[English] 日本語
 Yorodumi
Yorodumi- EMDB-26310: Cryo-electron microscopy structure of human mt-SerRS in complex w... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-electron microscopy structure of human mt-SerRS in complex with mt-tRNA (GCU) | |||||||||||||||
 Map data Map data | Cryo-electron microscopy structure of human mt-SerRS in complex with mt-tRNA (GCU) | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | tRNA / SerRS / mitochondria / aminoacylation / Ligase-RNA complex | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationmitochondrial seryl-tRNA aminoacylation / Mitochondrial tRNA aminoacylation / serine-tRNA ligase / serine-tRNA ligase activity / seryl-tRNA aminoacylation / tRNA binding / mitochondrial matrix / mitochondrion / RNA binding / ATP binding Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||||||||
 Authors Authors | Hirschi M / Kuhle B | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural basis for shape-selective recognition and aminoacylation of a D-armless human mitochondrial tRNA. Authors: Bernhard Kuhle / Marscha Hirschi / Lili K Doerfel / Gabriel C Lander / Paul Schimmel /  Abstract: Human mitochondrial gene expression relies on the specific recognition and aminoacylation of mitochondrial tRNAs (mtRNAs) by nuclear-encoded mitochondrial aminoacyl-tRNA synthetases (mt-aaRSs). ...Human mitochondrial gene expression relies on the specific recognition and aminoacylation of mitochondrial tRNAs (mtRNAs) by nuclear-encoded mitochondrial aminoacyl-tRNA synthetases (mt-aaRSs). Despite their essential role in cellular energy homeostasis, strong mutation pressure and genetic drift have led to an unparalleled sequence erosion of animal mtRNAs. The structural and functional consequences of this erosion are not understood. Here, we present cryo-EM structures of the human mitochondrial seryl-tRNA synthetase (mSerRS) in complex with mtRNA. These structures reveal a unique mechanism of substrate recognition and aminoacylation. The mtRNA is highly degenerated, having lost the entire D-arm, tertiary core, and stable L-shaped fold that define canonical tRNAs. Instead, mtRNA evolved unique structural innovations, including a radically altered T-arm topology that serves as critical identity determinant in an unusual shape-selective readout mechanism by mSerRS. Our results provide a molecular framework to understand the principles of mito-nuclear co-evolution and specialized mechanisms of tRNA recognition in mammalian mitochondrial gene expression. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26310.map.gz emd_26310.map.gz | 16.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26310-v30.xml emd-26310-v30.xml emd-26310.xml emd-26310.xml | 11.4 KB 11.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_26310.png emd_26310.png | 96.2 KB | ||
| Filedesc metadata |  emd-26310.cif.gz emd-26310.cif.gz | 5.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26310 http://ftp.pdbj.org/pub/emdb/structures/EMD-26310 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26310 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26310 | HTTPS FTP |
-Related structure data
| Related structure data |  7u2aMC  7tzbC  7u2bC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26310.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26310.map.gz / Format: CCP4 / Size: 18.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-electron microscopy structure of human mt-SerRS in complex with mt-tRNA (GCU) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Human mtSerRS in complex with mt-tRNA(GCU) and SerSA
| Entire | Name: Human mtSerRS in complex with mt-tRNA(GCU) and SerSA |
|---|---|
| Components |
|
-Supramolecule #1: Human mtSerRS in complex with mt-tRNA(GCU) and SerSA
| Supramolecule | Name: Human mtSerRS in complex with mt-tRNA(GCU) and SerSA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: RNA (38-MER)
| Macromolecule | Name: RNA (38-MER) / type: rna / ID: 1 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.089238 KDa |
| Sequence | String: GAGAAAGCUC CAUGUCUAAC AACAUGGCUU UCUCACCA |
-Macromolecule #2: Serine--tRNA ligase, mitochondrial
| Macromolecule | Name: Serine--tRNA ligase, mitochondrial / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: serine-tRNA ligase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 58.358391 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAASMARRLW PLLTRRGFRP RGGCISNDSP RRSFTTEKRN RNLLYEYARE GYSALPQLDI ERFCACPEEA AHALELRKGE LRSADLPAI ISTWQELRQL QEQIRSLEEE KAAVTEAVRA LLANQDSGEV QQDPKYQGLR ARGREIRKEL VHLYPREAQL E EQFYLQAL ...String: MAASMARRLW PLLTRRGFRP RGGCISNDSP RRSFTTEKRN RNLLYEYARE GYSALPQLDI ERFCACPEEA AHALELRKGE LRSADLPAI ISTWQELRQL QEQIRSLEEE KAAVTEAVRA LLANQDSGEV QQDPKYQGLR ARGREIRKEL VHLYPREAQL E EQFYLQAL KLPNQTHPDV PVGDESQARV LHMVGDKPVF SFQPRGHLEI GEKLDIIRQK RLSHVSGHRS YYLRGAGALL QH GLVNFTF NKLLRRGFTP MTVPDLLRGA VFEGCGMTPN ANPSQIYNID PARFKDLNLA GTAEVGLAGY FMDHTVAFRD LPV RMVCSS TCYRAETNTG QEPRGLYRVH HFTKVEMFGV TGPGLEQSSQ LLEEFLSLQM EILTELGLHF RVLDMPTQEL GLPA YRKFD IEAWMPGRGR FGEVTSASNC TDFQSRRLHI MFQTEAGELQ FAHTVNATAC AVPRLLIALL ESNQQKDGSV LVPPA LQSY LGTDRITAPT HVPLQYIGPN QPRKPGLPGQ PAVS UniProtKB: Serine--tRNA ligase, mitochondrial |
-Macromolecule #3: 5'-O-(N-(L-SERYL)-SULFAMOYL)ADENOSINE
| Macromolecule | Name: 5'-O-(N-(L-SERYL)-SULFAMOYL)ADENOSINE / type: ligand / ID: 3 / Number of copies: 2 / Formula: SSA |
|---|---|
| Molecular weight | Theoretical: 433.397 Da |
| Chemical component information | 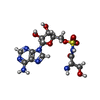 ChemComp-SSA: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 66.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 152066 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




















