+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23960 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Norovirus T=3 GII.4 HOV VLP | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | T=3 Capsid GII.4 HOV Norovirus / VIRUS | |||||||||
| Function / homology | Calicivirus coat protein C-terminal / Calicivirus coat protein C-terminal / Calicivirus coat protein / Calicivirus coat protein / virion component / Picornavirus/Calicivirus coat protein / Viral coat protein subunit / host cell cytoplasm / VP1 Function and homology information Function and homology information | |||||||||
| Biological species |  Norovirus GII.4 Norovirus GII.4 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Salmen W / Hu L | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Atomic structure of the predominant GII.4 human norovirus capsid reveals novel stability and plasticity. Authors: Liya Hu / Wilhelm Salmen / Rong Chen / Yi Zhou / Frederick Neill / James E Crowe / Robert L Atmar / Mary K Estes / B V Venkataram Prasad /  Abstract: Human noroviruses (HuNoVs) cause sporadic and epidemic viral gastroenteritis worldwide. The GII.4 variants are responsible for most HuNoV infections, and GII.4 virus-like particles (VLPs) are being ...Human noroviruses (HuNoVs) cause sporadic and epidemic viral gastroenteritis worldwide. The GII.4 variants are responsible for most HuNoV infections, and GII.4 virus-like particles (VLPs) are being used in vaccine development. The atomic structure of the GII.4 capsid in the native T = 3 state has not been determined. Here we present the GII.4 VLP structure with T = 3 symmetry determined using X-ray crystallography and cryo-EM at 3.0 Å and 3.8 Å resolution, respectively, which reveals unanticipated novel features. A novel aspect in the crystal structure determined without imposing icosahedral symmetry is the remarkable adaptability of the capsid protein VP1 driven by the flexible hinge between the shell and the protruding domains. In both crystal and cryo-EM structures, VP1 adopts a stable conformation with the protruding domain resting on the shell domain, in contrast to the 'rising' conformation observed in recent cryo-EM structures of other GII.4 VLPs. Our studies further revealed that the resting state of VP1 dimer is stabilized by a divalent ion, and chelation using EDTA increases capsid diameter, exposing new hydrophobic and antigenic sites and suggesting a transition to the rising conformation. These novel insights into GII.4 capsid structure, stability, and antigen presentation may be useful for ongoing vaccine development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23960.map.gz emd_23960.map.gz | 451.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23960-v30.xml emd-23960-v30.xml emd-23960.xml emd-23960.xml | 14.9 KB 14.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23960.png emd_23960.png | 224.3 KB | ||
| Masks |  emd_23960_msk_1.map emd_23960_msk_1.map | 476.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-23960.cif.gz emd-23960.cif.gz | 5.5 KB | ||
| Others |  emd_23960_half_map_1.map.gz emd_23960_half_map_1.map.gz emd_23960_half_map_2.map.gz emd_23960_half_map_2.map.gz | 442.1 MB 442.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23960 http://ftp.pdbj.org/pub/emdb/structures/EMD-23960 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23960 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23960 | HTTPS FTP |
-Validation report
| Summary document |  emd_23960_validation.pdf.gz emd_23960_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23960_full_validation.pdf.gz emd_23960_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_23960_validation.xml.gz emd_23960_validation.xml.gz | 18.2 KB | Display | |
| Data in CIF |  emd_23960_validation.cif.gz emd_23960_validation.cif.gz | 21.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23960 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23960 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23960 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23960 | HTTPS FTP |
-Related structure data
| Related structure data |  7mryMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23960.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23960.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.24 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
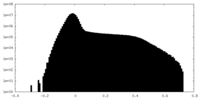
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_23960_msk_1.map emd_23960_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
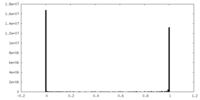
| Density Histograms |
-Half map: #1
| File | emd_23960_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_23960_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Norovirus GII.4
| Entire | Name:  Norovirus GII.4 Norovirus GII.4 |
|---|---|
| Components |
|
-Supramolecule #1: Norovirus GII.4
| Supramolecule | Name: Norovirus GII.4 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1 / NCBI-ID: 489821 / Sci species name: Norovirus GII.4 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: Yes |
|---|
-Macromolecule #1: VP1
| Macromolecule | Name: VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Norovirus GII.4 Norovirus GII.4 |
| Molecular weight | Theoretical: 58.760895 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MKMASSDASP SDGSTANLVP EVNNEVMALE PVVGAAIAAP VAGQQNVIDP WIRNNFVQAP GGEFTVSPRN APGEILWSAP LGPDLNPYL SHLARMYNGY AGGFEVQVIL AGNAFTAGKI IFAAVPPNFP TEGLSPSQVT MFPHIIVDVR QLEPVLIPLP D VRNNFYHY ...String: MKMASSDASP SDGSTANLVP EVNNEVMALE PVVGAAIAAP VAGQQNVIDP WIRNNFVQAP GGEFTVSPRN APGEILWSAP LGPDLNPYL SHLARMYNGY AGGFEVQVIL AGNAFTAGKI IFAAVPPNFP TEGLSPSQVT MFPHIIVDVR QLEPVLIPLP D VRNNFYHY NQSNDPTIKL IAMLYTPLRA NNAGDDVFTV SCRVLTRPSP DFDFIFLVPP TVESRTKPFT VPILTVEEMT NS RFPIPLE KLFTGPSGAF VVQPQNGRCT TDGVLLGTTQ LSPVNICTFR GDVTHIAGTH DYTMNLASQN WNNYDPTEEI PAP LGTPDF VGKIQGVLTQ TTRGDGSTRG HKATVSTGSV HFTPKLGSVQ FTTDTNNDLE TGQNTKFTPV GVVQDGNSAH QNEP QQWVL PNYSGRTGHN VHLAPAVAPT FPGEQLLFFR STMPGCSGYP NMNLDCLLPQ EWVLHFYQEA APAQSDVALL RFVNP DTGR VLFECKLHKS GYVTVAHTGP HDLVIPPNGY FRFDSWVNQF YTLAPMGNGA GRRRAL UniProtKB: VP1 |
-Macromolecule #2: water
| Macromolecule | Name: water / type: ligand / ID: 2 / Number of copies: 5 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.96 mg/mL |
|---|---|
| Buffer | pH: 6 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average exposure time: 10.0 sec. / Average electron dose: 55.96 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 2.15.0) Details: Using Cryosparc GSFSC, resolution calculated: No Mask = 4.2 A, Spherical = 4 A, Loose = 3.9 A, Tight = 3.8 A, corrected = 3.8 A Number images used: 51913 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2.15.0) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2.15.0) |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)