[English] 日本語
 Yorodumi
Yorodumi- EMDB-24807: Cryo-EM structure of human NKCC1 K289NA492EL671C bound with bumetanide -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human NKCC1 K289NA492EL671C bound with bumetanide | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cation-chloride cotransporter / ion transport / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of cell volume / positive regulation of aspartate secretion / transepithelial ammonium transport / regulation of matrix metallopeptidase secretion / cell body membrane / metal ion transmembrane transporter activity / inorganic anion import across plasma membrane / inorganic cation import across plasma membrane / chloride:monoatomic cation symporter activity / sodium:potassium:chloride symporter activity ...positive regulation of cell volume / positive regulation of aspartate secretion / transepithelial ammonium transport / regulation of matrix metallopeptidase secretion / cell body membrane / metal ion transmembrane transporter activity / inorganic anion import across plasma membrane / inorganic cation import across plasma membrane / chloride:monoatomic cation symporter activity / sodium:potassium:chloride symporter activity / Cation-coupled Chloride cotransporters / potassium ion transmembrane transporter activity / transepithelial chloride transport / intracellular chloride ion homeostasis / ammonium transmembrane transport / negative regulation of vascular wound healing / sodium ion homeostasis / ammonium channel activity / chloride ion homeostasis / cell projection membrane / cellular response to chemokine / T cell chemotaxis / potassium ion homeostasis / intracellular sodium ion homeostasis / sodium ion import across plasma membrane / cell volume homeostasis / cellular response to potassium ion / hyperosmotic response / regulation of spontaneous synaptic transmission / gamma-aminobutyric acid signaling pathway / maintenance of blood-brain barrier / potassium ion import across plasma membrane / intracellular potassium ion homeostasis / lateral plasma membrane / transport across blood-brain barrier / monoatomic ion transport / sodium ion transmembrane transport / basal plasma membrane / cytoplasmic vesicle membrane / chloride transmembrane transport / cell periphery / cell projection / Hsp90 protein binding / extracellular vesicle / protein-folding chaperone binding / cell body / basolateral plasma membrane / neuron projection / apical plasma membrane / neuronal cell body / protein kinase binding / extracellular exosome / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Zhao YX / Cao EH | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural basis for inhibition of the Cation-chloride cotransporter NKCC1 by the diuretic drug bumetanide Authors: Zhao Y / Roy K / Vidossich P / Cancedda L / De Vivo M / Forbush B / Cao E | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24807.map.gz emd_24807.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24807-v30.xml emd-24807-v30.xml emd-24807.xml emd-24807.xml | 16 KB 16 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_24807.png emd_24807.png | 37 KB | ||
| Filedesc metadata |  emd-24807.cif.gz emd-24807.cif.gz | 6.5 KB | ||
| Others |  emd_24807_additional_1.map.gz emd_24807_additional_1.map.gz | 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24807 http://ftp.pdbj.org/pub/emdb/structures/EMD-24807 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24807 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24807 | HTTPS FTP |
-Validation report
| Summary document |  emd_24807_validation.pdf.gz emd_24807_validation.pdf.gz | 542.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_24807_full_validation.pdf.gz emd_24807_full_validation.pdf.gz | 542.1 KB | Display | |
| Data in XML |  emd_24807_validation.xml.gz emd_24807_validation.xml.gz | 6.3 KB | Display | |
