[English] 日本語
 Yorodumi
Yorodumi- EMDB-24432: SARS-CoV-2 replication-transcription complex bound to nsp13 helic... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24432 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC dimer | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA-dependent RNA polymerase / viral replication-transcription complex / transcription / viral proteins / REPLICATION-TRANSCRIPTION complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein guanylyltransferase activity / RNA endonuclease activity producing 3'-phosphomonoesters, hydrolytic mechanism / mRNA guanylyltransferase activity / 5'-3' RNA helicase activity / Lyases; Phosphorus-oxygen lyases / Assembly of the SARS-CoV-2 Replication-Transcription Complex (RTC) / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of TBK1 activity / Maturation of replicase proteins / ISG15-specific peptidase activity / TRAF3-dependent IRF activation pathway ...protein guanylyltransferase activity / RNA endonuclease activity producing 3'-phosphomonoesters, hydrolytic mechanism / mRNA guanylyltransferase activity / 5'-3' RNA helicase activity / Lyases; Phosphorus-oxygen lyases / Assembly of the SARS-CoV-2 Replication-Transcription Complex (RTC) / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of TBK1 activity / Maturation of replicase proteins / ISG15-specific peptidase activity / TRAF3-dependent IRF activation pathway / Transcription of SARS-CoV-2 sgRNAs / Translation of Replicase and Assembly of the Replication Transcription Complex / snRNP Assembly / Replication of the SARS-CoV-2 genome / Hydrolases; Acting on ester bonds; Exoribonucleases producing 5'-phosphomonoesters / host cell endoplasmic reticulum-Golgi intermediate compartment / double membrane vesicle viral factory outer membrane / SARS coronavirus main proteinase / 5'-3' DNA helicase activity / 3'-5'-RNA exonuclease activity / host cell endosome / symbiont-mediated degradation of host mRNA / mRNA guanylyltransferase / symbiont-mediated suppression of host ISG15-protein conjugation / G-quadruplex RNA binding / symbiont-mediated suppression of host toll-like receptor signaling pathway / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of IRF3 activity / omega peptidase activity / mRNA (guanine-N7)-methyltransferase / SARS-CoV-2 modulates host translation machinery / methyltransferase cap1 / host cell Golgi apparatus / symbiont-mediated suppression of host NF-kappaB cascade / symbiont-mediated perturbation of host ubiquitin-like protein modification / DNA helicase / methyltransferase cap1 activity / ubiquitinyl hydrolase 1 / cysteine-type deubiquitinase activity / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / Hydrolases; Acting on peptide bonds (peptidases); Cysteine endopeptidases / single-stranded RNA binding / host cell perinuclear region of cytoplasm / regulation of autophagy / viral protein processing / lyase activity / host cell endoplasmic reticulum membrane / RNA helicase / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / symbiont-mediated suppression of host gene expression / copper ion binding / viral translational frameshifting / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / viral RNA genome replication / RNA-directed RNA polymerase activity / lipid binding / DNA-templated transcription / host cell nucleus / SARS-CoV-2 activates/modulates innate and adaptive immune responses / ATP hydrolysis activity / proteolysis / RNA binding / zinc ion binding / ATP binding / membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
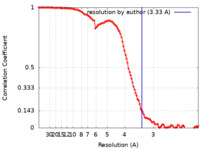
| Method | single particle reconstruction / cryo EM / Resolution: 3.33 Å | |||||||||
