登録情報 データベース : EMDB / ID : EMD-28629タイトル CX3CR1 nucleosome and wild type PU.1 complex 複合体 : nucleosome PU.1 complexタンパク質・ペプチド : Histone H3.1タンパク質・ペプチド : Histone H4タンパク質・ペプチド : Histone H2A type 2-Cタンパク質・ペプチド : Histone H2B type 2-EDNA : DNA (162-MER)DNA : DNA (162-MER)タンパク質・ペプチド : Single-chain variable fragmentタンパク質・ペプチド : Transcription factor PU.1 / / / / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト) / Escherichia coli (大腸菌) / Mus musculus (ハツカネズミ)手法 / / 解像度 : 2.85 Å Lian T / Guan R / Bai Y 資金援助 Organization Grant number 国 National Institutes of Health/National Cancer Institute (NIH/NCI)
ジャーナル : Nat Struct Mol Biol / 年 : 2024タイトル : Structural mechanism of synergistic targeting of the CX3CR1 nucleosome by PU.1 and C/EBPα.著者 : Tengfei Lian / Ruifang Guan / Bing-Rui Zhou / Yawen Bai / 要旨 : Pioneer transcription factors are vital for cell fate changes. PU.1 and C/EBPα work together to regulate hematopoietic stem cell differentiation. However, how they recognize in vivo nucleosomal DNA ... Pioneer transcription factors are vital for cell fate changes. PU.1 and C/EBPα work together to regulate hematopoietic stem cell differentiation. However, how they recognize in vivo nucleosomal DNA targets remains elusive. Here we report the structures of the nucleosome containing the mouse genomic CX3CR1 enhancer DNA and its complexes with PU.1 alone and with both PU.1 and the C/EBPα DNA binding domain. Our structures reveal that PU.1 binds the DNA motif at the exit linker, shifting 17 bp of DNA into the core region through interactions with H2A, unwrapping ~20 bp of nucleosomal DNA. C/EBPα binding, aided by PU.1's repositioning, unwraps ~25 bp of entry DNA. The PU.1 Q218H mutation, linked to acute myeloid leukemia, disrupts PU.1-H2A interactions. PU.1 and C/EBPα jointly displace linker histone H1 and open the H1-condensed nucleosome array. Our study unveils how two pioneer factors can work cooperatively to open closed chromatin by altering DNA positioning in the nucleosome. 履歴 登録 2022年10月20日 - ヘッダ(付随情報) 公開 2023年11月1日 - マップ公開 2023年11月1日 - 更新 2024年10月16日 - 現状 2024年10月16日 処理サイト : RCSB / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報
 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト) /
Homo sapiens (ヒト) / 

 データ登録者
データ登録者 米国, 1件
米国, 1件  引用
引用 ジャーナル: Nat Struct Mol Biol / 年: 2024
ジャーナル: Nat Struct Mol Biol / 年: 2024
 構造の表示
構造の表示 ダウンロードとリンク
ダウンロードとリンク emd_28629.map.gz
emd_28629.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-28629-v30.xml
emd-28629-v30.xml emd-28629.xml
emd-28629.xml EMDBヘッダ
EMDBヘッダ emd_28629.png
emd_28629.png emd-28629.cif.gz
emd-28629.cif.gz emd_28629_half_map_1.map.gz
emd_28629_half_map_1.map.gz emd_28629_half_map_2.map.gz
emd_28629_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-28629
http://ftp.pdbj.org/pub/emdb/structures/EMD-28629 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28629
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28629 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_28629.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_28629.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)






 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 画像解析
画像解析 ムービー
ムービー コントローラー
コントローラー



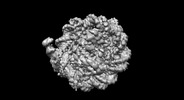
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)