+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Rad24-RFC ADP state | ||||||||||||
 Map data Map data | Rad24-RFC ADP state | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | DNA damage / DNA replication / DNA sliding clamp / REPLICATION | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationRad17 RFC-like complex / : / Elg1 RFC-like complex / DNA replication factor C complex / Ctf18 RFC-like complex / Polymerase switching / Translesion synthesis by REV1 / : / : / : ...Rad17 RFC-like complex / : / Elg1 RFC-like complex / DNA replication factor C complex / Ctf18 RFC-like complex / Polymerase switching / Translesion synthesis by REV1 / : / : / : / DNA clamp loader activity / DNA replication checkpoint signaling / : / Activation of ATR in response to replication stress / mitotic DNA replication checkpoint signaling / reciprocal meiotic recombination / mitotic sister chromatid cohesion / sister chromatid cohesion / leading strand elongation / Gap-filling DNA repair synthesis and ligation in TC-NER / Dual incision in TC-NER / mismatch repair / DNA damage checkpoint signaling / nucleotide-excision repair / DNA-templated DNA replication / DNA repair / chromatin binding / ATP hydrolysis activity / DNA binding / ATP binding / nucleus / cytosol Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.73 Å | ||||||||||||
 Authors Authors | Castaneda JC / Schrecker M | ||||||||||||
| Funding support | 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: Mechanisms of loading and release of the 9-1-1 checkpoint clamp. Authors: Juan C Castaneda / Marina Schrecker / Dirk Remus / Richard K Hite /  Abstract: Single-stranded or double-stranded DNA junctions with recessed 5' ends serve as loading sites for the checkpoint clamp, 9-1-1, which mediates activation of the apical checkpoint kinase, ATR. However, ...Single-stranded or double-stranded DNA junctions with recessed 5' ends serve as loading sites for the checkpoint clamp, 9-1-1, which mediates activation of the apical checkpoint kinase, ATR. However, the basis for 9-1-1's recruitment to 5' junctions is unclear. Here, we present structures of the yeast checkpoint clamp loader, Rad24-replication factor C (RFC), in complex with 9-1-1 and a 5' junction and in a post-ATP-hydrolysis state. Unexpectedly, 9-1-1 adopts both closed and planar open states in the presence of Rad24-RFC and DNA. Moreover, Rad24-RFC associates with the DNA junction in the opposite orientation of processivity clamp loaders with Rad24 exclusively coordinating the double-stranded region. ATP hydrolysis stimulates conformational changes in Rad24-RFC, leading to disengagement of DNA-loaded 9-1-1. Together, these structures explain 9-1-1's recruitment to 5' junctions and reveal new principles of sliding clamp loading. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25426.map.gz emd_25426.map.gz | 108.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25426-v30.xml emd-25426-v30.xml emd-25426.xml emd-25426.xml | 19.4 KB 19.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25426.png emd_25426.png | 119.7 KB | ||
