[English] 日本語
 Yorodumi
Yorodumi- EMDB-2822: Negative stain electron microscopy reconstruction of an intact Ph... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2822 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Negative stain electron microscopy reconstruction of an intact Phycobilisome in complex with photosystem II. | |||||||||
 Map data Map data | Negative stain electron microscopy reconstruction of the intact Phycobilisome from Anabaena sp. strain PCC 7120 in complex with photosystem II. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | light-harvesting antennae / photosynthesis | |||||||||
| Biological species |  Anabaena (bacteria) Anabaena (bacteria) | |||||||||
| Method | single particle reconstruction / Resolution: 34.0 Å | |||||||||
 Authors Authors | Chang L / Liu X / Li Y / Liu CC / Yang F / Zhao J / Sui SF | |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2015 Journal: Cell Res / Year: 2015Title: Structural organization of an intact phycobilisome and its association with photosystem II. Authors: Leifu Chang / Xianwei Liu / Yanbing Li / Cui-Cui Liu / Fan Yang / Jindong Zhao / Sen-Fang Sui /   Abstract: Phycobilisomes (PBSs) are light-harvesting antennae that transfer energy to photosynthetic reaction centers in cyanobacteria and red algae. PBSs are supermolecular complexes composed of ...Phycobilisomes (PBSs) are light-harvesting antennae that transfer energy to photosynthetic reaction centers in cyanobacteria and red algae. PBSs are supermolecular complexes composed of phycobiliproteins (PBPs) that bear chromophores for energy absorption and linker proteins. Although the structures of some individual components have been determined using crystallography, the three-dimensional structure of an entire PBS complex, which is critical for understanding the energy transfer mechanism, remains unknown. Here, we report the structures of an intact PBS and a PBS in complex with photosystem II (PSII) from Anabaena sp. strain PCC 7120 using single-particle electron microscopy in combination with biochemical and molecular analyses. In the PBS structure, all PBP trimers and the conserved linker protein domains were unambiguously located, and the global distribution of all chromophores was determined. We provide evidence that ApcE and ApcF are critical for the formation of a protrusion at the bottom of PBS, which plays an important role in mediating PBS interaction with PSII. Our results provide insights into the molecular architecture of an intact PBS at different assembly levels and provide the basis for understanding how the light energy absorbed by PBS is transferred to PSII. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2822.map.gz emd_2822.map.gz | 16.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2822-v30.xml emd-2822-v30.xml emd-2822.xml emd-2822.xml | 8.2 KB 8.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2822.jpg emd_2822.jpg | 42.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2822 http://ftp.pdbj.org/pub/emdb/structures/EMD-2822 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2822 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2822 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2822.map.gz / Format: CCP4 / Size: 21.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2822.map.gz / Format: CCP4 / Size: 21.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Negative stain electron microscopy reconstruction of the intact Phycobilisome from Anabaena sp. strain PCC 7120 in complex with photosystem II. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.46 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
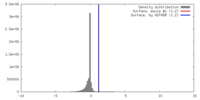
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Phycobilisome from Anabaena sp. strain PCC 7120 in complex with p...
| Entire | Name: Phycobilisome from Anabaena sp. strain PCC 7120 in complex with photosystem II. |
|---|---|
| Components |
|
-Supramolecule #1000: Phycobilisome from Anabaena sp. strain PCC 7120 in complex with p...
| Supramolecule | Name: Phycobilisome from Anabaena sp. strain PCC 7120 in complex with photosystem II. type: sample / ID: 1000 / Number unique components: 2 |
|---|---|
| Molecular weight | Theoretical: 6 MDa |
-Macromolecule #1: Phycobilisome
| Macromolecule | Name: Phycobilisome / type: protein_or_peptide / ID: 1 / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  Anabaena (bacteria) Anabaena (bacteria) |
| Molecular weight | Theoretical: 6 MDa |
-Macromolecule #2: photosystem II
| Macromolecule | Name: photosystem II / type: protein_or_peptide / ID: 2 / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  Anabaena (bacteria) Anabaena (bacteria) |
-Experimental details
-Structure determination
 Processing Processing | single particle reconstruction |
|---|---|
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 / Details: 0.8 M K/Na-PO4 |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Jun 6, 2011 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Average electron dose: 30 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 34.0 Å / Resolution method: OTHER / Software - Name: EMAN / Number images used: 2363 |
|---|
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)