFIEGVVRGY RNGLLSNNQY INLTQCDTLE DLKLQLSSTD YGNFLSSVSS ESLTTSLIQE YASSKLYHEF NYIRDQSSG STRKFMDYIT YGYMIDNVAL MITGTIHDRD KGEILQRCHP LGWFDTLPTL SVATDLESLY ETVLVDTPLA P YFKNCFDT ...String: MEGVYFNIDN GFIEGVVRGY RNGLLSNNQY INLTQCDTLE DLKLQLSSTD YGNFLSSVSS ESLTTSLIQE YASSKLYHEF NYIRDQSSG STRKFMDYIT YGYMIDNVAL MITGTIHDRD KGEILQRCHP LGWFDTLPTL SVATDLESLY ETVLVDTPLA P YFKNCFDT AEELDDMNIE IIRNKLYKAY LEDFYNFVTE EIPEPAKECM QTLLGFEADR RSINIALNSL QSSDIDPDLK SD LLPNIGK LYPLATFHLA QAQDFEGVRA ALANVYEYRG FLETGNLEDH FYQLEMELCR DAFTQQFAIS TVWAWMKSKE QEV RNITWI AECIAQNQRE RINNYISVY UniProtKB: V-type proton ATPase subunit d |
-Macromolecule #5: V-type proton ATPase subunit c''
| Macromolecule | Name: V-type proton ATPase subunit c'' / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 20.880686 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SFSHFLYYLV LIVVIVYGLY KLFTGHGSDI NFGKFLLRTS PYMWANLGIA LCVGLSVVGA AWGIFITGSS MIGAGVRAPR ITTKNLISI IFCEVVAIYG LIIAIVFSSK LTVATAENMY SKSNLYTGYS LFWAGITVGA SNLICGIAVG ITGATAAISD A ADSALFVK ...String: SFSHFLYYLV LIVVIVYGLY KLFTGHGSDI NFGKFLLRTS PYMWANLGIA LCVGLSVVGA AWGIFITGSS MIGAGVRAPR ITTKNLISI IFCEVVAIYG LIIAIVFSSK LTVATAENMY SKSNLYTGYS LFWAGITVGA SNLICGIAVG ITGATAAISD A ADSALFVK ILVIEIFGSI LGLLGLIVGL LMAGKASEFQ UniProtKB: V-type proton ATPase subunit c'' |
-Macromolecule #6: V-type proton ATPase subunit c'
| Macromolecule | Name: V-type proton ATPase subunit c' / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 16.414615 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SNIYAPLYAP FFGFAGCAAA MVLSCLGAAI GTAKSGIGIA GIGTFKPELI MKSLIPVVMS GILAIYGLVV AVLIAGNLSP TEDYTLFNG FMHLSCGLCV GFACLSSGYA IGMVGDVGVR KYMHQPRLFV GIVLILIFSE VLGLYGMIVA LILNTRGSE UniProtKB: V-type proton ATPase subunit c' |
-Macromolecule #7: V-type proton ATPase subunit c
| Macromolecule | Name: V-type proton ATPase subunit c / type: protein_or_peptide / ID: 7 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 16.254358 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTELCPVYAP FFGAIGCASA IIFTSLGAAY GTAKSGVGIC ATCVLRPDLL FKNIVPVIMA GIIAIYGLVV SVLVCYSLGQ KQALYTGFI QLGAGLSVGL SGLAAGFAIG IVGDAGVRGS SQQPRLFVGM ILILIFAEVL GLYGLIVALL LNSRATQDVV UniProtKB: V-type proton ATPase subunit c |
-Macromolecule #8: V0 assembly protein 1
| Macromolecule | Name: V0 assembly protein 1 / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 5.752846 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DDILSSIWTE GLLMCLIVSA LLLFILIVAL SWISNLDITY GALEKSTNPI KK UniProtKB: V0 assembly protein 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)