 Authors Authors | Chen J / Malone B | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: Ensemble cryo-EM reveals conformational states of the nsp13 helicase in the SARS-CoV-2 helicase replication-transcription complex. Authors: James Chen / Qi Wang / Brandon Malone / Eliza Llewellyn / Yakov Pechersky / Kashyap Maruthi / Ed T Eng / Jason K Perry / Elizabeth A Campbell / David E Shaw / Seth A Darst /  Abstract: The SARS-CoV-2 nonstructural proteins coordinate genome replication and gene expression. Structural analyses revealed the basis for coupling of the essential nsp13 helicase with the RNA-dependent RNA ...The SARS-CoV-2 nonstructural proteins coordinate genome replication and gene expression. Structural analyses revealed the basis for coupling of the essential nsp13 helicase with the RNA-dependent RNA polymerase (RdRp) where the holo-RdRp and RNA substrate (the replication-transcription complex or RTC) associated with two copies of nsp13 (nsp13-RTC). One copy of nsp13 interacts with the template-RNA in an opposing polarity to the RdRp and is envisaged to drive the RdRp backward on the RNA template (backtracking), prompting questions as to how the RdRp can efficiently synthesize RNA in the presence of nsp13. Here we use cryogenic-electron microscopy and molecular dynamics simulations to analyze the nsp13-RTC, revealing four distinct conformational states of the helicases. The results indicate a mechanism for the nsp13-RTC to turn backtracking on and off, using an allosteric mechanism to switch between RNA synthesis or backtracking in response to stimuli at the RdRp active site. #1:  Journal: Biorxiv / Year: 2021 Journal: Biorxiv / Year: 2021Title: Ensemble cryo-electron microscopy reveals conformational states of the nsp13 helicase in the SARS-CoV-2 helicase replication-transcription complex Authors: Chen J / Wang Q / Malone B / Llewellyn E / Pechersky Y / Maruthi K / Eng ET / Perry JK / Campbell EA / Shaw DE / Darst SA | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24432.map.gz emd_24432.map.gz | 64.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24432-v30.xml emd-24432-v30.xml emd-24432.xml emd-24432.xml | 25.1 KB 25.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24432_fsc.xml emd_24432_fsc.xml | 11.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_24432.png emd_24432.png | 101.9 KB | ||
| Filedesc metadata |  emd-24432.cif.gz emd-24432.cif.gz | 7.9 KB | ||
| Others |  emd_24432_additional_1.map.gz emd_24432_additional_1.map.gz | 71.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24432 http://ftp.pdbj.org/pub/emdb/structures/EMD-24432 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24432 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24432 | HTTPS FTP |
-Validation report
| Summary document |  emd_24432_validation.pdf.gz emd_24432_validation.pdf.gz | 596 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_24432_full_validation.pdf.gz emd_24432_full_validation.pdf.gz | 595.6 KB | Display | |
| Data in XML |  emd_24432_validation.xml.gz emd_24432_validation.xml.gz | 12.5 KB | Display | |
| Data in CIF |  emd_24432_validation.cif.gz emd_24432_validation.cif.gz | 16.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24432 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24432 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24432 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24432 | HTTPS FTP |
-Related structure data
| Related structure data |  7re3MC  7rdxC  7rdyC  7rdzC  7re0C  7re1C  7re2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24432.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24432.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
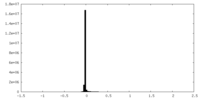
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: local resolution filtered map
| File | emd_24432_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | local resolution filtered map | ||||||||||||
| Projections & Slices |
| ||||||||||||
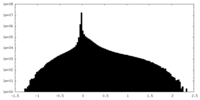
| Density Histograms |
- Sample components
Sample components
+Entire : SARS-CoV-2 replication/transcription complex bound to nsp13 helic...
+Supramolecule #1: SARS-CoV-2 replication/transcription complex bound to nsp13 helic...
+Macromolecule #1: RNA-directed RNA polymerase
+Macromolecule #2: Non-structural protein 8
+Macromolecule #3: Non-structural protein 7
+Macromolecule #4: Helicase
+Macromolecule #5: Product RNA
+Macromolecule #6: Template RNA
+Macromolecule #7: ZINC ION
+Macromolecule #8: MAGNESIUM ION
+Macromolecule #9: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #10: ALUMINUM FLUORIDE
+Macromolecule #11: CHAPSO
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 65.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)