| Filedesc metadata |  emd-25426.cif.gz emd-25426.cif.gz | 7.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25426 http://ftp.pdbj.org/pub/emdb/structures/EMD-25426 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25426 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25426 | HTTPS FTP |
-Related structure data
| Related structure data |  7steMC  7st9C  7stbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25426.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25426.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Rad24-RFC ADP state | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.826 Å | ||||||||||||||||||||||||||||||||||||
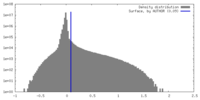
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Rad24-RFC:9-1-1:DNA
| Entire | Name: Rad24-RFC:9-1-1:DNA |
|---|---|
| Components |
|
-Supramolecule #1: Rad24-RFC:9-1-1:DNA
| Supramolecule | Name: Rad24-RFC:9-1-1:DNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Replication factor C subunit 4
| Macromolecule | Name: Replication factor C subunit 4 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 36.201039 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSKTLSLQLP WVEKYRPQVL SDIVGNKETI DRLQQIAKDG NMPHMIISGM PGIGKTTSVH CLAHELLGRS YADGVLELNA SDDRGIDVV RNQIKHFAQK KLHLPPGKHK IVILDEADSM TAGAQQALRR TMELYSNSTR FAFACNQSNK IIEPLQSRCA I LRYSKLSD ...String: MSKTLSLQLP WVEKYRPQVL SDIVGNKETI DRLQQIAKDG NMPHMIISGM PGIGKTTSVH CLAHELLGRS YADGVLELNA SDDRGIDVV RNQIKHFAQK KLHLPPGKHK IVILDEADSM TAGAQQALRR TMELYSNSTR FAFACNQSNK IIEPLQSRCA I LRYSKLSD EDVLKRLLQI IKLEDVKYTN DGLEAIIFTA EGDMRQAINN LQSTVAGHGL VNADNVFKIV DSPHPLIVKK ML LASNLED SIQILRTDLW KKGYSSIDIV TTSFRVTKNL AQVKESVRLE MIKEIGLTHM RILEGVGTYL QLASMLAKIH KLN NKA UniProtKB: Replication factor C subunit 4 |
-Macromolecule #2: Replication factor C subunit 3
| Macromolecule | Name: Replication factor C subunit 3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 38.155414 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSTSTEKRSK ENLPWVEKYR PETLDEVYGQ NEVITTVRKF VDEGKLPHLL FYGPPGTGKT STIVALAREI YGKNYSNMVL ELNASDDRG IDVVRNQIKD FASTRQIFSK GFKLIILDEA DAMTNAAQNA LRRVIERYTK NTRFCVLANY AHKLTPALLS R CTRFRFQP ...String: MSTSTEKRSK ENLPWVEKYR PETLDEVYGQ NEVITTVRKF VDEGKLPHLL FYGPPGTGKT STIVALAREI YGKNYSNMVL ELNASDDRG IDVVRNQIKD FASTRQIFSK GFKLIILDEA DAMTNAAQNA LRRVIERYTK NTRFCVLANY AHKLTPALLS R CTRFRFQP LPQEAIERRI ANVLVHEKLK LSPNAEKALI ELSNGDMRRV LNVLQSCKAT LDNPDEDEIS DDVIYECCGA PR PSDLKAV LKSILEDDWG TAHYTLNKVR SAKGLALIDL IEGIVKILED YELQNEETRV HLLTKLADIE YSISKGGNDQ IQG SAVIGA IKASFENETV KAN UniProtKB: Replication factor C subunit 3 |
-Macromolecule #3: Replication factor C subunit 2
| Macromolecule | Name: Replication factor C subunit 2 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 39.794473 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MFEGFGPNKK RKISKLAAEQ SLAQQPWVEK YRPKNLDEVT AQDHAVTVLK KTLKSANLPH MLFYGPPGTG KTSTILALTK ELYGPDLMK SRILELNASD ERGISIVREK VKNFARLTVS KPSKHDLENY PCPPYKIIIL DEADSMTADA QSALRRTMET Y SGVTRFCL ...String: MFEGFGPNKK RKISKLAAEQ SLAQQPWVEK YRPKNLDEVT AQDHAVTVLK KTLKSANLPH MLFYGPPGTG KTSTILALTK ELYGPDLMK SRILELNASD ERGISIVREK VKNFARLTVS KPSKHDLENY PCPPYKIIIL DEADSMTADA QSALRRTMET Y SGVTRFCL ICNYVTRIID PLASRCSKFR FKALDASNAI DRLRFISEQE NVKCDDGVLE RILDISAGDL RRGITLLQSA SK GAQYLGD GKNITSTQVE ELAGVVPHDI LIEIVEKVKS GDFDEIKKYV NTFMKSGWSA ASVVNQLHEY YITNDNFDTN FKN QISWLL FTTDSRLNNG TNEHIQLLNL LVKISQL UniProtKB: Replication factor C subunit 2 |
