+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Strand-transfer complex of TnsB from ShCAST | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CAST / transposase / strand-transfer complex / CRISPR / Cas / DNA BINDING PROTEIN-DNA complex | |||||||||
| Biological species |  [Scytonema hofmanni] UTEX 2349 (bacteria) / synthetic construct (others) [Scytonema hofmanni] UTEX 2349 (bacteria) / synthetic construct (others) | |||||||||
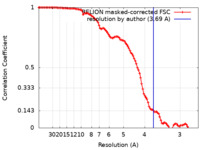
| Method | single particle reconstruction / cryo EM / Resolution: 3.69 Å | |||||||||
 Authors Authors | Park J / Tsai AWT | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Mechanistic details of CRISPR-associated transposon recruitment and integration revealed by cryo-EM. Authors: Jung-Un Park / Amy Wei-Lun Tsai / Tiffany H Chen / Joseph E Peters / Elizabeth H Kellogg /  Abstract: CRISPR-associated transposons (CASTs) are Tn7-like elements that are capable of RNA-guided DNA integration. Although structural data are known for nearly all core transposition components, the ...CRISPR-associated transposons (CASTs) are Tn7-like elements that are capable of RNA-guided DNA integration. Although structural data are known for nearly all core transposition components, the transposase component, TnsB, remains uncharacterized. Using cryo-electron microscopy (cryo-EM) structure determination, we reveal the conformation of TnsB during transposon integration for the type V-K CAST system from (ShCAST). Our structure of TnsB is a tetramer, revealing strong mechanistic relationships with the overall architecture of RNaseH transposases/integrases in general, and in particular the MuA transposase from bacteriophage Mu. However, key structural differences in the C-terminal domains indicate that TnsB's tetrameric architecture is stabilized by a different set of protein-protein interactions compared with MuA. We describe the base-specific interactions along the TnsB binding site, which explain how different CAST elements can function on cognate mobile elements independent of one another. We observe that melting of the 5' nontransferred strand of the transposon end is a structural feature stabilized by TnsB and furthermore is crucial for donor-DNA integration. Although not observed in the TnsB strand-transfer complex, the C-terminal end of TnsB serves a crucial role in transposase recruitment to the target site. The C-terminal end of TnsB adopts a short, structured 15-residue "hook" that decorates TnsC filaments. Unlike full-length TnsB, C-terminal fragments do not appear to stimulate filament disassembly using two different assays, suggesting that additional interactions between TnsB and TnsC are required for redistributing TnsC to appropriate targets. The structural information presented here will help guide future work in modifying these important systems as programmable gene integration tools. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25455.map.gz emd_25455.map.gz | 7.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25455-v30.xml emd-25455-v30.xml emd-25455.xml emd-25455.xml | 23.7 KB 23.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25455_fsc.xml emd_25455_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_25455.png emd_25455.png | 65.7 KB | ||
| Masks |  emd_25455_msk_1.map emd_25455_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-25455.cif.gz emd-25455.cif.gz | 6.7 KB | ||
