[English] 日本語
 Yorodumi
Yorodumi- EMDB-24463: Cryo-EM structure of human rod CNGA1/B1 channel in L-cis-Diltiaze... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24463 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human rod CNGA1/B1 channel in L-cis-Diltiazem-blocked open state | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | ion channel / TRANSPORT PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationolfactory nerve maturation / non-motile cilium membrane / detection of chemical stimulus involved in sensory perception of smell / photoreceptor cell outer segment organization / response to odorant / intracellular cyclic nucleotide activated cation channel complex / protein localization to organelle / intracellularly cGMP-activated cation channel activity / : / detection of light stimulus involved in visual perception ...olfactory nerve maturation / non-motile cilium membrane / detection of chemical stimulus involved in sensory perception of smell / photoreceptor cell outer segment organization / response to odorant / intracellular cyclic nucleotide activated cation channel complex / protein localization to organelle / intracellularly cGMP-activated cation channel activity / : / detection of light stimulus involved in visual perception / intracellularly cAMP-activated cation channel activity / rod photoreceptor outer segment / VxPx cargo-targeting to cilium / Golgi-associated vesicle membrane / transmembrane transporter complex / photoreceptor cell maintenance / retina homeostasis / ciliary membrane / photoreceptor outer segment membrane / ligand-gated monoatomic ion channel activity / sodium ion transport / sodium channel activity / monoatomic cation transmembrane transport / cGMP binding / monoatomic cation transport / membrane depolarization / photoreceptor outer segment / phototransduction / regulation of cytosolic calcium ion concentration / cAMP binding / visual perception / potassium ion transport / terminal bouton / calcium channel activity / Olfactory Signaling Pathway / Activation of the phototransduction cascade / calcium ion transport / sensory perception of smell / Inactivation, recovery and regulation of the phototransduction cascade / positive regulation of gene expression / protein-containing complex binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.88 Å | ||||||||||||
 Authors Authors | Xue J / Han Y | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Neuron / Year: 2022 Journal: Neuron / Year: 2022Title: Structural mechanisms of assembly, permeation, gating, and pharmacology of native human rod CNG channel. Authors: Jing Xue / Yan Han / Weizhong Zeng / Youxing Jiang /  Abstract: Mammalian cyclic nucleotide-gated (CNG) channels are nonselective cation channels activated by cGMP or cAMP and play essential roles in the signal transduction of the visual and olfactory sensory ...Mammalian cyclic nucleotide-gated (CNG) channels are nonselective cation channels activated by cGMP or cAMP and play essential roles in the signal transduction of the visual and olfactory sensory systems. CNGA1, the principal component of the CNG channel from rod photoreceptors, can by itself form a functional homotetrameric channel and has been used as the model system in the majority of rod CNG studies. However, the native rod CNG functions as a heterotetramer consisting of three A1 and one B1 subunits and exhibits different functional properties than the CNGA1 homomer. Here we present the functional analysis of human rod CNGA1/B1 heterotetramer and its cryo-EM structures in apo, cGMP-bound, cAMP-bound, and L-cis-Diltiazem-blocked states. These structures, with resolution ranging from 2.6 to 3.3 Å, elucidate the structural mechanisms underlying the 3:1 subunit stoichiometry, the asymmetrical gating upon cGMP activation, and the unique pharmacological property of the native rod CNG channel. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24463.map.gz emd_24463.map.gz | 85.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24463-v30.xml emd-24463-v30.xml emd-24463.xml emd-24463.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_24463.png emd_24463.png | 194.7 KB | ||