-Macromolecule #4: Replication factor C subunit 5
| Macromolecule | Name: Replication factor C subunit 5 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 39.993582 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSLWVDKYRP KSLNALSHNE ELTNFLKSLS DQPRDLPHLL LYGPNGTGKK TRCMALLESI FGPGVYRLKI DVRQFVTASN RKLELNVVS SPYHLEITPS DMGNNDRIVI QELLKEVAQM EQVDFQDSKD GLAHRYKCVI INEANSLTKD AQAALRRTME K YSKNIRLI ...String: MSLWVDKYRP KSLNALSHNE ELTNFLKSLS DQPRDLPHLL LYGPNGTGKK TRCMALLESI FGPGVYRLKI DVRQFVTASN RKLELNVVS SPYHLEITPS DMGNNDRIVI QELLKEVAQM EQVDFQDSKD GLAHRYKCVI INEANSLTKD AQAALRRTME K YSKNIRLI MVCDSMSPII APIKSRCLLI RCPAPSDSEI STILSDVVTN ERIQLETKDI LKRIAQASNG NLRVSLLMLE SM ALNNELA LKSSSPIIKP DWIIVIHKLT RKIVKERSVN SLIECRAVLY DLLAHCIPAN IILKELTFSL LDVETLNTTN KSS IIEYSS VFDERLSLGN KAIFHLEGFI AKVMCCLD UniProtKB: Replication factor C subunit 5 |
-Macromolecule #5: Checkpoint protein RAD24
| Macromolecule | Name: Checkpoint protein RAD24 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 80.096828 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDSTNLNKRP LLQYSLSSLG SQITKWSSSR PTSPVRKARS TENDFLSKQD TSSILPSIND DGGEQWYEKF KPNCLEQVAI HKRKLKDVQ EALDAMFLPN AKHRILLLSG PSGCSKSTVI KELSKILVPK YRQNSNGTSF RSTPNEHKVT EFRGDCIVND L PQMESFSE ...String: MDSTNLNKRP LLQYSLSSLG SQITKWSSSR PTSPVRKARS TENDFLSKQD TSSILPSIND DGGEQWYEKF KPNCLEQVAI HKRKLKDVQ EALDAMFLPN AKHRILLLSG PSGCSKSTVI KELSKILVPK YRQNSNGTSF RSTPNEHKVT EFRGDCIVND L PQMESFSE FLKGARYLVM SNLSLILIED LPNVFHIDTR RRFQQLILQW LYSSEPLLPP LVICITECEI PENDNNYRKF GI DYTFSAE TIMNKEILMH PRLKRIKFNP INSTLLKKHL KFICVQNMKM LKEKNKWNKR QEVIDYIAQE TGDIRSAITT LQF WATSSG SLPISTREST ISYFHAIGKV IHGSHSTNND NEMINNLFEN SNNLLSKEDF KLGILENYNT FNKGEFSISD ASSI VDCLS ECDNMNGLPE SNEYGLREVR KTFRNISKQG HNHGTVYFPR EWKVRKLQNS FKVQAEDWLN VSLYKYNAVH SFRNI TLEF GYYAPLIRKC QSYKKKYILY YLKNLPSGSS GPKQTMDKFS DIMKVENGID VVDRIGGPIE ALSVEDGLAP LMDNDS NNC DHLEDQKKER DRRLRMLIDQ YERNVMMAND DLEDEETSFN DDPIVDSDSD NSNNIGNETF GRSDEDESLC EILSQRQ PR KAPVISESLS DSDLEILGLN LEVLFQGPGG DYKDDDDKDY KDDDDKDYKD DDDK UniProtKB: Checkpoint protein RAD24 |
-Macromolecule #6: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 4 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #7: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 7 / Number of copies: 1 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #8: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 8 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.0 mg/mL | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.6 Component:
| ||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GRAPHENE OXIDE / Mesh: 400 / Support film - Material: GRAPHENE OXIDE / Support film - topology: HOLEY ARRAY | ||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 297 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 66.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)