[English] 日本語
 Yorodumi
Yorodumi- EMDB-25690: SARS-CoV-2 S (Spike Glycoprotein) D614G with Three (3) RBDs Up, B... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 S (Spike Glycoprotein) D614G with Three (3) RBDs Up, Bound to Antibody 2-7 scFv, local refinement map | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
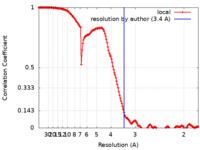
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Byrne PO / McLellan JS | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: IgG-like bispecific antibodies with potent and synergistic neutralization against circulating SARS-CoV-2 variants of concern. Authors: Matthew R Chang / Luke Tomasovic / Natalia A Kuzmina / Adam J Ronk / Patrick O Byrne / Rebecca Johnson / Nadia Storm / Eduardo Olmedillas / Yixuan J Hou / Alexandra Schäfer / Sarah R Leist ...Authors: Matthew R Chang / Luke Tomasovic / Natalia A Kuzmina / Adam J Ronk / Patrick O Byrne / Rebecca Johnson / Nadia Storm / Eduardo Olmedillas / Yixuan J Hou / Alexandra Schäfer / Sarah R Leist / Longping V Tse / Hanzhong Ke / Christian Coherd / Katrina Nguyen / Maliwan Kamkaew / Anna Honko / Quan Zhu / Galit Alter / Erica Ollmann Saphire / Jason S McLellan / Anthony Griffiths / Ralph S Baric / Alexander Bukreyev / Wayne A Marasco /  Abstract: Monoclonal antibodies are a promising approach to treat COVID-19, however the emergence of SARS-CoV-2 variants has challenged the efficacy and future of these therapies. Antibody cocktails are being ...Monoclonal antibodies are a promising approach to treat COVID-19, however the emergence of SARS-CoV-2 variants has challenged the efficacy and future of these therapies. Antibody cocktails are being employed to mitigate these challenges, but neutralization escape remains a major challenge and alternative strategies are needed. Here we present two anti-SARS-CoV-2 spike binding antibodies, one Class 1 and one Class 4, selected from our non-immune human single-chain variable fragment (scFv) phage library, that are engineered into four, fully-human IgG-like bispecific antibodies (BsAb). Prophylaxis of hACE2 mice and post-infection treatment of golden hamsters demonstrates the efficacy of the monospecific antibodies against the original Wuhan strain, while promising in vitro results with the BsAbs demonstrate enhanced binding and distinct synergistic effects on neutralizing activity against circulating variants of concern. In particular, one BsAb engineered in a tandem scFv-Fc configuration shows synergistic neutralization activity against several variants of concern including B.1.617.2. This work provides evidence that synergistic neutralization can be achieved using a BsAb scaffold, and serves as a foundation for the future development of broadly reactive BsAbs against emerging variants of concern. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25690.map.gz emd_25690.map.gz | 160.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25690-v30.xml emd-25690-v30.xml emd-25690.xml emd-25690.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25690_fsc.xml emd_25690_fsc.xml | 15.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_25690.png emd_25690.png | 74.8 KB | ||
| Masks |  emd_25690_msk_1.map emd_25690_msk_1.map | 325 MB |  Mask map Mask map | |
| Others |  emd_25690_additional_1.map.gz emd_25690_additional_1.map.gz emd_25690_half_map_1.map.gz emd_25690_half_map_1.map.gz emd_25690_half_map_2.map.gz emd_25690_half_map_2.map.gz | 306.5 MB 300.9 MB 300.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25690 http://ftp.pdbj.org/pub/emdb/structures/EMD-25690 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25690 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25690 | HTTPS FTP |
-Validation report
| Summary document |  emd_25690_validation.pdf.gz emd_25690_validation.pdf.gz | 789 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25690_full_validation.pdf.gz emd_25690_full_validation.pdf.gz | 788.6 KB | Display | |
| Data in XML |  emd_25690_validation.xml.gz emd_25690_validation.xml.gz | 23.7 KB | Display | |
| Data in CIF |  emd_25690_validation.cif.gz emd_25690_validation.cif.gz | 30.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25690 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25690 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25690 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25690 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25690.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25690.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.9 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25690_msk_1.map emd_25690_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_25690_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_25690_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_25690_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of SARS CoV-2 S, Spike Glycoprotein, variant D614G with A...
| Entire | Name: Complex of SARS CoV-2 S, Spike Glycoprotein, variant D614G with Antibody 2-7 scFv |
|---|---|
| Components |
|
-Supramolecule #1: Complex of SARS CoV-2 S, Spike Glycoprotein, variant D614G with A...
| Supramolecule | Name: Complex of SARS CoV-2 S, Spike Glycoprotein, variant D614G with Antibody 2-7 scFv type: complex / ID: 1 / Chimera: Yes / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 180 sec. / Pretreatment - Atmosphere: OTHER | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)