[English] 日本語
 Yorodumi
Yorodumi- EMDB-27529: Cryo-EM structure of SARS-CoV-2 Omicron BA.2 spike protein in com... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2 | |||||||||
 Map data Map data | Cryo-EM structure of SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / glycoprotein / fusion protein / viral protein / Viral Protein-Immune System complex / Omicron / BA.2 / mouse / ACE2 | |||||||||
| Function / homology |  Function and homology information Function and homology informationMetabolism of Angiotensinogen to Angiotensins / ceramide biosynthetic process / positive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin ...Metabolism of Angiotensinogen to Angiotensins / ceramide biosynthetic process / positive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / maternal process involved in female pregnancy / regulation of cardiac conduction / peptidyl-dipeptidase activity / transporter activator activity / carboxypeptidase activity / angiotensin maturation / metallocarboxypeptidase activity / positive regulation of cardiac muscle contraction / negative regulation of smooth muscle cell proliferation / brush border membrane / negative regulation of ERK1 and ERK2 cascade / metallopeptidase activity / virus receptor activity / regulation of inflammatory response / symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / endopeptidase activity / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / apical plasma membrane / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / cilium / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / cell surface / negative regulation of transcription by RNA polymerase II / : / extracellular region / membrane / metal ion binding / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   | |||||||||
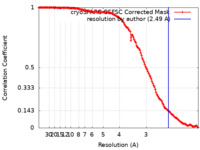
| Method | single particle reconstruction / cryo EM / Resolution: 2.49 Å | |||||||||
 Authors Authors | Zhu X / Saville JW / Mannar D / Berezuk AM / Cholak S / Tuttle KS / Vahdatihassani F / Subramaniam S | |||||||||
| Funding support |  Canada, 2 items Canada, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2023 Journal: Cell Rep / Year: 2023Title: Structural analysis of receptor engagement and antigenic drift within the BA.2 spike protein. Authors: James W Saville / Dhiraj Mannar / Xing Zhu / Alison M Berezuk / Spencer Cholak / Katharine S Tuttle / Faezeh Vahdatihassani / Sriram Subramaniam /  Abstract: The BA.2 sub-lineage of the Omicron (B.1.1.529) severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variant rapidly supplanted the original BA.1 sub-lineage in early 2022. Both lineages ...The BA.2 sub-lineage of the Omicron (B.1.1.529) severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variant rapidly supplanted the original BA.1 sub-lineage in early 2022. Both lineages threatened the efficacy of vaccine-elicited antibodies and acquired increased binding to several mammalian ACE2 receptors. Cryoelectron microscopy (cryo-EM) analysis of the BA.2 spike (S) glycoprotein in complex with mouse ACE2 (mACE2) identifies BA.1- and BA.2-mutated residues Q493R, N501Y, and Y505H as complementing non-conserved residues between human and mouse ACE2, rationalizing the enhanced S protein-mACE2 interaction for Omicron variants. Cryo-EM structures of the BA.2 S-human ACE2 complex and of the extensively mutated BA.2 amino-terminal domain (NTD) reveal a dramatic reorganization of the highly antigenic N1 loop into a β-strand, providing an explanation for decreased binding of the BA.2 S protein to antibodies isolated from BA.1-convalescent patients. Our analysis reveals structural mechanisms underlying the antigenic drift in the rapidly evolving Omicron variant landscape. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27529.map.gz emd_27529.map.gz | 123.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27529-v30.xml emd-27529-v30.xml emd-27529.xml emd-27529.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27529_fsc.xml emd_27529_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_27529.png emd_27529.png | 107.8 KB | ||
| Filedesc metadata |  emd-27529.cif.gz emd-27529.cif.gz | 6.7 KB | ||
| Others |  emd_27529_half_map_1.map.gz emd_27529_half_map_1.map.gz emd_27529_half_map_2.map.gz emd_27529_half_map_2.map.gz | 226.3 MB 226.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27529 http://ftp.pdbj.org/pub/emdb/structures/EMD-27529 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27529 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27529 | HTTPS FTP |
-Related structure data
| Related structure data |  8dm7MC  8dm1C  8dm2C  8dm3C  8dm4C  8dm5C  8dm6C  8dm8C  8dm9C  8dmaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27529.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27529.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Cryo-EM structure of SARS-CoV-2 Omicron BA.2 spike protein...
| File | emd_27529_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryo-EM structure of SARS-CoV-2 Omicron BA.2 spike protein...
