[English] 日本語
 Yorodumi
Yorodumi- EMDB-28546: Structure of SARS-CoV-2 Orf3a in late endosome/lysosome-like envi... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of SARS-CoV-2 Orf3a in late endosome/lysosome-like environment, Saposin A nanodisc | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationhost cell lysosome / symbiont-mediated activation of host reticulophagy / Maturation of protein 3a / ganglioside GM1 transport to membrane / ganglioside GM2 binding / ganglioside GM3 binding / ganglioside GP1c binding / ganglioside GM1 binding / ganglioside GT1b binding / SARS-CoV-2 modulates autophagy ...host cell lysosome / symbiont-mediated activation of host reticulophagy / Maturation of protein 3a / ganglioside GM1 transport to membrane / ganglioside GM2 binding / ganglioside GM3 binding / ganglioside GP1c binding / ganglioside GM1 binding / ganglioside GT1b binding / SARS-CoV-2 modulates autophagy / sphingolipid metabolic process / epithelial cell differentiation involved in prostate gland development / Glycosphingolipid catabolism / prostate gland growth / : / lysosomal transport / voltage-gated calcium channel complex / azurophil granule membrane / host cell endoplasmic reticulum / regulation of lipid metabolic process / monoatomic ion channel activity / SARS-CoV-2 targets host intracellular signalling and regulatory pathways / voltage-gated potassium channel complex / lysosomal lumen / molecular function activator activity / Peptide ligand-binding receptors / regulation of autophagy / phospholipid binding / enzyme activator activity / cytoplasmic side of plasma membrane / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / late endosome / Platelet degranulation / protease binding / scaffold protein binding / Translation of Structural Proteins / Virion Assembly and Release / host cell endosome / Induction of Cell-Cell Fusion / G alpha (i) signalling events / gene expression / Attachment and Entry / lysosome / host cell endoplasmic reticulum membrane / lysosomal membrane / Neutrophil degranulation / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / protein homodimerization activity / : / extracellular exosome / extracellular region / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Miller AN / Houlihan PR / Matamala E / Cabezas-Bratesco D / Lee GY / Cristofori-Armstrong B / Dilan TL / Sanchez-Martinez S / Matthies D / Yan R ...Miller AN / Houlihan PR / Matamala E / Cabezas-Bratesco D / Lee GY / Cristofori-Armstrong B / Dilan TL / Sanchez-Martinez S / Matthies D / Yan R / Yu Z / Ren D / Brauchi SE / Clapham DE | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation | Journal: bioRxiv / Year: 2022 Title: The SARS-CoV-2 accessory protein Orf3a is not an ion channel, but does interact with trafficking proteins. Authors: Alexandria N Miller / Patrick R Houlihan / Ella Matamala / Deny Cabezas-Bratesco / Gi Young Lee / Ben Cristofori-Armstrong / Tanya L Dilan / Silvia Sanchez-Martinez / Doreen Matthies / Rui ...Authors: Alexandria N Miller / Patrick R Houlihan / Ella Matamala / Deny Cabezas-Bratesco / Gi Young Lee / Ben Cristofori-Armstrong / Tanya L Dilan / Silvia Sanchez-Martinez / Doreen Matthies / Rui Yan / Zhiheng Yu / Dejian Ren / Sebastian E Brauchi / David E Clapham Abstract: The severe acute respiratory syndrome associated coronavirus 2 (SARS-CoV-2) and SARS-CoV-1 accessory protein Orf3a colocalizes with markers of the plasma membrane, endocytic pathway, and Golgi ...The severe acute respiratory syndrome associated coronavirus 2 (SARS-CoV-2) and SARS-CoV-1 accessory protein Orf3a colocalizes with markers of the plasma membrane, endocytic pathway, and Golgi apparatus. Some reports have led to annotation of both Orf3a proteins as a viroporin. Here we show that neither SARS-CoV-2 nor SARS-CoV-1 form functional ion conducting pores and that the conductances measured are common contaminants in overexpression and with high levels of protein in reconstitution studies. Cryo-EM structures of both SARS-CoV-2 and SARS-CoV-1 Orf3a display a narrow constriction and the presence of a basic aqueous vestibule, which would not favor cation permeation. We observe enrichment of the late endosomal marker Rab7 upon SARS-CoV-2 Orf3a overexpression, and co-immunoprecipitation with VPS39. Interestingly, SARS-CoV-1 Orf3a does not cause the same cellular phenotype as SARS-CoV-2 Orf3a and does not interact with VPS39. To explain this difference, we find that a divergent, unstructured loop of SARS-CoV-2 Orf3a facilitates its binding with VPS39, a HOPS complex tethering protein involved in late endosome and autophagosome fusion with lysosomes. We suggest that the added loop enhances SARS-CoV-2 Orf3a ability to co-opt host cellular trafficking mechanisms for viral exit or host immune evasion. