[English] 日本語
 Yorodumi
Yorodumi- EMDB-24731: afTMEM16 in C18 lipid nanodiscs with MSP1E3 scaffold protein in t... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | afTMEM16 in C18 lipid nanodiscs with MSP1E3 scaffold protein in the presence of Ca2+, monomer with extra lipids | |||||||||
 Map data Map data | unsharpened map from a focused reconstruction on one monomer from symmetry expanded particles, used to build additional lipids. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TMEM16 / lipid scrambling / LIPID TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationphospholipid scramblase activity / cortical endoplasmic reticulum / phospholipid translocation / chloride channel activity / voltage-gated calcium channel activity / chloride transmembrane transport / monoatomic ion transmembrane transport / membrane Similarity search - Function | |||||||||
| Biological species |   | |||||||||
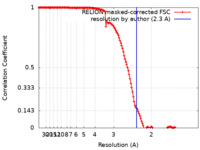
| Method | single particle reconstruction / cryo EM / Resolution: 2.3 Å | |||||||||
 Authors Authors | Falzone ME / Accardi A | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: TMEM16 scramblases thin the membrane to enable lipid scrambling. Authors: Maria E Falzone / Zhang Feng / Omar E Alvarenga / Yangang Pan / ByoungCheol Lee / Xiaolu Cheng / Eva Fortea / Simon Scheuring / Alessio Accardi /   Abstract: TMEM16 scramblases dissipate the plasma membrane lipid asymmetry to activate multiple eukaryotic cellular pathways. Scrambling was proposed to occur with lipid headgroups moving between leaflets ...TMEM16 scramblases dissipate the plasma membrane lipid asymmetry to activate multiple eukaryotic cellular pathways. Scrambling was proposed to occur with lipid headgroups moving between leaflets through a membrane-spanning hydrophilic groove. Direct information on lipid-groove interactions is lacking. We report the 2.3 Å resolution cryogenic electron microscopy structure of the nanodisc-reconstituted Ca-bound afTMEM16 scramblase showing how rearrangement of individual lipids at the open pathway results in pronounced membrane thinning. Only the groove's intracellular vestibule contacts lipids, and mutagenesis suggests scrambling does not require specific protein-lipid interactions with the extracellular vestibule. We find scrambling can occur outside a closed groove in thinner membranes and is inhibited in thicker membranes, despite an open pathway. Our results show afTMEM16 thins the membrane to enable scrambling and that an open hydrophilic pathway is not a structural requirement to allow rapid transbilayer movement of lipids. This mechanism could be extended to other scramblases lacking a hydrophilic groove. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24731.map.gz emd_24731.map.gz | 169.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24731-v30.xml emd-24731-v30.xml emd-24731.xml emd-24731.xml | 15.2 KB 15.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24731_fsc.xml emd_24731_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_24731.png emd_24731.png | 137.9 KB | ||
| Masks |  emd_24731_msk_1.map emd_24731_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-24731.cif.gz emd-24731.cif.gz | 6.2 KB | ||
| Others |  emd_24731_additional_1.map.gz emd_24731_additional_1.map.gz | 202 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24731 http://ftp.pdbj.org/pub/emdb/structures/EMD-24731 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24731 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24731 | HTTPS FTP |
-Related structure data
| Related structure data |  7rxhMC  7rwjC  7rx2C  7rx3C  7rxaC  7rxbC  7rxgC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_24731.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24731.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map from a focused reconstruction on one monomer from symmetry expanded particles, used to build additional lipids. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.7067 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_24731_msk_1.map emd_24731_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: unsharpened map from a focused reconstruction on one...
| File | emd_24731_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map from a focused reconstruction on one monomer from symmetry expanded particles | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : afTMEM16 scramblase C18 lipid nanodiscs with MSP1E3 scaffold in t...
| Entire | Name: afTMEM16 scramblase C18 lipid nanodiscs with MSP1E3 scaffold in the presence of Ca2+, monomer with additional lipids |
|---|---|
| Components |
|
-Supramolecule #1: afTMEM16 scramblase C18 lipid nanodiscs with MSP1E3 scaffold in t...
| Supramolecule | Name: afTMEM16 scramblase C18 lipid nanodiscs with MSP1E3 scaffold in the presence of Ca2+, monomer with additional lipids type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: afTMEM16 lipid scramblase
| Macromolecule | Name: afTMEM16 lipid scramblase / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: CEA10 / CBS 144.89 / FGSC A1163 |
| Molecular weight | Theoretical: 84.616859 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAFNPAPKAV QENHHVDYVI RFNYGDIDTP EAIKKFEVLL LELSEVGLQT EVRQGDENSL FVFVRAASKK KLKRAVYQSR VRDWLYGVR NTEPEPASSA KPQSEAERLL VIYHLITVPK AEGGAGITPR HGEWKNVDAI FPLHDEETNR QCMREWSKKT F LSTEDLDR ...String: MAFNPAPKAV QENHHVDYVI RFNYGDIDTP EAIKKFEVLL LELSEVGLQT EVRQGDENSL FVFVRAASKK KLKRAVYQSR VRDWLYGVR NTEPEPASSA KPQSEAERLL VIYHLITVPK AEGGAGITPR HGEWKNVDAI FPLHDEETNR QCMREWSKKT F LSTEDLDR IRNTFGEHVG FYFAFLQSYF RFLMFPAAFG FSCWLLLGSF SIIYTVVNCL WCIVFIEYWK RQEEDLSCRW QT KGVSAVH EKRAEFKPEK EIRDESTGEV RGVFPATKRM YRQLLQVPFA LLAAVALGAI IATCFAIEIF ISEVYNGPLK GYL VFIPTI LVSALIPTMS AVLLTVATKL NDYENYETQD AYKVALTQKI FVVNFITSYL PIILTAFVYV PFASRIVPYL DVFH LTVRP FVSKEHAIKA RTEFSINPDR LRKQVIYFTV TAQIVGFALE TIVPFVKQRV FREYKEYTKK QHAKAEPGNG AGEKK TVSL GDDEDEARFL TRVRNEAELE DYDVTDDLRE MCIQFGYLAL FSPVWPLVPV SFLINNWVEL RSDFFKICVE CKRPWP QRA DTIGPWLDSL GFLSWVGSIT SSALVYMFSN GHEGPNGEPT TIRCWALLLT IFFSEHLYLI VRYAVRSALA KLEPPNT RR ERIERFMMRK RYLDTVLSAE SDDDADEVKG VVSSIPPSEI TRESLEQDAR DWSKQGTDPT ERFWMRQRGW KESAEVGL S LITKAKGDET KKQQ UniProtKB: Plasma membrane channel protein (Aqy1), putative |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #3: (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]o...
| Macromolecule | Name: (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate type: ligand / ID: 3 / Number of copies: 16 / Formula: PGW |
|---|---|
| Molecular weight | Theoretical: 749.007 Da |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 24 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 42.8227 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)