| Filedesc metadata |  emd-24463.cif.gz emd-24463.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24463 http://ftp.pdbj.org/pub/emdb/structures/EMD-24463 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24463 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24463 | HTTPS FTP |
-Related structure data
| Related structure data |  7rhjMC  7rh9C  7rhgC  7rhhC  7rhiC  7rhkC  7rhlC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24463.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24463.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.839 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
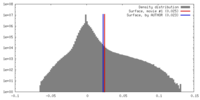
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human rod CNGA1/B1 channel in L-cis-Diltiazem-blocked open state
| Entire | Name: human rod CNGA1/B1 channel in L-cis-Diltiazem-blocked open state |
|---|---|
| Components |
|
-Supramolecule #1: human rod CNGA1/B1 channel in L-cis-Diltiazem-blocked open state
| Supramolecule | Name: human rod CNGA1/B1 channel in L-cis-Diltiazem-blocked open state type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: cGMP-gated cation channel alpha-1
| Macromolecule | Name: cGMP-gated cation channel alpha-1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 64.650434 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDYKDDDDKG GSASKDKKEE EKKEVVVIDP SGNTYYNWLF CITLPVMYNW TMVIARACFD ELQSDYLEYW LILDYVSDIV YLIDMFVRT RTGYLEQGLL VKEELKLINK YKSNLQFKLD VLSLIPTDLL YFKLGWNYPE IRLNRLLRFS RMFEFFQRTE T RTNYPNIF ...String: MDYKDDDDKG GSASKDKKEE EKKEVVVIDP SGNTYYNWLF CITLPVMYNW TMVIARACFD ELQSDYLEYW LILDYVSDIV YLIDMFVRT RTGYLEQGLL VKEELKLINK YKSNLQFKLD VLSLIPTDLL YFKLGWNYPE IRLNRLLRFS RMFEFFQRTE T RTNYPNIF RISNLVMYIV IIIHWNACVF YSISKAIGFG NDTWVYPDIN DPEFGRLARK YVYSLYWSTL TLTTIGETPP PV RDSEYVF VVVDFLIGVL IFATIVGNIG SMISNMNAAR AEFQARIDAI KQYMHFRNVS KDMEKRVIKW FDYLWTNKKT VDE KEVLKY LPDKLRAEIA INVHLDTLKK VRIFADCEAG LLVELVLKLQ PQVYSPGDYI CKKGDIGREM YIIKEGKLAV VADD GVTQF VVLSDGSYFG EISILNIKGS KAGNRRTANI KSIGYSDLFC LSKDDLMEAL TEYPDAKTML EEKGKQILMK DGLLD LNIA NAGSDPKDLE EKVTRMEGSV DLLQTRFARI LAEYESMQQK LKQRLTKVEK FLKPLIDTEF SSIEGPGAES GPIDST UniProtKB: Cyclic nucleotide-gated channel alpha-1 |
-Macromolecule #2: Cyclic nucleotide-gated cation channel beta-1
| Macromolecule | Name: Cyclic nucleotide-gated cation channel beta-1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 91.039305 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDYKDDDDKG GSASSGVPAT KQHPEVQVED TDADSCPLMA EENPPSTVLP PPSPAKSDTL IVPSSASGTH RKKLPSEDDE AEELKALSP AESPVVAWSD PTTPKDTDGQ DRAASTASTN SAIINDRLQE LVKLFKERTE KVKEKLIDPD VTSDEESPKP S PAKKAPEP ...String: MDYKDDDDKG GSASSGVPAT KQHPEVQVED TDADSCPLMA EENPPSTVLP PPSPAKSDTL IVPSSASGTH RKKLPSEDDE AEELKALSP AESPVVAWSD PTTPKDTDGQ DRAASTASTN SAIINDRLQE LVKLFKERTE KVKEKLIDPD VTSDEESPKP S PAKKAPEP APDTKPAEAE PVEEEHYCDM LCCKFKHRPW KKYQFPQSID PLTNLMYVLW LFFVVMAWNW NCWLIPVRWA FP YQTPDNI HHWLLMDYLC DLIYFLDITV FQTRLQFVRG GDIITDKKDM RNNYLKSRRF KMDLLSLLPL DFLYLKVGVN PLL RLPRCL KYMAFFEFNS RLESILSKAY VYRVIRTTAY LLYSLHLNSC LYYWASAYQG LGSTHWVYDG VGNSYIRCYY FAVK TLITI GGLPDPKTLF EIVFQLLNYF TGVFAFSVMI GQMRDVVGAA TAGQTYYRSC MDSTVKYMNF YKIPKSVQNR VKTWY EYTW HSQGMLDESE LMVQLPDKMR LDLAIDVNYN IVSKVALFQG CDRQMIFDML KRLRSVVYLP NDYVCKKGEI GREMYI IQA GQVQVLGGPD GKSVLVTLKA GSVFGEISLL AVGGGNRRTA NVVAHGFTNL FILDKKDLNE ILVHYPESQK LLRKKAR RM LRSNNKPKEE KSVLILPPRA GTPKLFNAAL AMTGKMGGKG AKGGKLAHLR ARLKELAALE AAAKQQELVE QAKSSQDV K GEEGSAAPDQ HTHPKEAATD PPAPRTPPEP PGSPPSSPPP ASLGRPEGEE EGPAEPEEHS VRICMSPGPE PGEQILSVK MPEEREEKAE UniProtKB: Cyclic nucleotide-gated channel beta-1 |
-Macromolecule #3: CYCLIC GUANOSINE MONOPHOSPHATE
| Macromolecule | Name: CYCLIC GUANOSINE MONOPHOSPHATE / type: ligand / ID: 3 / Number of copies: 4 / Formula: PCG |
|---|---|
| Molecular weight | Theoretical: 345.205 Da |
| Chemical component information |  ChemComp-PCG: |
-Macromolecule #4: (2R,3R)-5-[2-(dimethylamino)ethyl]-2-(4-methoxyphenyl)-4-oxo-2,3,...
| Macromolecule | Name: (2R,3R)-5-[2-(dimethylamino)ethyl]-2-(4-methoxyphenyl)-4-oxo-2,3,4,5-tetrahydro-1,5-benzothiazepin-3-yl acetate type: ligand / ID: 4 / Number of copies: 1 / Formula: 5H0 |
|---|---|
| Molecular weight | Theoretical: 414.518 Da |
| Chemical component information |  ChemComp-5H0: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)