+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | MicroED structure of proteinase K recorded on K2 | |||||||||
 Map data Map data | 2mFo-DFc | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | serine protease / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationphosphorelay signal transduction system / regulation of DNA-templated transcription / DNA binding / membrane Similarity search - Function | |||||||||
| Biological species |  Parengyodontium album (fungus) Parengyodontium album (fungus) | |||||||||
| Method | electron crystallography / cryo EM | |||||||||
 Authors Authors | Clabbers MTB / Martynowycz MW / Hattne J / Nannenga BL / Gonen T | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2022 Journal: J Struct Biol / Year: 2022Title: Electron-counting MicroED data with the K2 and K3 direct electron detectors. Authors: Max T B Clabbers / Michael W Martynowycz / Johan Hattne / Brent L Nannenga / Tamir Gonen /  Abstract: Microcrystal electron diffraction (MicroED) uses electron cryo-microscopy (cryo-EM) to collect diffraction data from small crystals during continuous rotation of the sample. As a result of advances ...Microcrystal electron diffraction (MicroED) uses electron cryo-microscopy (cryo-EM) to collect diffraction data from small crystals during continuous rotation of the sample. As a result of advances in hardware as well as methods development, the data quality has continuously improved over the past decade, to the point where even macromolecular structures can be determined ab initio. Detectors suitable for electron diffraction should ideally have fast readout to record data in movie mode, and high sensitivity at low exposure rates to accurately report the intensities. Direct electron detectors are commonly used in cryo-EM imaging for their sensitivity and speed, but despite their availability are generally not used in diffraction. Primary concerns with diffraction experiments are the dynamic range and coincidence loss, which will corrupt the measurement if the flux exceeds the count rate of the detector. Here, we describe instrument setup and low-exposure MicroED data collection in electron-counting mode using K2 and K3 direct electron detectors and show that the integrated intensities can be effectively used to solve structures of two macromolecules between 1.2 Å and 2.8 Å resolution. Even though a beam stop was not used with the K3 studies we did not observe damage to the camera. As these cameras are already available in many cryo-EM facilities, this provides opportunities for users who do not have access to dedicated facilities for MicroED. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27900.map.gz emd_27900.map.gz | 2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27900-v30.xml emd-27900-v30.xml emd-27900.xml emd-27900.xml | 13.5 KB 13.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27900.png emd_27900.png | 52.9 KB | ||
| Filedesc metadata |  emd-27900.cif.gz emd-27900.cif.gz | 5.5 KB | ||
| Filedesc structureFactors |  emd_27900_sf.cif.gz emd_27900_sf.cif.gz | 167.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27900 http://ftp.pdbj.org/pub/emdb/structures/EMD-27900 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27900 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27900 | HTTPS FTP |
-Validation report
| Summary document |  emd_27900_validation.pdf.gz emd_27900_validation.pdf.gz | 578 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27900_full_validation.pdf.gz emd_27900_full_validation.pdf.gz | 577.5 KB | Display | |
| Data in XML |  emd_27900_validation.xml.gz emd_27900_validation.xml.gz | 4.4 KB | Display | |
| Data in CIF |  emd_27900_validation.cif.gz emd_27900_validation.cif.gz | 5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27900 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27900 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27900 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27900 | HTTPS FTP |
-Related structure data
| Related structure data |  8e52MC  8e53C  8e54C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27900.map.gz / Format: CCP4 / Size: 2.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27900.map.gz / Format: CCP4 / Size: 2.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 2mFo-DFc | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X: 0.6757 Å / Y: 0.6757 Å / Z: 0.6307 Å | ||||||||||||||||||||||||||||||||||||
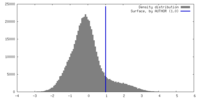
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Proteinase K
| Entire | Name: Proteinase K |
|---|---|
| Components |
|
-Supramolecule #1: Proteinase K
| Supramolecule | Name: Proteinase K / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: Serine protease |
|---|---|
| Source (natural) | Organism:  Parengyodontium album (fungus) Parengyodontium album (fungus) |
| Molecular weight | Theoretical: 28.9 KDa |
-Macromolecule #1: Proteinase K
| Macromolecule | Name: Proteinase K / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: peptidase K |
|---|---|
| Source (natural) | Organism:  Parengyodontium album (fungus) Parengyodontium album (fungus) |
| Molecular weight | Theoretical: 28.930783 KDa |
| Recombinant expression | Organism:  Parengyodontium album (fungus) Parengyodontium album (fungus) |
| Sequence | String: AAQTNAPWGL ARISSTSPGT STYYYDESAG QGSCVYVIDT GIEASHPEFE GRAQMVKTYY YSSRDGNGHG THCAGTVGSR TYGVAKKTQ LFGVKVLDDN GSGQYSTIIA GMDFVASDKN NRNCPKGVVA SLSLGGGYSS SVNSAAARLQ SSGVMVAVAA G NNNADARN ...String: AAQTNAPWGL ARISSTSPGT STYYYDESAG QGSCVYVIDT GIEASHPEFE GRAQMVKTYY YSSRDGNGHG THCAGTVGSR TYGVAKKTQ LFGVKVLDDN GSGQYSTIIA GMDFVASDKN NRNCPKGVVA SLSLGGGYSS SVNSAAARLQ SSGVMVAVAA G NNNADARN YSPASEPSVC TVGASDRYDR RSSFSNYGSV LDIFGPGTSI LSTWIGGSTR SISGTSMATP HVAGLAAYLM TL GKTTAAS ACRYIADTAN KGDLSNIPFG TVNLLAYNNY QA UniProtKB: Putative transcriptional regulator ToxR |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron crystallography |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Concentration | 50 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 10 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: LEICA PLUNGER |
| Details | Microcrystals |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77.0 K / Max: 90.0 K |
| Image recording | Film or detector model: GATAN K2 BASE (4k x 4k) / Digitization - Dimensions - Width: 1650 pixel / Digitization - Dimensions - Height: 1470 pixel / Number grids imaged: 1 / Number real images: 1 / Number diffraction images: 1600 / Average exposure time: 0.025 sec. / Average electron dose: 0.01 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: DIFFRACTION / Cs: 2.7 mm / Camera length: 590 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution method: DIFFRACTION PATTERN/LAYERLINES / Software - Name: REFMAC |
|---|---|
| Merging software list | Software - Name: AIMLESS |
| Crystallography statistics | Number intensities measured: 30326 / Number structure factors: 5453 / Fourier space coverage: 83 / R sym: 30 / R merge: 64 / Overall phase error: 28 / Overall phase residual: 0 / Phase error rejection criteria: None / High resolution: 2.7 Å / Shell - Shell ID: 1 / Shell - High resolution: 2.7 Å / Shell - Low resolution: 3.4 Å / Shell - Number structure factors: 2499 / Shell - Phase residual: 33 / Shell - Fourier space coverage: 83 / Shell - Multiplicity: 6 |
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: RIGID BODY FIT / Overall B value: 20.32 / Target criteria: Maximum liklihood |
|---|---|
| Output model |  PDB-8e52: |
 Movie
Movie Controller
Controller






 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)