+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23520 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of RSV preF bound by Fabs 32.4K and 01.4B | |||||||||
 Map data Map data | Full map, sharpened with DeepEMhancer | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RSV / viral fusogen / fusion protein / antibody / IMMUNE SYSTEM / IMMUNE SYSTEM-VIRAL PROTEIN complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated induction of syncytium formation / host cell Golgi membrane / entry receptor-mediated virion attachment to host cell / fusion of virus membrane with host plasma membrane / viral envelope / symbiont entry into host cell / host cell plasma membrane / virion membrane Similarity search - Function | |||||||||
| Biological species |  Respiratory syncytial virus / Respiratory syncytial virus /  Homo sapiens (human) Homo sapiens (human) | |||||||||
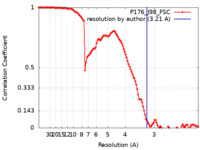
| Method | single particle reconstruction / cryo EM / Resolution: 3.21 Å | |||||||||
 Authors Authors | Wrapp D / McLellan JS | |||||||||
 Citation Citation |  Journal: Immunity / Year: 2021 Journal: Immunity / Year: 2021Title: Vaccination with prefusion-stabilized respiratory syncytial virus fusion protein induces genetically and antigenically diverse antibody responses. Authors: Maryam Mukhamedova / Daniel Wrapp / Chen-Hsiang Shen / Morgan S A Gilman / Tracy J Ruckwardt / Chaim A Schramm / Larissa Ault / Lauren Chang / Alexandrine Derrien-Colemyn / Sarah A M Lucas / ...Authors: Maryam Mukhamedova / Daniel Wrapp / Chen-Hsiang Shen / Morgan S A Gilman / Tracy J Ruckwardt / Chaim A Schramm / Larissa Ault / Lauren Chang / Alexandrine Derrien-Colemyn / Sarah A M Lucas / Amy Ransier / Samuel Darko / Emily Phung / Lingshu Wang / Yi Zhang / Scott A Rush / Bharat Madan / Guillaume B E Stewart-Jones / Pamela J Costner / LaSonji A Holman / Somia P Hickman / Nina M Berkowitz / Nicole A Doria-Rose / Kaitlyn M Morabito / Brandon J DeKosky / Martin R Gaudinski / Grace L Chen / Michelle C Crank / John Misasi / Nancy J Sullivan / Daniel C Douek / Peter D Kwong / Barney S Graham / Jason S McLellan / John R Mascola /  Abstract: An effective vaccine for respiratory syncytial virus (RSV) is an unrealized public health goal. A single dose of the prefusion-stabilized fusion (F) glycoprotein subunit vaccine (DS-Cav1) ...An effective vaccine for respiratory syncytial virus (RSV) is an unrealized public health goal. A single dose of the prefusion-stabilized fusion (F) glycoprotein subunit vaccine (DS-Cav1) substantially increases serum-neutralizing activity in healthy adults. We sought to determine whether DS-Cav1 vaccination induces a repertoire mirroring the pre-existing diversity from natural infection or whether antibody lineages targeting specific epitopes predominate. We evaluated RSV F-specific B cell responses before and after vaccination in six participants using complementary B cell sequencing methodologies and identified 555 clonal lineages. DS-Cav1-induced lineages recognized the prefusion conformation of F (pre-F) and were genetically diverse. Expressed antibodies recognized all six antigenic sites on the pre-F trimer. We identified 34 public clonotypes, and structural analysis of two antibodies from a predominant clonotype revealed a common mode of recognition. Thus, vaccination with DS-Cav1 generates a diverse polyclonal response targeting the antigenic sites on pre-F, supporting the development and advanced testing of pre-F-based vaccines against RSV. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23520.map.gz emd_23520.map.gz | 124.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23520-v30.xml emd-23520-v30.xml emd-23520.xml emd-23520.xml | 23.1 KB 23.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23520_fsc.xml emd_23520_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_23520.png emd_23520.png | 155.1 KB | ||
| Filedesc metadata |  emd-23520.cif.gz emd-23520.cif.gz | 6.5 KB | ||
| Others |  emd_23520_additional_1.map.gz emd_23520_additional_1.map.gz emd_23520_half_map_1.map.gz emd_23520_half_map_1.map.gz emd_23520_half_map_2.map.gz emd_23520_half_map_2.map.gz | 136.6 MB 134.2 MB 134.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23520 http://ftp.pdbj.org/pub/emdb/structures/EMD-23520 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23520 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23520 | HTTPS FTP |
