[English] 日本語
 Yorodumi
Yorodumi- EMDB-21133: Cryo-EM SPA : Structure of the PCBP2/Stem Loop IVm complex underl... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21133 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM SPA : Structure of the PCBP2/Stem Loop IVm complex underlying the functionality of the poliovirus type I IRES. | |||||||||
 Map data Map data | Structure of the PCBP2/Stem Loop | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Poliovirus type 1 (strain Mahoney) / Poliovirus type 1 (strain Mahoney) /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.1 Å | |||||||||
 Authors Authors | Wilce MCJ / Wilce JA | |||||||||
| Funding support |  Australia, 1 items Australia, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2020 Journal: Nucleic Acids Res / Year: 2020Title: Structure of the PCBP2/stem-loop IV complex underlying translation initiation mediated by the poliovirus type I IRES. Authors: Simone A Beckham / Mehdi Y Matak / Matthew J Belousoff / Hariprasad Venugopal / Neelam Shah / Naveen Vankadari / Hans Elmlund / Joseph H C Nguyen / Bert L Semler / Matthew C J Wilce / Jacqueline A Wilce /   Abstract: The poliovirus type I IRES is able to recruit ribosomal machinery only in the presence of host factor PCBP2 that binds to stem-loop IV of the IRES. When PCBP2 is cleaved in its linker region by viral ...The poliovirus type I IRES is able to recruit ribosomal machinery only in the presence of host factor PCBP2 that binds to stem-loop IV of the IRES. When PCBP2 is cleaved in its linker region by viral proteinase 3CD, translation initiation ceases allowing the next stage of replication to commence. Here, we investigate the interaction of PCBP2 with the apical region of stem-loop IV (SLIVm) of poliovirus RNA in its full-length and truncated form. CryoEM structure reconstruction of the full-length PCBP2 in complex with SLIVm solved to 6.1 Å resolution reveals a compact globular complex of PCBP2 interacting with the cruciform RNA via KH domains and featuring a prominent GNRA tetraloop. SEC-SAXS, SHAPE and hydroxyl-radical cleavage establish that PCBP2 stabilizes the SLIVm structure, but upon cleavage in the linker domain the complex becomes more flexible and base accessible. Limited proteolysis and REMSA demonstrate the accessibility of the linker region in the PCBP2/SLIVm complex and consequent loss of affinity of PCBP2 for the SLIVm upon cleavage. Together this study sheds light on the structural features of the PCBP2/SLIV complex vital for ribosomal docking, and the way in which this key functional interaction is regulated following translation of the poliovirus genome. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21133.map.gz emd_21133.map.gz | 8.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21133-v30.xml emd-21133-v30.xml emd-21133.xml emd-21133.xml | 11.7 KB 11.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21133.png emd_21133.png | 72.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21133 http://ftp.pdbj.org/pub/emdb/structures/EMD-21133 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21133 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21133 | HTTPS FTP |
-Validation report
| Summary document |  emd_21133_validation.pdf.gz emd_21133_validation.pdf.gz | 78.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21133_full_validation.pdf.gz emd_21133_full_validation.pdf.gz | 77.7 KB | Display | |
| Data in XML |  emd_21133_validation.xml.gz emd_21133_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21133 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21133 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21133 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21133 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21133.map.gz / Format: CCP4 / Size: 11.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21133.map.gz / Format: CCP4 / Size: 11.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the PCBP2/Stem Loop | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
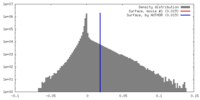
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : truncated stem loop IV of the poliovirus IRES, residues, 278-398 ...
| Entire | Name: truncated stem loop IV of the poliovirus IRES, residues, 278-398 in complex with PolyC binding protein 2 |
|---|---|
| Components |
|
-Supramolecule #1: truncated stem loop IV of the poliovirus IRES, residues, 278-398 ...
| Supramolecule | Name: truncated stem loop IV of the poliovirus IRES, residues, 278-398 in complex with PolyC binding protein 2 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Poliovirus type 1 (strain Mahoney) / Strain: Mahoney Poliovirus type 1 (strain Mahoney) / Strain: Mahoney |
-Macromolecule #1: PolyC Binding Protein 2
| Macromolecule | Name: PolyC Binding Protein 2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MDTGVIEGGL NVTLTIRLLM HGKEVGSIIG KKGESVKKMR EESGARINIS EGNCPERIIT LAGPTNAIF KAFAMIIDKL EEDISSSMTN STAASRPPVT LRLVVPASQC GSLIGKGGCK I KEIRESTG AQVQVAGDML PNSTERAITI AGIPQSIIEC VKQICVVMLE ...String: MDTGVIEGGL NVTLTIRLLM HGKEVGSIIG KKGESVKKMR EESGARINIS EGNCPERIIT LAGPTNAIF KAFAMIIDKL EEDISSSMTN STAASRPPVT LRLVVPASQC GSLIGKGGCK I KEIRESTG AQVQVAGDML PNSTERAITI AGIPQSIIEC VKQICVVMLE TLSQSPPKGV TI PYRPKPS SSPVIFAGGQ DRYSTGSDSA SFPHTTPSMC LNPDLEGPPL EAYTIQGQYA IPQ PDLTKL HQLAMQQSHF PMTHGNTGFS GIESSSPEVK GYWGLDASAQ TTSHELTIPN DLIG CIIGR QGAKINEIRQ MSGAQIKIAN PVEGSTDRQV TITGSAASIS LAQYLINVRL SSETG GMGS SLEHHHHHH |
-Macromolecule #2: Poliovirus (Mahoney) : truncated Stemloop IV from IRES
| Macromolecule | Name: Poliovirus (Mahoney) : truncated Stemloop IV from IRES type: rna / ID: 2 Details: Produced using in vitro transcription : T7 RiboMAX (Promega) |
|---|---|
| Source (natural) | Organism:   Poliovirus type 1 (strain Mahoney) / Strain: Mahoney Poliovirus type 1 (strain Mahoney) / Strain: Mahoney |
| Sequence | String: CGCGCGUUGC GCUCAGCACU CAACCCCAGA GUGUAGCUUA GGCUGAUGAG UCUGGACAUC CCUCACCGGU GACGGUGGUC CAGGCUGCGU UGGCGGCCUA CCUAUGGCUA ACGCCAUGGG ACGCGCG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Concentration | 0.14 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 5 mM HEPES-KOH, 25 mM KCl, 2 mM MgCl2, 2 mM DTT, 4 % glycerol, 0.1 mM EDTA, pH 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV / Details: 2.5 second blot. |
| Details | A mono disperse sample was applied |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Software - Name: CTFFIND / Software - details: CTFFIND |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 6.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 888988 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)