| Others |  emd_25455_additional_1.map.gz emd_25455_additional_1.map.gz emd_25455_half_map_1.map.gz emd_25455_half_map_1.map.gz emd_25455_half_map_2.map.gz emd_25455_half_map_2.map.gz | 79.3 MB 79.4 MB 79.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25455 http://ftp.pdbj.org/pub/emdb/structures/EMD-25455 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25455 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25455 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25455.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25455.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.33 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25455_msk_1.map emd_25455_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: full map
| File | emd_25455_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | full map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap2
| File | emd_25455_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap1
| File | emd_25455_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : strand-transfer complex of TnsB from ShCAST element
| Entire | Name: strand-transfer complex of TnsB from ShCAST element |
|---|---|
| Components |
|
-Supramolecule #1: strand-transfer complex of TnsB from ShCAST element
| Supramolecule | Name: strand-transfer complex of TnsB from ShCAST element / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  [Scytonema hofmanni] UTEX 2349 (bacteria) [Scytonema hofmanni] UTEX 2349 (bacteria) |
| Molecular weight | Theoretical: 350 KDa |
-Macromolecule #1: STC_LE_For
| Macromolecule | Name: STC_LE_For / type: dna / ID: 1 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 21.57983 KDa |
| Sequence | String: (DA)(DT)(DG)(DA)(DC)(DA)(DT)(DT)(DA)(DA) (DT)(DC)(DT)(DG)(DT)(DC)(DA)(DC)(DC)(DG) (DA)(DC)(DG)(DA)(DC)(DA)(DG)(DA)(DT) (DA)(DA)(DT)(DT)(DT)(DG)(DT)(DC)(DA)(DC) (DT) (DG)(DT)(DA)(DC)(DA)(DG) ...String: (DA)(DT)(DG)(DA)(DC)(DA)(DT)(DT)(DA)(DA) (DT)(DC)(DT)(DG)(DT)(DC)(DA)(DC)(DC)(DG) (DA)(DC)(DG)(DA)(DC)(DA)(DG)(DA)(DT) (DA)(DA)(DT)(DT)(DT)(DG)(DT)(DC)(DA)(DC) (DT) (DG)(DT)(DA)(DC)(DA)(DG)(DG)(DC) (DC)(DC)(DT)(DA)(DG)(DG)(DT)(DC)(DT)(DA) (DC)(DG) (DG)(DT)(DT)(DA)(DG)(DA)(DG) (DG)(DC)(DT) |
-Macromolecule #2: STC_LE_Rev1
| Macromolecule | Name: STC_LE_Rev1 / type: dna / ID: 2 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 13.905951 KDa |
| Sequence | String: (DT)(DG)(DT)(DA)(DC)(DA)(DG)(DT)(DG)(DA) (DC)(DA)(DA)(DA)(DT)(DT)(DA)(DT)(DC)(DT) (DG)(DT)(DC)(DG)(DT)(DC)(DG)(DG)(DT) (DG)(DA)(DC)(DA)(DG)(DA)(DT)(DT)(DA)(DA) (DT) (DG)(DT)(DC)(DA)(DT) |
-Macromolecule #3: STC_LE_Rev2
| Macromolecule | Name: STC_LE_Rev2 / type: dna / ID: 3 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 6.062945 KDa |
| Sequence | String: (DA)(DG)(DC)(DC)(DT)(DC)(DT)(DA)(DA)(DC) (DC)(DG)(DT)(DA)(DG)(DA)(DC)(DC)(DT)(DA) |
-Macromolecule #4: TnsB
| Macromolecule | Name: TnsB / type: protein_or_peptide / ID: 4 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  [Scytonema hofmanni] UTEX 2349 (bacteria) [Scytonema hofmanni] UTEX 2349 (bacteria) |
| Molecular weight | Theoretical: 66.63707 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNSQQNPDLA VHPLAIPMEG LLGESATTLE KNVIATQLSE EAQVKLEVIQ SLLEPCDRTT YGQKLREAAE KLNVSLRTVQ RLVKNWEQD GLVGLTQTSR ADKGKHRIGE FWENFITKTY KEGNKGSKRM TPKQVALRVE AKARELKDSK PPNYKTVLRV L APILEKQQ ...String: MNSQQNPDLA VHPLAIPMEG LLGESATTLE KNVIATQLSE EAQVKLEVIQ SLLEPCDRTT YGQKLREAAE KLNVSLRTVQ RLVKNWEQD GLVGLTQTSR ADKGKHRIGE FWENFITKTY KEGNKGSKRM TPKQVALRVE AKARELKDSK PPNYKTVLRV L APILEKQQ KAKSIRSPGW RGTTLSVKTR EGKDLSVDYS NHVWQCDHTR VDVLLVDQHG EILSRPWLTT VIDTYSRCIM GI NLGFDAP SSGVVALALR HAILPKRYGS EYKLHCEWGT YGKPEHFYTD GGKDFRSNHL SQIGAQLGFV CHLRDRPSEG GVV ERPFKT LNDQLFSTLP GYTGSNVQER PEDAEKDARL TLRELEQLLV RYIVDRYNQS IDARMGDQTR FERWEAGLPT VPVP IPERD LDICLMKQSR RTVQRGGCLQ FQNLMYRGEY LAGYAGETVN LRFDPRDITT ILVYRQENNQ EVFLTRAHAQ GLETE QLAL DEAEAASRRL RTAGKTISNQ SLLQEVVDRD ALVATKKSRK ERQKLEQTVL RSAAVDESNR ESLPSQIVEP DEVEST ETV HSQYEDIEVW DYEQLREEYG F |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 49.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7svw: |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)