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28546.map.gz emd_28546.map.gz | 5.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28546-v30.xml emd-28546-v30.xml emd-28546.xml emd-28546.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28546.png emd_28546.png | 91.9 KB | ||
| Others |  emd_28546_half_map_1.map.gz emd_28546_half_map_1.map.gz emd_28546_half_map_2.map.gz emd_28546_half_map_2.map.gz | 48.6 MB 48.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28546 http://ftp.pdbj.org/pub/emdb/structures/EMD-28546 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28546 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28546 | HTTPS FTP |
-Related structure data
| Related structure data |  8equMC  8eqjC  8eqsC  8eqtC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28546.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28546.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.844 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_28546_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
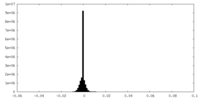
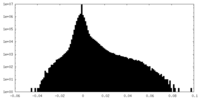
| Density Histograms |
-Half map: #2
| File | emd_28546_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
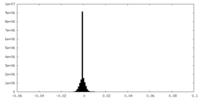
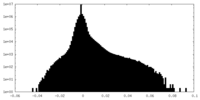
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of SARS-CoV-2 Orf3a in late endosome/lysosome-like envi...
| Entire | Name: Structure of SARS-CoV-2 Orf3a in late endosome/lysosome-like environment, Saposin A nanodisc |
|---|---|
| Components |
|
-Supramolecule #1: Structure of SARS-CoV-2 Orf3a in late endosome/lysosome-like envi...
| Supramolecule | Name: Structure of SARS-CoV-2 Orf3a in late endosome/lysosome-like environment, Saposin A nanodisc type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: ORF3a protein
| Macromolecule | Name: ORF3a protein / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.489445 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDLFMRIFTI GTVTLKQGEI KDATPSDFVR ATATIPIQAS LPFGWLIVGV ALLAVFQSAS KIITLKKRWQ LALSKGVHFV CNLLLLFVT VYSHLLLVAA GLEAPFLYLY ALVYFLQSIN FVRIIMRLWL CWKCRSKNPL LYDANYFLCW HTNCYDYCIP Y NSVTSSIV ...String: MDLFMRIFTI GTVTLKQGEI KDATPSDFVR ATATIPIQAS LPFGWLIVGV ALLAVFQSAS KIITLKKRWQ LALSKGVHFV CNLLLLFVT VYSHLLLVAA GLEAPFLYLY ALVYFLQSIN FVRIIMRLWL CWKCRSKNPL LYDANYFLCW HTNCYDYCIP Y NSVTSSIV ITSGDGTTSP ISEHDYQIGG YTEKWESGVK DCVVLHSYFT SDYYQLYSTQ LSTDTGVEHV TFFIYNKIVD EP EEHVQIH TIDGSSGVVN PVMEPIYDEP TTTTSVPLGG RGLEVLFQGP GSGQLVGSGG LEGGGGWSHP QFEKGGGSGG GSG GGSWSH PQFEK |
-Macromolecule #2: Saposin-A
| Macromolecule | Name: Saposin-A / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.677292 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGGHHHHHHS SGVDLGTENL YFQSMSLPCD ICKDVVTAAG DMLKDNATEE EILVYLEKTC DWLPKPNMSA SCKEIVDSYL PVILDIIKG EMSRPGEVCS ALNLCES |
-Macromolecule #3: Saposin A, polyalanine model
| Macromolecule | Name: Saposin A, polyalanine model / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 6.741301 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) |
-Macromolecule #4: 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine
| Macromolecule | Name: 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine / type: ligand / ID: 4 / Number of copies: 2 / Formula: PEE |
|---|---|
| Molecular weight | Theoretical: 744.034 Da |
| Chemical component information |  ChemComp-PEE: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 2 / Number real images: 15946 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 2.8 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 135280 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 3.0) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8equ: |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)