[English] 日本語
 Yorodumi
Yorodumi- EMDB-27914: Escherichia coli Rho-dependent transcription pre-termination comp... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part | |||||||||
 Map data Map data | Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | factor-dependent termination / Rho / transcription termination / transcription elongation complex / helicase / ATPase / TRANSCRIPTION-RNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationATP-dependent activity, acting on RNA / helicase activity / DNA-templated transcription termination / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / ATP hydrolysis activity / RNA binding / ATP binding / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Molodtsov V / Wang C | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Structural basis of Rho-dependent transcription termination. Authors: Vadim Molodtsov / Chengyuan Wang / Emre Firlar / Jason T Kaelber / Richard H Ebright /  Abstract: Rho is a ring-shaped hexameric ATP-dependent molecular motor. Together with the transcription elongation factor NusG, Rho mediates factor-dependent transcription termination and transcription- ...Rho is a ring-shaped hexameric ATP-dependent molecular motor. Together with the transcription elongation factor NusG, Rho mediates factor-dependent transcription termination and transcription-translation-coupling quality control in Escherichia coli. Here we report the preparation of complexes that are functional in factor-dependent transcription termination from Rho, NusG, RNA polymerase (RNAP), and synthetic nucleic acid scaffolds, and we report cryogenic electron microscopy structures of the complexes. The structures show that functional factor-dependent pre-termination complexes contain a closed-ring Rho hexamer; have RNA threaded through the central channel of Rho; have 60 nucleotides of RNA interacting sequence-specifically with the exterior of Rho and 6 nucleotides of RNA interacting sequence-specifically with the central channel of Rho; have Rho oriented relative to RNAP such that ATP-dependent translocation by Rho exerts mechanical force on RNAP; and have NusG bridging Rho and RNAP. The results explain five decades of research on Rho and provide a foundation for understanding Rho's function. #1:  Journal: Biorxiv / Year: 2022 Journal: Biorxiv / Year: 2022Title: Structural basis of Rho-dependent transcription termination Authors: Molodtsov V / Wang C / Firlar E / Kaelber JT / Ebright RH | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27914.map.gz emd_27914.map.gz | 135.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27914-v30.xml emd-27914-v30.xml emd-27914.xml emd-27914.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27914.png emd_27914.png | 20.7 KB | ||
| Filedesc metadata |  emd-27914.cif.gz emd-27914.cif.gz | 5.7 KB | ||
| Others |  emd_27914_half_map_1.map.gz emd_27914_half_map_1.map.gz emd_27914_half_map_2.map.gz emd_27914_half_map_2.map.gz | 146.1 MB 146.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27914 http://ftp.pdbj.org/pub/emdb/structures/EMD-27914 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27914 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27914 | HTTPS FTP |
-Related structure data
| Related structure data |  8e5lMC  8e3fC  8e3hC  8e5kC  8e5oC  8e5pC  8e6wC  8e6xC  8e6zC  8e70C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27914.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27914.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.038 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half-map 1 Escherichia coli Rho-dependent transcription pre-termination complex...
| File | emd_27914_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1 Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part | ||||||||||||
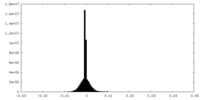
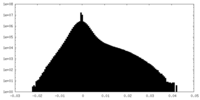
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map 2 Escherichia coli Rho-dependent transcription pre-termination complex...
| File | emd_27914_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2 Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part | ||||||||||||
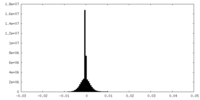
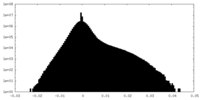
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Escherichia coli Rho-dependent transcription pre-termination comp...
| Entire | Name: Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part |
|---|---|
| Components |
|
-Supramolecule #1: Escherichia coli Rho-dependent transcription pre-termination comp...
| Supramolecule | Name: Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: RNA with 21 nt long spacer
| Macromolecule | Name: RNA with 21 nt long spacer / type: rna / ID: 1 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 11.99382 KDa |
| Sequence | String: AUGUUUUUUU UUUUUUUUUU UUUUGAUUUG GUGAGAGG |
-Macromolecule #2: Transcription termination factor Rho
| Macromolecule | Name: Transcription termination factor Rho / type: protein_or_peptide / ID: 2 / Number of copies: 6 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.070168 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNLTELKNTP VSELITLGEN MGLENLARMR KQDIIFAILK QHAKSGEDIF GDGVLEILQD GFGFLRSADS SYLAGPDDIY VSPSQIRRF NLRTGDTISG KIRPPKEGER YFALLKVNEV NFDKPENARN KILFENLTPL HANSRLRMER GNGSTEDLTA R VLDLASPI ...String: MNLTELKNTP VSELITLGEN MGLENLARMR KQDIIFAILK QHAKSGEDIF GDGVLEILQD GFGFLRSADS SYLAGPDDIY VSPSQIRRF NLRTGDTISG KIRPPKEGER YFALLKVNEV NFDKPENARN KILFENLTPL HANSRLRMER GNGSTEDLTA R VLDLASPI GRGQRGLIVA PPKAGKTMLL QNIAQSIAYN HPDCVLMVLL IDERPEEVTE MQRLVKGEVV ASTFDEPASR HV QVAEMVI EKAKRLVEHK KDVIILLDSI TRLARAYNTV VPASGKVLTG GVDANALHRP KRFFGAARNV EEGGSLTIIA TAL IDTGSK MDEVIYEEFK GTGNMELHLS RKIAEKRVFP AIDYNRSGTR KEELLTTQEE LQKMWILRKI IHPMGEIDAM EFLI NKLAM TKTNDDFFEM MKRS UniProtKB: Transcription termination factor Rho |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 6 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 6 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: BERYLLIUM TRIFLUORIDE ION
| Macromolecule | Name: BERYLLIUM TRIFLUORIDE ION / type: ligand / ID: 5 / Number of copies: 6 / Formula: BEF |
|---|---|
| Molecular weight | Theoretical: 66.007 Da |
| Chemical component information |  ChemComp-BEF: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 28.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.25 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.2 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 31098 |
| Initial angle assignment | Type: COMMON LINE |
| Final angle assignment | Type: COMMON LINE |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)