+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22402 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human DPP9-CARD8 complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CARD8 / DPP9 / inflammasome / Val-boroPro (VbP) / talabostat / innate immunity / IMMUNE SYSTEM / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationCARD8 inflammasome complex assembly / CARD8 inflammasome complex / NACHT domain binding / Formation of apoptosome / CARD domain binding / self proteolysis / NLRP3 inflammasome complex / dipeptidyl-peptidase IV / negative regulation of lipopolysaccharide-mediated signaling pathway / negative regulation of NLRP3 inflammasome complex assembly ...CARD8 inflammasome complex assembly / CARD8 inflammasome complex / NACHT domain binding / Formation of apoptosome / CARD domain binding / self proteolysis / NLRP3 inflammasome complex / dipeptidyl-peptidase IV / negative regulation of lipopolysaccharide-mediated signaling pathway / negative regulation of NLRP3 inflammasome complex assembly / cysteine-type endopeptidase activator activity / dipeptidyl-peptidase activity / Regulation of the apoptosome activity / Hydrolases; Acting on peptide bonds (peptidases) / negative regulation of interleukin-1 beta production / negative regulation of programmed cell death / pattern recognition receptor activity / cell leading edge / pyroptotic inflammatory response / aminopeptidase activity / negative regulation of tumor necrosis factor-mediated signaling pathway / activation of innate immune response / intrinsic apoptotic signaling pathway / serine-type peptidase activity / antiviral innate immune response / positive regulation of interleukin-1 beta production / negative regulation of canonical NF-kappaB signal transduction / molecular condensate scaffold activity / protein destabilization / positive regulation of inflammatory response / peptidase activity / regulation of apoptotic process / defense response to virus / microtubule / inflammatory response / protein homodimerization activity / protein-containing complex / proteolysis / nucleoplasm / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Sharif H / Hollingsworth LR | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Immunity / Year: 2021 Journal: Immunity / Year: 2021Title: Dipeptidyl peptidase 9 sets a threshold for CARD8 inflammasome formation by sequestering its active C-terminal fragment. Authors: Humayun Sharif / L Robert Hollingsworth / Andrew R Griswold / Jeffrey C Hsiao / Qinghui Wang / Daniel A Bachovchin / Hao Wu /  Abstract: CARD8 detects intracellular danger signals and forms a caspase-1 activating inflammasome. Like the related inflammasome sensor NLRP1, CARD8 autoprocesses into noncovalently associated N-terminal (NT) ...CARD8 detects intracellular danger signals and forms a caspase-1 activating inflammasome. Like the related inflammasome sensor NLRP1, CARD8 autoprocesses into noncovalently associated N-terminal (NT) and C-terminal (CT) fragments and binds the cellular dipeptidyl peptidases DPP8 and 9 (DPP8/9). Certain danger-associated signals, including the DPP8/9 inhibitor Val-boroPro (VbP) and HIV protease, induce proteasome-mediated NT degradation and thereby liberate the inflammasome-forming CT. Here, we report cryoelectron microscopy (cryo-EM) structures of CARD8 bound to DPP9, revealing a repressive ternary complex consisting of DPP9, full-length CARD8, and CARD8-CT. Unlike NLRP1-CT, CARD8-CT does not interact with the DPP8/9 active site and is not directly displaced by VbP. However, larger DPP8/9 active-site probes can directly weaken this complex in vitro, and VbP itself nevertheless appears to disrupt this complex, perhaps indirectly, in cells. Thus, DPP8/9 inhibitors can activate the CARD8 inflammasome by promoting CARD8 NT degradation and by weakening ternary complex stability. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22402.map.gz emd_22402.map.gz | 15.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22402-v30.xml emd-22402-v30.xml emd-22402.xml emd-22402.xml | 15.8 KB 15.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22402_fsc.xml emd_22402_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_22402.png emd_22402.png | 167.4 KB | ||
| Filedesc metadata |  emd-22402.cif.gz emd-22402.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22402 http://ftp.pdbj.org/pub/emdb/structures/EMD-22402 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22402 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22402 | HTTPS FTP |
-Related structure data