| Data in CIF |  emd_24807_validation.cif.gz emd_24807_validation.cif.gz | 7.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24807 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24807 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24807 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24807 | HTTPS FTP |
-Related structure data
| Related structure data |  7s1xMC  7s1yC  7s1zC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
| EM raw data |  EMPIAR-11049 (Title: Cryo-EM structure of human NKCC1 K289NA492EL671C bound with bumetanide EMPIAR-11049 (Title: Cryo-EM structure of human NKCC1 K289NA492EL671C bound with bumetanideData size: 1.8 TB Data #1: Cryo-EM structure of human NKCC1 K289NA492EL671C bound with bumetanide [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_24807.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24807.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_24807_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human NKCC1 K289NA492EL671C bound with bumetanide
| Entire | Name: human NKCC1 K289NA492EL671C bound with bumetanide |
|---|---|
| Components |
|
-Supramolecule #1: human NKCC1 K289NA492EL671C bound with bumetanide
| Supramolecule | Name: human NKCC1 K289NA492EL671C bound with bumetanide / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 263.34 KDa |
-Macromolecule #1: Solute carrier family 12 member 2
| Macromolecule | Name: Solute carrier family 12 member 2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 131.888641 KDa |
| Recombinant expression | Organism: Mammalian expression vector BsrGI-MCS-pcDNA3.1 (others) |
| Sequence | String: GAMGSEPRPT APSSGAPGLA GVGETPSAAA LAAARVELPG TAVPSVPEDA APASRDGGGV RDEGPAAAGD GLGRPLGPTP SQSRFQVDL VSENAGRAAA AAAAAAAAAA AAGAGAGAKQ TPADGEASGE SEPAKGSEEA KGRFRVNFVD PAASSSAEDS L SDAAGVGV ...String: GAMGSEPRPT APSSGAPGLA GVGETPSAAA LAAARVELPG TAVPSVPEDA APASRDGGGV RDEGPAAAGD GLGRPLGPTP SQSRFQVDL VSENAGRAAA AAAAAAAAAA AAGAGAGAKQ TPADGEASGE SEPAKGSEEA KGRFRVNFVD PAASSSAEDS L SDAAGVGV DGPNVSFQNG GDTVLSEGSS LHSGGGGGSG HHQHYYYDTH TNTYYLRTFG HNTMDAVPRI DHYRHTAAQL GE KLLRPSL AELHDELEKE PFEDGFANGE ESTPTRDAVV TYTAESKGVV KFGWINGVLV RCMLNIWGVM LFIRLSWIVG QAG IGLSVL VIMMATVVTT ITGLSTSAIA TNGFVRGGGA YYLISRSLGP EFGGAIGLIF AFANAVAVAM YVVGFAETVV ELLK EHSIL MIDEINDIRI IGAITVVILL GISVAGMEWE AKAQIVLLVI LLLAIGDFVI GTFIPLESKK PKGFFGYKSE IFNEN FGPD FREEETFFSV FEIFFPAATG ILAGANISGD LADPQSAIPK GTLLAILITT LVYVGIAVSV GSCVVRDATG NVNDTI VTE LTNCTSAACK LNFDFSSCES SPCSYGLMNN FQVMSMVSGF TPLISAGIFS ATLSSALASL VSAPKIFQAL CKDNIYP AF QMFAKGYGKN NEPLRGYILT FLIALGFILI AECNVIAPII SNFFLASYAL INFSVFHASL AKSPGWRPAF KYYNMWIS L LGAILCCIVM FVINWWAALL TYVIVLGLYI YVTYKKPDVN WGSSTQALTY LNALQHSIRL SGVEDHVKNF RPQCLVMTG APNSRPALLH LVHDFTKNVG LMICGHVHMG PRRQAMKEMS IDQAKYQRWL IKNKMKAFYA PVHADDLREG AQYLMQAAGL GRMKPNTLV LGFKKDWLQA DMRDVDMYIN LFHDAFDIQY GVVVIRLKEG LDISHLQGQE ELLSSQEKSP GTKDVVVSVE Y SKKSDLDT SKPLSEKPIT HKVEEEDGKT ATQPLLKKES KGPIVPLNVA DQKLLEASTQ FQKKQGKNTI DVWWLFDDGG LT LLIPYLL TTKKKWKDCK IRVFIGGKIN RIDHDRRAMA TLLSKFRIDF SDIMVLGDIN TKPKKENIIA FEEIIEPYRL HED DKEQDI ADKMKEDEPW RITDNELELY KTKTYRQIRL NELLKEHSST ANIIVMSLPV ARKGAVSSAL YMAWLEALSK DLPP ILLVR GNHQSVLTFY S UniProtKB: Solute carrier family 12 member 2 |
-Macromolecule #2: 3-(butylamino)-4-phenoxy-5-sulfamoylbenzoic acid
| Macromolecule | Name: 3-(butylamino)-4-phenoxy-5-sulfamoylbenzoic acid / type: ligand / ID: 2 / Number of copies: 2 / Formula: 82U |
|---|---|
| Molecular weight | Theoretical: 364.416 Da |
| Chemical component information | 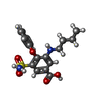 ChemComp-82U: |
-Macromolecule #3: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Macromolecule #4: CHLORIDE ION
| Macromolecule | Name: CHLORIDE ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: CL |
|---|---|
| Molecular weight | Theoretical: 35.453 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 1.175 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)