-Related structure data
| Related structure data |  7lucMC  7ludC  7lueC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23520.map.gz / Format: CCP4 / Size: 144.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23520.map.gz / Format: CCP4 / Size: 144.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map, sharpened with DeepEMhancer | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.073 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
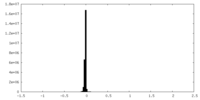
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Focused map (C1)
| File | emd_23520_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused map (C1) | ||||||||||||
| Projections & Slices |
| ||||||||||||
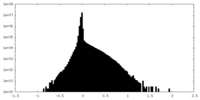
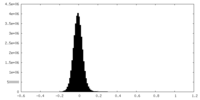
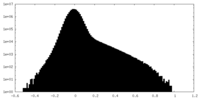
| Density Histograms |
-Half map: Half map B
| File | emd_23520_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
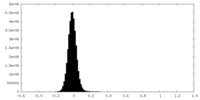
| Density Histograms |
-Half map: Half map A
| File | emd_23520_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
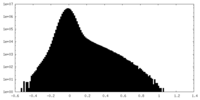
| Density Histograms |
- Sample components
Sample components
-Entire : Prefusion RSV F bound by Fabs 01.4B and 32.4K
| Entire | Name: Prefusion RSV F bound by Fabs 01.4B and 32.4K |
|---|---|
| Components |
|
-Supramolecule #1: Prefusion RSV F bound by Fabs 01.4B and 32.4K
| Supramolecule | Name: Prefusion RSV F bound by Fabs 01.4B and 32.4K / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Respiratory syncytial virus Respiratory syncytial virus |
| Molecular weight | Theoretical: 440 KDa |
-Macromolecule #1: Fusion glycoprotein F0
| Macromolecule | Name: Fusion glycoprotein F0 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Respiratory syncytial virus Respiratory syncytial virus |
| Molecular weight | Theoretical: 58.615941 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QNITEEFYQS TCSAVSKGYL SALRTGWYTS VITIELSNIK EIKCNGTDAK VKLIKQELDK YKNAVTELQL LMQSTPATNN RARRELPRF MNYTLNNAKK TNVTLSKKRK RRFLGFLLGV GSAIASGVAV SKVLHLEGEV NKIKSALLST NKAVVSLSNG V SVLTSKVL ...String: QNITEEFYQS TCSAVSKGYL SALRTGWYTS VITIELSNIK EIKCNGTDAK VKLIKQELDK YKNAVTELQL LMQSTPATNN RARRELPRF MNYTLNNAKK TNVTLSKKRK RRFLGFLLGV GSAIASGVAV SKVLHLEGEV NKIKSALLST NKAVVSLSNG V SVLTSKVL DLKNYIDKQL LPIVNKQSCS IPNIETVIEF QQKNNRLLEI TREFSVNAGV TTPVSTYMLT NSELLSLIND MP ITNDQKK LMSNNVQIVR QQSYSIMSII KEEVLAYVVQ LPLYGVIDTP CWKLHTSPLC TTNTKEGSNI CLTRTDRGWY CDN AGSVSF FPQAETCKVQ SNRVFCDTMN SLTLPSEVNL CNVDIFNPKY DCKIMTSKTD VSSSVITSLG AIVSCYGKTK CTAS NKNRG IIKTFSNGCD YVSNKGVDTV SVGNTLYYVN KQEGKSLYVK GEPIINFYDP LVFPSDEFDA SISQVNEKIN QSLAF IRKS DELLSAIGGY IPEAPRDGQA YVRKDGEWVL LSTFLGSLEV LFQ UniProtKB: Fusion glycoprotein F0 |
-Macromolecule #2: 01.4B Fab Heavy chain
| Macromolecule | Name: 01.4B Fab Heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.154789 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLVQSGAE VKKPGASVKL SCQASGYTFN NYGVSWLRQA PGQGLEWMGW ISAYNGNKKY APKFQGRLTL TTVTSTGTAY MELRSLKSD DTALYFCARD PPAVAAAMFD FWGQGTQVTV SS |
-Macromolecule #3: 01.4B Fab Light chain
| Macromolecule | Name: 01.4B Fab Light chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.499002 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DVVLTQSPLS LPVTLGQPAS ISCRSSQSLV LSDGNTYLSW FHQRPGHSPR RLIYRISHRD SGVPDRFSGS ESGTDFTLKI SRVEAEDVG IYYCMQGTHW PRTFGQGTKV EIK |
-Macromolecule #4: 32.4K Fab Heavy chain
| Macromolecule | Name: 32.4K Fab Heavy chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.140781 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLVQSGAE VKKPGESLKI SCKGSADSFT NHWIGWVRQT PGKGLEWMGM IYPGDSDTRY SPSFQGQVTL SVDKSVTTVY LQWNSLKAS DTAIYYCARQ VGGVVVAEPP PYYYYGMDAW GQGTTVTVSS |
-Macromolecule #5: 32.4K Fab Light chain
| Macromolecule | Name: 32.4K Fab Light chain / type: protein_or_peptide / ID: 5 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.664103 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQLTQSPSS LSASVGDRVT LTCRASQSIA TFLNWFQQRP GKAPKLLMFD ASKLQTGVPS RFSGSGSGTH FTLTISTLQP EDFATYYCQ QSYDLPLTFG PGTKVEIK |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 30 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 37.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)