[English] 日本語
 Yorodumi
Yorodumi- EMDB-28589: Cryo-EM structure of the organic cation transporter 2 in complex ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the organic cation transporter 2 in complex with 1-methyl-4-phenylpyridinium | |||||||||
 Map data Map data | main map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cation / transport / membrane protein / TRANSPORT PROTEIN | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.61 Å | |||||||||
 Authors Authors | Suo Y / Wright NJ / Lee S-Y | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation | Journal: bioRxiv / Year: 2023 Title: Molecular basis of polyspecific drug binding and transport by OCT1 and OCT2. Authors: Yang Suo / Nicholas J Wright / Hugo Guterres / Justin G Fedor / Kevin John Butay / Mario J Borgnia / Wonpil Im / Seok-Yong Lee /  Abstract: A wide range of endogenous and xenobiotic organic ions require facilitated transport systems to cross the plasma membrane for their disposition . In mammals, organic cation transporter subtypes 1 ...A wide range of endogenous and xenobiotic organic ions require facilitated transport systems to cross the plasma membrane for their disposition . In mammals, organic cation transporter subtypes 1 and 2 (OCT1 and OCT2, also known as SLC22A1 and SLC22A2, respectively) are polyspecific transporters responsible for the uptake and clearance of structurally diverse cationic compounds in the liver and kidneys, respectively . Notably, it is well established that human OCT1 and OCT2 play central roles in the pharmacokinetics, pharmacodynamics, and drug-drug interactions (DDI) of many prescription medications, including metformin . Despite their importance, the basis of polyspecific cationic drug recognition and the alternating access mechanism for OCTs have remained a mystery. Here, we present four cryo-EM structures of apo, substrate-bound, and drug-bound OCT1 and OCT2 in outward-facing and outward-occluded states. Together with functional experiments, docking, and molecular dynamics simulations, these structures uncover general principles of organic cation recognition by OCTs and illuminate unexpected features of the OCT alternating access mechanism. Our findings set the stage for a comprehensive structure-based understanding of OCT-mediated DDI, which will prove critical in the preclinical evaluation of emerging therapeutics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28589.map.gz emd_28589.map.gz | 20.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28589-v30.xml emd-28589-v30.xml emd-28589.xml emd-28589.xml | 19.8 KB 19.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28589.png emd_28589.png | 18.5 KB | ||
| Filedesc metadata |  emd-28589.cif.gz emd-28589.cif.gz | 6.4 KB | ||
| Others |  emd_28589_half_map_1.map.gz emd_28589_half_map_1.map.gz emd_28589_half_map_2.map.gz emd_28589_half_map_2.map.gz | 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28589 http://ftp.pdbj.org/pub/emdb/structures/EMD-28589 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28589 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28589 | HTTPS FTP |
-Related structure data
| Related structure data |  8et9MC  8et6C  8et7C  8et8C M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28589.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28589.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half 1
| File | emd_28589_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half 2
| File | emd_28589_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : OCT2
| Entire | Name: OCT2 |
|---|---|
| Components |
|
-Supramolecule #1: OCT2
| Supramolecule | Name: OCT2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 60 kDa/nm |
-Macromolecule #1: OCT2
| Macromolecule | Name: OCT2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 62.030547 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPTTFDDILE HIGEFGRFQK QTFFLLCLLS AAFAPIYVGI VFLGFTPDHR CRSPGVAELS QRCGWSLEEE LNYTVPGVGS SGEASPSQC RRYEVDWNQT GLSCTDPLAS LAANRSHLPL GPCQDGWVYD TPGSSIVTEF NLVCADSWML DLVQAAVNVG F FVGSMSIG ...String: MPTTFDDILE HIGEFGRFQK QTFFLLCLLS AAFAPIYVGI VFLGFTPDHR CRSPGVAELS QRCGWSLEEE LNYTVPGVGS SGEASPSQC RRYEVDWNQT GLSCTDPLAS LAANRSHLPL GPCQDGWVYD TPGSSIVTEF NLVCADSWML DLVQAAVNVG F FVGSMSIG YIADRFGRKL CLLVTILINA ISGVLMAVAP NYTWMLIFRL IQGLVSKGGW LIGYTLPTEF VGLEYRRTVG IL YQMAFSV GLLVLAGVAY AIPHWRWLQL AVTLPNFFFL LYYWCIPESP RWLISQNKNA KAMKIIKHIA KKNGKKLPKS LQE ERKETE VGEKLNPSFL DLVRTPQIRK HTLILMYNWF TSSVLYQGLI MHMGIAGGNI YLDFFYSALV EFPAAFIIIL TIDR IGRRY PWAAANMVAG AACLITAFIP DGLFWLKTTV ACLGRMGITM AFEMVCFVNT ELYPTFIRNL GVLVCSSLCD IGGII TPFL VYRLASIWLE LPLVVFAVLG LIAGGLVLLL PETKGKPLPE TIEDAENPHR PRKNKEKMIY LQVLLSDIPD N |
-Macromolecule #2: 1-methyl-4-phenylpyridin-1-ium
| Macromolecule | Name: 1-methyl-4-phenylpyridin-1-ium / type: ligand / ID: 2 / Number of copies: 1 / Formula: WRF |
|---|---|
| Molecular weight | Theoretical: 170.23 Da |
| Chemical component information | 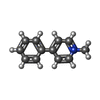 ChemComp-WRF: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 240 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 8029 / Average exposure time: 3.7 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 141.9 |
|---|---|
| Output model |  PDB-8et9: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)