+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21026 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Truncated Tetrahedral RNA Nanostructures | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species | unidentified, produced by invitro transcription of synthetic templates (unknown) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 11.0 Å | |||||||||
 Authors Authors | Wu W / Zakrevsky P / Heinz W / Khant H / Kasprzak WK / Bindewald E / Dorjsuren N / Fields E / De Val N / Jaeger L / Shapiro BA | |||||||||
 Citation Citation |  Journal: Nanoscale / Year: 2020 Journal: Nanoscale / Year: 2020Title: Truncated tetrahedral RNA nanostructures exhibit enhanced features for delivery of RNAi substrates. Authors: Paul Zakrevsky / Wojciech K Kasprzak / William F Heinz / Weimin Wu / Htet Khant / Eckart Bindewald / Nomongo Dorjsuren / Eric A Fields / Natalia de Val / Luc Jaeger / Bruce A Shapiro /  Abstract: Using RNA as a material for nanoparticle construction provides control over particle size and shape at the nano-scale. RNA nano-architectures have shown promise as delivery vehicles for RNA ...Using RNA as a material for nanoparticle construction provides control over particle size and shape at the nano-scale. RNA nano-architectures have shown promise as delivery vehicles for RNA interference (RNAi) substrates, allowing multiple functional entities to be combined on a single particle in a programmable fashion. Rather than employing a completely bottom-up approach to scaffold design, here multiple copies of an existing synthetic supramolecular RNA nano-architecture serve as building blocks along with additional motifs for the design of a novel truncated tetrahedral RNA scaffold, demonstrating that rationally designed RNA assemblies can themselves serve as modular pieces in the construction of larger rationally designed structures. The resulting tetrahedral scaffold displays enhanced characteristics for RNAi-substrate delivery in comparison to similar RNA-based scaffolds, as evidenced by its increased functional capacity, increased cellular uptake and ultimately an increased RNAi efficacy of its adorned Dicer substrate siRNAs. The unique truncated tetrahedral shape of the nanoparticle core appears to contribute to this particle's enhanced function, indicating the physical characteristics of RNA scaffolds merit significant consideration when designing platforms for delivery of functional RNAs via RNA nanoparticles. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21026.map.gz emd_21026.map.gz | 59.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21026-v30.xml emd-21026-v30.xml emd-21026.xml emd-21026.xml | 9.3 KB 9.3 KB | Display Display |  EMDB header EMDB header |
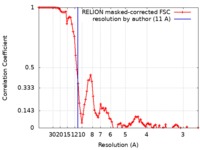
| FSC (resolution estimation) |  emd_21026_fsc.xml emd_21026_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_21026.png emd_21026.png | 157.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21026 http://ftp.pdbj.org/pub/emdb/structures/EMD-21026 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21026 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21026 | HTTPS FTP |
-Validation report
| Summary document |  emd_21026_validation.pdf.gz emd_21026_validation.pdf.gz | 79.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21026_full_validation.pdf.gz emd_21026_full_validation.pdf.gz | 78.2 KB | Display | |
| Data in XML |  emd_21026_validation.xml.gz emd_21026_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21026 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21026 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21026 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21026 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21026.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21026.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.36 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Tetrahedral RNA nanoparticle
| Entire | Name: Tetrahedral RNA nanoparticle |
|---|---|
| Components |
|
-Supramolecule #1: Tetrahedral RNA nanoparticle
| Supramolecule | Name: Tetrahedral RNA nanoparticle / type: complex / ID: 1 / Parent: 0 Details: Supra-molecular RNA structure assembled from rationally designed RNA strands produced by in vitro transcription of synthetic DNA templates |
|---|---|
| Source (natural) | Organism: unidentified, produced by invitro transcription of synthetic templates (unknown) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)