[English] 日本語
 Yorodumi
Yorodumi- EMDB-28954: Asymmetric structure of cleaved HIV-1 AE2 envelope glycoprotein t... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Asymmetric structure of cleaved HIV-1 AE2 envelope glycoprotein trimer in styrene-maleic acid lipid nanoparticles (AE2.1) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HIV-1 / envelope glycoprotein / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane ...positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Wang K / Zhang S / Sodroski J / Mao Y | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2023 Journal: Commun Biol / Year: 2023Title: Asymmetric conformations of cleaved HIV-1 envelope glycoprotein trimers in styrene-maleic acid lipid nanoparticles. Authors: Kunyu Wang / Shijian Zhang / Eden P Go / Haitao Ding / Wei Li Wang / Hanh T Nguyen / John C Kappes / Heather Desaire / Joseph Sodroski / Youdong Mao /   Abstract: During virus entry, the pretriggered human immunodeficiency virus (HIV-1) envelope glycoprotein (Env) trimer initially transits into a default intermediate state (DIS) that remains structurally ...During virus entry, the pretriggered human immunodeficiency virus (HIV-1) envelope glycoprotein (Env) trimer initially transits into a default intermediate state (DIS) that remains structurally uncharacterized. Here, we present cryo-EM structures at near-atomic resolution of two cleaved full-length HIV-1 Env trimers purified from cell membranes in styrene-maleic acid lipid nanoparticles without antibodies or receptors. The cleaved Env trimers exhibited tighter subunit packing than uncleaved trimers. Cleaved and uncleaved Env trimers assumed remarkably consistent yet distinct asymmetric conformations, with one smaller and two larger opening angles. Breaking conformational symmetry is allosterically coupled with dynamic helical transformations of the gp41 N-terminal heptad repeat (HR1) regions in two protomers and with trimer tilting in the membrane. The broken symmetry of the DIS potentially assists Env binding to two CD4 receptors-while resisting antibody binding-and promotes extension of the gp41 HR1 helical coiled-coil, which relocates the fusion peptide closer to the target cell membrane. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28954.map.gz emd_28954.map.gz | 14.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28954-v30.xml emd-28954-v30.xml emd-28954.xml emd-28954.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28954.png emd_28954.png | 121 KB | ||
| Filedesc metadata |  emd-28954.cif.gz emd-28954.cif.gz | 6.1 KB | ||
| Others |  emd_28954_half_map_1.map.gz emd_28954_half_map_1.map.gz emd_28954_half_map_2.map.gz emd_28954_half_map_2.map.gz | 11.9 MB 11.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28954 http://ftp.pdbj.org/pub/emdb/structures/EMD-28954 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28954 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28954 | HTTPS FTP |
-Validation report
| Summary document |  emd_28954_validation.pdf.gz emd_28954_validation.pdf.gz | 801.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28954_full_validation.pdf.gz emd_28954_full_validation.pdf.gz | 801.4 KB | Display | |
| Data in XML |  emd_28954_validation.xml.gz emd_28954_validation.xml.gz | 9.5 KB | Display | |
| Data in CIF |  emd_28954_validation.cif.gz emd_28954_validation.cif.gz | 11.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28954 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28954 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28954 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28954 | HTTPS FTP |
-Related structure data
| Related structure data |  8faeMC  8fadC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28954.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28954.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.65 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
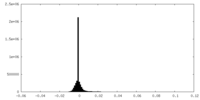
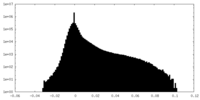
-Half map: #1
| File | emd_28954_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
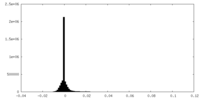
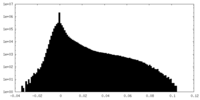
-Half map: #2
| File | emd_28954_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : HIV-1 Env AE2.1
| Entire | Name: HIV-1 Env AE2.1 |
|---|---|
| Components |
|
-Supramolecule #1: HIV-1 Env AE2.1
| Supramolecule | Name: HIV-1 Env AE2.1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Macromolecule #1: Envelope glycoprotein gp41
| Macromolecule | Name: Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 16.639076 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: FLGFLGAAGS TMGAASITLT VQARLLLSGI VQQQNNLLKA IEAQQHLLKL TVWGIKQLQA RVLTVERYLR DQQLLGIWGC SGKLICTTA VPWNASWSNK TLDMIWNNMT WMEWEKEIDN YTGLIYTLIE ESQNQQEKNE KELLELD UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #2: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 53.025961 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EKLWVTVYYG VPVWKEATTT LFCASDAKAY DTEVHNVWAT HACVPTDPNP QEVVLENVTE NFNMWKNNMV EQMHEDIISL WDESLKPCV KLTPLCVTLN CTDLRNVTNI NNSSEGMRGE IKNCSFNITT SIKDKVKKDY ALFYKLDVVP IDNDNTSYRL I NCNTSTIT ...String: EKLWVTVYYG VPVWKEATTT LFCASDAKAY DTEVHNVWAT HACVPTDPNP QEVVLENVTE NFNMWKNNMV EQMHEDIISL WDESLKPCV KLTPLCVTLN CTDLRNVTNI NNSSEGMRGE IKNCSFNITT SIKDKVKKDY ALFYKLDVVP IDNDNTSYRL I NCNTSTIT QACPKVSFEP IPIHYCTPAG FAILKCKDKK FNGTGPCKNV STVQCTHGIK PVVSTQLLLN GSLAEEEVVI RS SNFTDNA KNIIVQLKES VEINCTRPNN NTRKSIHIGP GKAFYTTGDI IGDIRQAHCN ISRTKWNNTL NQIATKLKEQ FGN NKTIVF NQSSGGDPEI VMHSFNCGGE FFYCNSTQLF NSTWNFNGTW NLTQSNGTEG NDTITLPCKI KQIINMWQEV GKAM YAPPI RGQIRCSSNI TGLILTRDGG NNHNNDTETF RPGGGDMRDN WRSELYKYKV VKIEPLGVAP TKAKRRV UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #18: 1-[(2R)-4-(benzenecarbonyl)-2-methylpiperazin-1-yl]-2-(4-methoxy-...
| Macromolecule | Name: 1-[(2R)-4-(benzenecarbonyl)-2-methylpiperazin-1-yl]-2-(4-methoxy-1H-pyrrolo[2,3-b]pyridin-3-yl)ethane-1,2-dione type: ligand / ID: 18 / Number of copies: 3 / Formula: 83G |
|---|---|
| Molecular weight | Theoretical: 406.434 Da |
| Chemical component information | 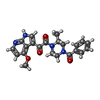 ChemComp-83G: |
-Macromolecule #19: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 19 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 57.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 140035 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)