[English] 日本語
 Yorodumi
Yorodumi- EMDB-27332: Human GABAA receptor alpha1-beta2-gamma2 subtype in complex with ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human GABAA receptor alpha1-beta2-gamma2 subtype in complex with GABA plus Zolpidem | |||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | GABAA receptor / Zolpidem / MEMBRANE PROTEIN-IMMUNE SYSTEM complex | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationbenzodiazepine receptor activity / GABA receptor complex / cellular response to histamine / GABA receptor activation / inner ear receptor cell development / inhibitory synapse assembly / GABA-A receptor activity / GABA-gated chloride ion channel activity / GABA-A receptor complex / innervation ...benzodiazepine receptor activity / GABA receptor complex / cellular response to histamine / GABA receptor activation / inner ear receptor cell development / inhibitory synapse assembly / GABA-A receptor activity / GABA-gated chloride ion channel activity / GABA-A receptor complex / innervation / postsynaptic specialization membrane / gamma-aminobutyric acid signaling pathway / synaptic transmission, GABAergic / chloride channel activity / adult behavior / Signaling by ERBB4 / cochlea development / chloride channel complex / dendrite membrane / cytoplasmic vesicle membrane / chloride transmembrane transport / post-embryonic development / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / GABA-ergic synapse / chemical synaptic transmission / dendritic spine / postsynaptic membrane / postsynapse / axon / extracellular exosome / plasma membrane Similarity search - Function | |||||||||||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||||||||||||||
 Authors Authors | Zhu S / Hibbs RE | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural and dynamic mechanisms of GABA receptor modulators with opposing activities. Authors: Shaotong Zhu / Akshay Sridhar / Jinfeng Teng / Rebecca J Howard / Erik Lindahl / Ryan E Hibbs /   Abstract: γ-Aminobutyric acid type A (GABA) receptors are pentameric ligand-gated ion channels abundant in the central nervous system and are prolific drug targets for treating anxiety, sleep disorders and ...γ-Aminobutyric acid type A (GABA) receptors are pentameric ligand-gated ion channels abundant in the central nervous system and are prolific drug targets for treating anxiety, sleep disorders and epilepsy. Diverse small molecules exert a spectrum of effects on γ-aminobutyric acid type A (GABA) receptors by acting at the classical benzodiazepine site. They can potentiate the response to GABA, attenuate channel activity, or counteract modulation by other ligands. Structural mechanisms underlying the actions of these drugs are not fully understood. Here we present two high-resolution structures of GABA receptors in complex with zolpidem, a positive allosteric modulator and heavily prescribed hypnotic, and DMCM, a negative allosteric modulator with convulsant and anxiogenic properties. These two drugs share the extracellular benzodiazepine site at the α/γ subunit interface and two transmembrane sites at β/α interfaces. Structural analyses reveal a basis for the subtype selectivity of zolpidem that underlies its clinical success. Molecular dynamics simulations provide insight into how DMCM switches from a negative to a positive modulator as a function of binding site occupancy. Together, these findings expand our understanding of how GABA receptor allosteric modulators acting through a common site can have diverging activities. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27332.map.gz emd_27332.map.gz | 10.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27332-v30.xml emd-27332-v30.xml emd-27332.xml emd-27332.xml | 23.8 KB 23.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27332.png emd_27332.png | 148 KB | ||
| Filedesc metadata |  emd-27332.cif.gz emd-27332.cif.gz | 7.5 KB | ||
| Others |  emd_27332_half_map_1.map.gz emd_27332_half_map_1.map.gz emd_27332_half_map_2.map.gz emd_27332_half_map_2.map.gz | 65.2 MB 65.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27332 http://ftp.pdbj.org/pub/emdb/structures/EMD-27332 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27332 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27332 | HTTPS FTP |
-Validation report
| Summary document |  emd_27332_validation.pdf.gz emd_27332_validation.pdf.gz | 885.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27332_full_validation.pdf.gz emd_27332_full_validation.pdf.gz | 885.4 KB | Display | |
| Data in XML |  emd_27332_validation.xml.gz emd_27332_validation.xml.gz | 12.8 KB | Display | |
| Data in CIF |  emd_27332_validation.cif.gz emd_27332_validation.cif.gz | 15.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27332 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27332 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27332 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27332 | HTTPS FTP |
-Related structure data
| Related structure data |  8dd2MC  8dd3C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27332.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27332.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.834 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_27332_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_27332_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human GABAA receptor alpha1-beta2-gamma2 subtype in complex with ...