| Related structure data |  7jn7MC  7jkqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10596 (Title: Human DPP9-CARD8 complex-VbP / Data size: 2.1 TB EMPIAR-10596 (Title: Human DPP9-CARD8 complex-VbP / Data size: 2.1 TBData #1: Unaligned multi frame micographs of CARD8-DPP9-VbP-noTILT [micrographs - multiframe] Data #2: Unaligned multi frame micographs of CARD8-DPP9-VbP-TILT [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22402.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22402.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8187 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
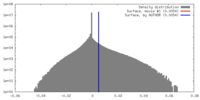
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : DPP9-CARD8 complex
| Entire | Name: DPP9-CARD8 complex |
|---|---|
| Components |
|
-Supramolecule #1: DPP9-CARD8 complex
| Supramolecule | Name: DPP9-CARD8 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Dipeptidyl peptidase 9
| Macromolecule | Name: Dipeptidyl peptidase 9 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: dipeptidyl-peptidase IV |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 101.761984 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSYYHHHHHH DYDIPTTENL YFQGAMGSMA TTGTPTADRG DAAATDDPAA RFQVQKHSWD GLRSIIHGSR KYSGLIVNKA PHDFQFVQK TDESGPHSHR LYYLGMPYGS RENSLLYSEI PKKVRKEALL LLSWKQMLDH FQATPHHGVY SREEELLRER K RLGVFGIT ...String: MSYYHHHHHH DYDIPTTENL YFQGAMGSMA TTGTPTADRG DAAATDDPAA RFQVQKHSWD GLRSIIHGSR KYSGLIVNKA PHDFQFVQK TDESGPHSHR LYYLGMPYGS RENSLLYSEI PKKVRKEALL LLSWKQMLDH FQATPHHGVY SREEELLRER K RLGVFGIT SYDFHSESGL FLFQASNSLF HCRDGGKNGF MVSPMKPLEI KTQCSGPRMD PKICPADPAF FSFINNSDLW VA NIETGEE RRLTFCHQGL SNVLDDPKSA GVATFVIQEE FDRFTGYWWC PTASWEGSEG LKTLRILYEE VDESEVEVIH VPS PALEER KTDSYRYPRT GSKNPKIALK LAEFQTDSQG KIVSTQEKEL VQPFSSLFPK VEYIARAGWT RDGKYAWAMF LDRP QQWLQ LVLLPPALFI PSTENEEQRL ASARAVPRNV QPYVVYEEVT NVWINVHDIF YPFPQSEGED ELCFLRANEC KTGFC HLYK VTAVLKSQGY DWSEPFSPGE DEFKCPIKEE IALTSGEWEV LARHGSKIWV NEETKLVYFQ GTKDTPLEHH LYVVSY EAA GEIVRLTTPG FSHSCSMSQN FDMFVSHYSS VSTPPCVHVY KLSGPDDDPL HKQPRFWASM MEAASCPPDY VPPEIFH FH TRSDVRLYGM IYKPHALQPG KKHPTVLFVY GGPQVQLVNN SFKGIKYLRL NTLASLGYAV VVIDGRGSCQ RGLRFEGA L KNQMGQVEIE DQVEGLQFVA EKYGFIDLSR VAIHGWSYGG FLSLMGLIHK PQVFKVAIAG APVTVWMAYD TGYTERYMD VPENNQHGYE AGSVALHVEK LPNEPNRLLI LHGFLDENVH FFHTNFLVSQ LIRAGKPYQL QIYPNERHSI RCPESGEHYE VTLLHFLQE YL UniProtKB: Dipeptidyl peptidase 9 |
-Macromolecule #2: Caspase recruitment domain-containing protein 8
| Macromolecule | Name: Caspase recruitment domain-containing protein 8 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 60.716875 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEKKECPEKS SSSEEELPRR DSGSSRNIDA SKLIRLQGSR KLLVDNSIRE LQYTKTGIFF QAEACVTNDT VYRELPCVSE TLCDISHFF QEDDETEAEP LLFRAVPECQ LSGGDIPSVS EEQESSEGQD SGDICSEENQ IVSSYASKVC FEIEEDYKNR Q FLGPEGNV ...String: MEKKECPEKS SSSEEELPRR DSGSSRNIDA SKLIRLQGSR KLLVDNSIRE LQYTKTGIFF QAEACVTNDT VYRELPCVSE TLCDISHFF QEDDETEAEP LLFRAVPECQ LSGGDIPSVS EEQESSEGQD SGDICSEENQ IVSSYASKVC FEIEEDYKNR Q FLGPEGNV DVELIDKSTN RYSVWFPTAG WYLWSATGLG FLVRDEVTVT IAFGSWSQHL ALDLQHHEQW LVGGPLFDVT AE PEEAVAE IHLPHFISLQ AGEVDVSWFL VAHFKNEGMV LEHPARVEPF YAVLESPSFS LMGILLRIAS GTRLSIPITS NTL IYYHPH PEDIKFHLYL VPSDALLTKA IDDEEDRFHG VRLQTSPPME PLNFGSSYIV SNSANLKVMP KELKLSYRSP GEIQ HFSKF YAGQMKEPIQ LEITEKRHGT LVWDTEVKPV DLQLVAASAP PPFSGAAFVK ENHRQLQARM GDLKGVLDDL QDNEV LTEN EKELVEQEKT RQSKNEALLS MVEKKGDLAL DVLFRSISER DPYLVSYLRQ QNL UniProtKB: Caspase recruitment domain-containing protein 8 |
-Macromolecule #3: [(2~{R})-1-[(2~{R})-2-azanyl-3-methyl-butanoyl]pyrrolidin-2-yl]bo...
| Macromolecule | Name: [(2~{R})-1-[(2~{R})-2-azanyl-3-methyl-butanoyl]pyrrolidin-2-yl]boronic acid type: ligand / ID: 3 / Number of copies: 2 / Formula: GK2 |
|---|---|
| Molecular weight | Theoretical: 214.07 Da |
| Chemical component information | 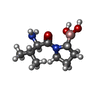 ChemComp-GK2: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 25 mM HEPES, pH 7.5, 150 mM NaCl, 1 mM TCEP |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | #0 - Image recording ID: 1 / #0 - Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / #0 - Number grids imaged: 4 / #0 - Number real images: 3306 / #0 - Average exposure time: 2.22 sec. / #0 - Average electron dose: 58.5 e/Å2 / #0 - Details: stage tilt 0 degrees / #1 - Image recording ID: 2 / #1 - Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / #1 - Number grids imaged: 4 / #1 - Number real images: 2488 / #1 - Average exposure time: 2.25 sec. / #1 - Average electron dose: 64.99 e/Å2 / #1 - Details: stage tilt 37 degrees |
| Electron beam | Acceleration voltage: 300 kV / Electron source: OTHER |
| Electron optics | Calibrated magnification: 10500 / Illumination mode: SPOT SCAN / Imaging mode: OTHER / Cs: 2.7 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: -0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)