+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the Haloferax volcanii archaeal type IV pilus | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Archaea / type IV pili / STRUCTURAL PROTEIN | |||||||||
| Function / homology | Archaeal Type IV pilin, N-terminal / Archaeal Type IV pilin, N-terminal / Flagellin/pilin, N-terminal / membrane / Archaeal Type IV pilin N-terminal domain-containing protein Function and homology information Function and homology information | |||||||||
| Biological species |  Haloferax volcanii (archaea) / Haloferax volcanii (archaea) /  Haloferax volcanii (strain ATCC 29605 / DSM 3757 / JCM 8879 / NBRC 14742 / NCIMB 2012 / VKM B-1768 / DS2) (archaea) Haloferax volcanii (strain ATCC 29605 / DSM 3757 / JCM 8879 / NBRC 14742 / NCIMB 2012 / VKM B-1768 / DS2) (archaea) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Wang F / Kreutzberger MA / Baquero DP / Krupovic M / Egelman EH | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: The evolution of archaeal flagellar filaments. Authors: Mark A B Kreutzberger / Virginija Cvirkaite-Krupovic / Ying Liu / Diana P Baquero / Junfeng Liu / Ravi R Sonani / Chris R Calladine / Fengbin Wang / Mart Krupovic / Edward H Egelman /    Abstract: Flagellar motility has independently arisen three times during evolution: in bacteria, archaea, and eukaryotes. In prokaryotes, the supercoiled flagellar filaments are composed largely of a single ...Flagellar motility has independently arisen three times during evolution: in bacteria, archaea, and eukaryotes. In prokaryotes, the supercoiled flagellar filaments are composed largely of a single protein, bacterial or archaeal flagellin, although these two proteins are not homologous, while in eukaryotes, the flagellum contains hundreds of proteins. Archaeal flagellin and archaeal type IV pilin are homologous, but how archaeal flagellar filaments (AFFs) and archaeal type IV pili (AT4Ps) diverged is not understood, in part, due to the paucity of structures for AFFs and AT4Ps. Despite having similar structures, AFFs supercoil, while AT4Ps do not, and supercoiling is essential for the function of AFFs. We used cryo-electron microscopy to determine the atomic structure of two additional AT4Ps and reanalyzed previous structures. We find that all AFFs have a prominent 10-strand packing, while AT4Ps show a striking structural diversity in their subunit packing. A clear distinction between all AFF and all AT4P structures involves the extension of the N-terminal α-helix with polar residues in the AFFs. Additionally, we characterize a flagellar-like AT4P from with filament and subunit structure similar to that of AFFs which can be viewed as an evolutionary link, showing how the structural diversity of AT4Ps likely allowed for an AT4P to evolve into a supercoiling AFF. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29215.map.gz emd_29215.map.gz | 6.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29215-v30.xml emd-29215-v30.xml emd-29215.xml emd-29215.xml | 14 KB 14 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_29215.png emd_29215.png | 111.7 KB | ||
| Masks |  emd_29215_msk_1.map emd_29215_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-29215.cif.gz emd-29215.cif.gz | 5.2 KB | ||
| Others |  emd_29215_half_map_1.map.gz emd_29215_half_map_1.map.gz emd_29215_half_map_2.map.gz emd_29215_half_map_2.map.gz | 116 MB 116 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29215 http://ftp.pdbj.org/pub/emdb/structures/EMD-29215 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29215 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29215 | HTTPS FTP |
-Validation report
| Summary document |  emd_29215_validation.pdf.gz emd_29215_validation.pdf.gz | 830 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29215_full_validation.pdf.gz emd_29215_full_validation.pdf.gz | 829.6 KB | Display | |
| Data in XML |  emd_29215_validation.xml.gz emd_29215_validation.xml.gz | 14 KB | Display | |
| Data in CIF |  emd_29215_validation.cif.gz emd_29215_validation.cif.gz | 16.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29215 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29215 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29215 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29215 | HTTPS FTP |
-Related structure data
| Related structure data |  8fj5MC  7txiC  8fjsC  8fk0C  8fk7C  8gi2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_29215.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29215.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||||||||||||||||||
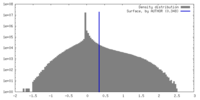
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29215_msk_1.map emd_29215_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_29215_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_29215_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Archaeal type IV pilus
| Entire | Name: Archaeal type IV pilus |
|---|---|
| Components |
|
-Supramolecule #1: Archaeal type IV pilus
| Supramolecule | Name: Archaeal type IV pilus / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Haloferax volcanii (archaea) Haloferax volcanii (archaea)Strain: ATCC 29605 / DSM 3757 / JCM 8879 / NBRC 14742 / NCIMB 2012 / VKM B-1768 / DS2 |
-Macromolecule #1: Pilin_N domain-containing protein
| Macromolecule | Name: Pilin_N domain-containing protein / type: protein_or_peptide / ID: 1 / Number of copies: 20 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Haloferax volcanii (strain ATCC 29605 / DSM 3757 / JCM 8879 / NBRC 14742 / NCIMB 2012 / VKM B-1768 / DS2) (archaea) Haloferax volcanii (strain ATCC 29605 / DSM 3757 / JCM 8879 / NBRC 14742 / NCIMB 2012 / VKM B-1768 / DS2) (archaea)Strain: ATCC 29605 / DSM 3757 / JCM 8879 / NBRC 14742 / NCIMB 2012 / VKM B-1768 / DS2 |
| Molecular weight | Theoretical: 14.841991 KDa |
| Recombinant expression | Organism:  Haloferax volcanii (archaea) Haloferax volcanii (archaea) |
| Sequence | String: MQLTTLFNDD SAVSPVIGVI LMVAITVILA AVIGTFVLGL GDQVSETSPQ ASFDFDYTNT SGNLTITHES GTSIDADSVS ISGPVGDDG KTWADIDGSA TEITAGSSIT VTANGSSFDS GETVRVIWTS DSGSSSSTLQ SWTYNG UniProtKB: Archaeal Type IV pilin N-terminal domain-containing protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 5.155 Å Applied symmetry - Helical parameters - Δ&Phi: 106.297 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 1660828 |
|---|---|
| Startup model | Type of model: OTHER |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)