| Entire | Name: Human GABAA receptor alpha1-beta2-gamma2 subtype in complex with GABA plus zolpidem |
|---|---|
| Components |
|
-Supramolecule #1: Human GABAA receptor alpha1-beta2-gamma2 subtype in complex with ...
| Supramolecule | Name: Human GABAA receptor alpha1-beta2-gamma2 subtype in complex with GABA plus zolpidem type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Gamma-aminobutyric acid receptor subunit beta-2
| Macromolecule | Name: Gamma-aminobutyric acid receptor subunit beta-2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.810086 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSVNDPSNMS LVKETVDRLL KGYDIRLRPD FGGPPVAVGM NIDIASIDMV SEVNMDYTLT MYFQQAWRDK RLSYNVIPLN LTLDNRVAD QLWVPDTYFL NDKKSFVHGV TVKNRMIRLH PDGTVLYGLR ITTTAACMMD LRRYPLDEQN CTLEIESYGY T TDDIEFYW ...String: QSVNDPSNMS LVKETVDRLL KGYDIRLRPD FGGPPVAVGM NIDIASIDMV SEVNMDYTLT MYFQQAWRDK RLSYNVIPLN LTLDNRVAD QLWVPDTYFL NDKKSFVHGV TVKNRMIRLH PDGTVLYGLR ITTTAACMMD LRRYPLDEQN CTLEIESYGY T TDDIEFYW RGDDNAVTGV TKIELPQFSI VDYKLITKKV VFSTGSYPRL SLSFKLKRNI GYFILQTYMP SILITILSWV SF WINYDAS AARVALGITT VLTMTTINTH LRETLPKIPY VKAIDMYLMG CFVFVFMALL EYALVNYIFF SQPARAAAID RWS RIFFPV VFSFFNIVYW LYYVNVDGSG ATNFSLLKQA GDVEENPG UniProtKB: Gamma-aminobutyric acid receptor subunit beta-2, Gamma-aminobutyric acid receptor subunit beta-2 |
-Macromolecule #2: Gamma-aminobutyric acid receptor subunit alpha-1
| Macromolecule | Name: Gamma-aminobutyric acid receptor subunit alpha-1 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.061211 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QPSLQDELKD NTTVFTRILD RLLDGYDNRL RPGLGERVTE VKTDIFVTSF GPVSDHDMEY TIDVFFRQSW KDERLKFKGP MTVLRLNNL MASKIWTPDT FFHNGKKSVA HNMTMPNKLL RITEDGTLLY TMRLTVRAEC PMHLEDFPMD AHACPLKFGS Y AYTRAEVV ...String: QPSLQDELKD NTTVFTRILD RLLDGYDNRL RPGLGERVTE VKTDIFVTSF GPVSDHDMEY TIDVFFRQSW KDERLKFKGP MTVLRLNNL MASKIWTPDT FFHNGKKSVA HNMTMPNKLL RITEDGTLLY TMRLTVRAEC PMHLEDFPMD AHACPLKFGS Y AYTRAEVV YEWTREPARS VVVAEDGSRL NQYDLLGQTV DSGIVQSSTG EYVVMTTHFH LKRKIGYFVI QTYLPCIMTV IL SQVSFWL NRESVPARTV FGVTTVLTMT TLSISARNSL PKVAYATAMD WFIAVCYAFV FSALIEFATV NYFTKSQPAR AAK IDRLSR IAFPLLFGIF NLVYWATYLN REPQLKAPTP HQ UniProtKB: Gamma-aminobutyric acid receptor subunit alpha-1 |
-Macromolecule #3: Gamma-aminobutyric acid receptor subunit gamma-2
| Macromolecule | Name: Gamma-aminobutyric acid receptor subunit gamma-2 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.673109 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: WSHPQFEKGG GSGGGSGGSS AWSHPQFEKL EVLFQGPQKS DDDYEDYASN KTWVLTPKVP EGDVTVILNN LLEGYDNKLR PDIGVKPTL IHTDMYVNSI GPVNAINMEY TIDIFFAQTW YDRRLKFNST IKVLRLNSNM VGKIWIPDTF FRNSKKADAH W ITTPNRML ...String: WSHPQFEKGG GSGGGSGGSS AWSHPQFEKL EVLFQGPQKS DDDYEDYASN KTWVLTPKVP EGDVTVILNN LLEGYDNKLR PDIGVKPTL IHTDMYVNSI GPVNAINMEY TIDIFFAQTW YDRRLKFNST IKVLRLNSNM VGKIWIPDTF FRNSKKADAH W ITTPNRML RIWNDGRVLY TLRLTIDAEC QLQLHNFPMD EHSCPLEFSS YGYPREEIVY QWKRSSVEVG DTRSWRLYQF SF VGLRNTT EVVKTTSGDY VVMSVYFDLS RRMGYFTIQT YIPCTLIVVL SWVSFWINKD AVPARTSLGI TTVLTMTTLS TIA RKSLPK VSYVTAMDLF VSVCFIFVFS ALVEYGTLHY FVSSQPARAA KMDSYARIFF PTAFCLFNLV YWVSYLYLSR GSGA TNFSL LKQAGDVEEN PG UniProtKB: Gamma-aminobutyric acid receptor subunit gamma-2 |