| File | emd_27529_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2
| Entire | Name: SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2 |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2
| Supramolecule | Name: SARS-CoV-2 Omicron BA.2 spike protein in complex with mouse ACE2 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|
-Supramolecule #2: SARS-CoV-2 Omicron BA.2 spike protein
| Supramolecule | Name: SARS-CoV-2 Omicron BA.2 spike protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: mouse ACE2
| Supramolecule | Name: mouse ACE2 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 142.441812 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPV LPFNDGVYF ASTEKSNIIR GWIFGTTLDS KTQSLLIVNN ATNVVIKVCE FQFCNDPFLD VYYHKNNKSW MESEFRVYSS A NNCTFEYV ...String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPV LPFNDGVYF ASTEKSNIIR GWIFGTTLDS KTQSLLIVNN ATNVVIKVCE FQFCNDPFLD VYYHKNNKSW MESEFRVYSS A NNCTFEYV SQPFLMDLEG KQGNFKNLRE FVFKNIDGYF KIYSKHTPIN LGRDLPQGFS ALEPLVDLPI GINITRFQTL LA LHRSYLT PGDSSSGWTA GAAAYYVGYL QPRTFLLKYN ENGTITDAVD CALDPLSETK CTLKSFTVEK GIYQTSNFRV QPT ESIVRF PNITNLCPFD EVFNATRFAS VYAWNRKRIS NCVADYSVLY NFAPFFAFKC YGVSPTKLND LCFTNVYADS FVIR GNEVS QIAPGQTGNI ADYNYKLPDD FTGCVIAWNS NKLDSKVGGN YNYLYRLFRK SNLKPFERDI STEIYQAGNK PCNGV AGFN CYFPLRSYGF RPTYGVGHQP YRVVVLSFEL LHAPATVCGP KKSTNLVKNK CVNFNFNGLT GTGVLTESNK KFLPFQ QFG RDIADTTDAV RDPQTLEILD ITPCSFGGVS VITPGTNTSN QVAVLYQGVN CTEVPVAIHA DQLTPTWRVY STGSNVF QT RAGCLIGAEY VNNSYECDIP IGAGICASYQ TQTKSHGSAS SVASQSIIAY TMSLGAENSV AYSNNSIAIP TNFTISVT T EILPVSMTKT SVDCTMYICG DSTECSNLLL QYGSFCTQLK RALTGIAVEQ DKNTQEVFAQ VKQIYKTPPI KYFGGFNFS QILPDPSKPS KRSPIEDLLF NKVTLADAGF IKQYGDCLGD IAARDLICAQ KFNGLTVLPP LLTDEMIAQY TSALLAGTIT SGWTFGAGP ALQIPFPMQM AYRFNGIGVT QNVLYENQKL IANQFNSAIG KIQDSLSSTP SALGKLQDVV NHNAQALNTL V KQLSSKFG AISSVLNDIL SRLDPPEAEV QIDRLITGRL QSLQTYVTQQ LIRAAEIRAS ANLAATKMSE CVLGQSKRVD FC GKGYHLM SFPQSAPHGV VFLHVTYVPA QEKNFTTAPA ICHDGKAHFP REGVFVSNGT HWFVTQRNFY EPQIITTDNT FVS GNCDVV IGIVNNTVYD PLQPELDSFK EELDKYFKNH TSPDVDLGDI SGINASVVNI QKEIDRLNEV AKNLNESLID LQEL GKYEQ GSGYIPEAPR DGQAYVRKDG EWVLLSTFLG RSLEVLFQGP GHHHHHHHHS AWSHPQFEKG GGSGGGGSGG SAWSH PQFE K UniProtKB: Spike glycoprotein |
-Macromolecule #2: Angiotensin-converting enzyme 2
| Macromolecule | Name: Angiotensin-converting enzyme 2 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: angiotensin-converting enzyme 2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 71.795523 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSSSSWLLLS LVAVTTAQSL TEENAKTFLN NFNQEAEDLS YQSSLASWNY NTNITEENAQ KMSEAAAKWS AFYEEQSKTA QSFSLQEIQ TPIIKRQLQA LQQSGSSALS ADKNKQLNTI LNTMSTIYST GKVCNPKNPQ ECLLLEPGLD EIMATSTDYN S RLWAWEGW ...String: MSSSSWLLLS LVAVTTAQSL TEENAKTFLN NFNQEAEDLS YQSSLASWNY NTNITEENAQ KMSEAAAKWS AFYEEQSKTA QSFSLQEIQ TPIIKRQLQA LQQSGSSALS ADKNKQLNTI LNTMSTIYST GKVCNPKNPQ ECLLLEPGLD EIMATSTDYN S RLWAWEGW RAEVGKQLRP LYEEYVVLKN EMARANNYND YGDYWRGDYE AEGADGYNYN RNQLIEDVER TFAEIKPLYE HL HAYVRRK LMDTYPSYIS PTGCLPAHLL GDMWGRFWTN LYPLTVPFAQ KPNIDVTDAM MNQGWDAERI FQEAEKFFVS VGL PHMTQG FWANSMLTEP ADGRKVVCHP TAWDLGHGDF RIKMCTKVTM DNFLTAHHEM GHIQYDMAYA RQPFLLRNGA NEGF HEAVG EIMSLSAATP KHLKSIGLLP SDFQEDSETE INFLLKQALT IVGTLPFTYM LEKWRWMVFR GEIPKEQWMK KWWEM KREI VGVVEPLPHD ETYCDPASLF HVSNDYSFIR YYTRTIYQFQ FQEALCQAAK YNGSLHKCDI SNSTEAGQKL LKMLSL GNS EPWTKALENV VGARNMDVKP LLNYFQPLFD WLKEQNRNSF VGWNTEWSPY ADHHHHHH UniProtKB: Angiotensin-converting enzyme 2 |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 25 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)