-Macromolecule #4: Kappa Fab Light Chain
| Macromolecule | Name: Kappa Fab Light Chain / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.505943 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: NIVMTQSPKS MSMSVGERVT LSCKASEYVG TYVSWYQQKP EQSPKLLIYG ASNRYTGVPD RFTGSGSATD FTLTIGSVQA EDLADYHCG QSYSYPTFGA GTKLELKRAD AAPTVSIFPP SSEQLTSGGA SVVCFLNNFY PKDINVKWKI DGSERQNGVL N SWTDQDSK ...String: NIVMTQSPKS MSMSVGERVT LSCKASEYVG TYVSWYQQKP EQSPKLLIYG ASNRYTGVPD RFTGSGSATD FTLTIGSVQA EDLADYHCG QSYSYPTFGA GTKLELKRAD AAPTVSIFPP SSEQLTSGGA SVVCFLNNFY PKDINVKWKI DGSERQNGVL N SWTDQDSK DSTYSMSSTL TLTKDEYERH NSYTCEATHK TSTSPIVKSF NRNEC |
-Macromolecule #5: IgG2b Fab Heavy Chain
| Macromolecule | Name: IgG2b Fab Heavy Chain / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 49.811043 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EVQLQQSGAE LVKPGASVKL SCTASGFNIK DTYMYWVKQR PEQGLEWIGR IDPANGDTKY DPKFQGKATI TTDTFSNTAY LQLSSLTSE DTAVYYCARK GLRWAMDYWG QGTSVTVSTA KTTPPSVYPL APGCGDTTGS SVTLGCLVKG YFPESVTVTW N SGSLSSSV ...String: EVQLQQSGAE LVKPGASVKL SCTASGFNIK DTYMYWVKQR PEQGLEWIGR IDPANGDTKY DPKFQGKATI TTDTFSNTAY LQLSSLTSE DTAVYYCARK GLRWAMDYWG QGTSVTVSTA KTTPPSVYPL APGCGDTTGS SVTLGCLVKG YFPESVTVTW N SGSLSSSV HTFPALLQSG LYTMSSSVTV PSSTWPSQTV TCSVAHPASS TTVDKKLEPS GPISTINPCP PCKECHKCPA PN LEGGPSV FIFPPNIKDV LMISLTPKVT CVVVDVSEDD PDVQISWFVN NVEVHTAQTQ THREDYNSTI RVVSTLPIQH QDW MSGKEF KCKVNNKDLP SPIERTISKI KGLVRAPQVY ILPPPAEQLS RKDVSLTCLV VGFNPGDISV EWTSNGHTEE NYKD TAPVL DSDGSYFIYS KLNMKTSKWE KTDSFSCNVR HEGLKNYYLK KTISRSPGK |
-Macromolecule #9: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 9 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #10: Zolpidem
| Macromolecule | Name: Zolpidem / type: ligand / ID: 10 / Number of copies: 3 / Formula: R5R |
|---|---|
| Molecular weight | Theoretical: 307.39 Da |
| Chemical component information | 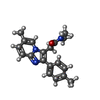 ChemComp-R5R: |
-Macromolecule #11: GAMMA-AMINO-BUTANOIC ACID
| Macromolecule | Name: GAMMA-AMINO-BUTANOIC ACID / type: ligand / ID: 11 / Number of copies: 2 / Formula: ABU |
|---|---|
| Molecular weight | Theoretical: 103.12 Da |
| Chemical component information |  ChemComp-ABU: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 3.5 second blot. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 14621 / Average electron dose: 55.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: De novo initial model from Relion |
|---|---|
| Final reconstruction | Number classes used: 1 / Applied symmetry - Point group: C1 (asymmetric) / Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 413941 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)