+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20834 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of itraconazole-bound NPC1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Niemann-Pick C disease / cholesterol transport / sterol-sensing domain / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcyclodextrin metabolic process / cholesterol storage / membrane raft organization / intracellular cholesterol transport / intracellular lipid transport / sterol transport / intestinal cholesterol absorption / LDL clearance / negative regulation of epithelial cell apoptotic process / glycoprotein biosynthetic process ...cyclodextrin metabolic process / cholesterol storage / membrane raft organization / intracellular cholesterol transport / intracellular lipid transport / sterol transport / intestinal cholesterol absorption / LDL clearance / negative regulation of epithelial cell apoptotic process / glycoprotein biosynthetic process / cholesterol transfer activity / cholesterol transport / programmed cell death / bile acid metabolic process / establishment of protein localization to membrane / cholesterol efflux / adult walking behavior / lysosomal transport / cholesterol binding / cellular response to steroid hormone stimulus / negative regulation of macroautophagy / response to cadmium ion / cellular response to low-density lipoprotein particle stimulus / cholesterol metabolic process / negative regulation of TORC1 signaling / neurogenesis / cholesterol homeostasis / macroautophagy / liver development / autophagy / endocytosis / transmembrane signaling receptor activity / nuclear envelope / late endosome membrane / virus receptor activity / signaling receptor activity / gene expression / lysosome / membrane raft / response to xenobiotic stimulus / lysosomal membrane / symbiont entry into host cell / perinuclear region of cytoplasm / endoplasmic reticulum / Golgi apparatus / extracellular exosome / extracellular region / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.02 Å | |||||||||
 Authors Authors | Long T / Li X | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Structural basis for itraconazole-mediated NPC1 inhibition. Authors: Tao Long / Xiaofeng Qi / Abdirahman Hassan / Qiren Liang / Jef K De Brabander / Xiaochun Li /  Abstract: Niemann-Pick C1 (NPC1), a lysosomal protein of 13 transmembrane helices (TMs) and three lumenal domains, exports low-density-lipoprotein (LDL)-derived cholesterol from lysosomes. TMs 3-7 of NPC1 ...Niemann-Pick C1 (NPC1), a lysosomal protein of 13 transmembrane helices (TMs) and three lumenal domains, exports low-density-lipoprotein (LDL)-derived cholesterol from lysosomes. TMs 3-7 of NPC1 comprise the Sterol-Sensing Domain (SSD). Previous studies suggest that mutation of the NPC1-SSD or the addition of the anti-fungal drug itraconazole abolishes NPC1 activity in cells. However, the itraconazole binding site and the mechanism of NPC1-mediated cholesterol transport remain unknown. Here, we report a cryo-EM structure of human NPC1 bound to itraconazole, which reveals how this binding site in the center of NPC1 blocks a putative lumenal tunnel linked to the SSD. Functional assays confirm that blocking this tunnel abolishes NPC1-mediated cholesterol egress. Intriguingly, the palmitate anchor of Hedgehog occupies a similar site in the homologous tunnel of Patched, suggesting a conserved mechanism for sterol transport in this family of proteins and establishing a central function of their SSDs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20834.map.gz emd_20834.map.gz | 150 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20834-v30.xml emd-20834-v30.xml emd-20834.xml emd-20834.xml | 10.3 KB 10.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20834.png emd_20834.png | 100.7 KB | ||
| Filedesc metadata |  emd-20834.cif.gz emd-20834.cif.gz | 5.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20834 http://ftp.pdbj.org/pub/emdb/structures/EMD-20834 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20834 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20834 | HTTPS FTP |
-Related structure data
| Related structure data |  6uoxMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20834.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20834.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.66 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : NPC1 and itraconazole-Br complex
| Entire | Name: NPC1 and itraconazole-Br complex |
|---|---|
| Components |
|
-Supramolecule #1: NPC1 and itraconazole-Br complex
| Supramolecule | Name: NPC1 and itraconazole-Br complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: NPC intracellular cholesterol transporter 1
| Macromolecule | Name: NPC intracellular cholesterol transporter 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 143.315016 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MTARGLALGL LLLLLCPAQV FSQSCVWYGE CGIAYGDKRY NCEYSGPPKP LPKDGYDLVQ ELCPGFFFGN VSLCCDVRQL QTLKDNLQL PLQFLSRCPS CFYNLLNLFC ELTCSPRQSQ FLNVTATEDY VDPVTNQTKT NVKELQYYVG QSFANAMYNA C RDVEAPSS ...String: MTARGLALGL LLLLLCPAQV FSQSCVWYGE CGIAYGDKRY NCEYSGPPKP LPKDGYDLVQ ELCPGFFFGN VSLCCDVRQL QTLKDNLQL PLQFLSRCPS CFYNLLNLFC ELTCSPRQSQ FLNVTATEDY VDPVTNQTKT NVKELQYYVG QSFANAMYNA C RDVEAPSS NDKALGLLCG KDADACNATN WIEYMFNKDN GQAPFTITPV FSDFPVHGME PMNNATKGCD ESVDEVTAPC SC QDCSIVC GPKPQPPPPP APWTILGLDA MYVIMWITYM AFLLVFFGAF FAVWCYRKRY FVSEYTPIDS NIAFSVNASD KGE ASCCDP VSAAFEGCLR RLFTRWGSFC VRNPGCVIFF SLVFITACSS GLVFVRVTTN PVDLWSAPSS QARLEKEYFD QHFG PFFRT EQLIIRAPLT DKHIYQPYPS GADVPFGPPL DIQILHQVLD LQIAIENITA SYDNETVTLQ DICLAPLSPY NTNCT ILSV LNYFQNSHSV LDHKKGDDFF VYADYHTHFL YCVRAPASLN DTSLLHDPCL GTFGGPVFPW LVLGGYDDQN YNNATA LVI TFPVNNYYND TEKLQRAQAW EKEFINFVKN YKNPNLTISF TAERSIEDEL NRESDSDVFT VVISYAIMFL YISLALG HM KSCRRLLVDS KVSLGIAGIL IVLSSVACSL GVFSYIGLPL TLIVIEVIPF LVLAVGVDNI FILVQAYQRD ERLQGETL D QQLGRVLGEV APSMFLSSFS ETVAFFLGAL SVMPAVHTFS LFAGLAVFID FLLQITCFVS LLGLDIKRQE KNRLDIFCC VRGAEDGTSV QASESCLFRF FKNSYSPLLL KDWMRPIVIA IFVGVLSFSI AVLNKVDIGL DQSLSMPDDS YMVDYFKSIS QYLHAGPPV YFVLEEGHDY TSSKGQNMVC GGMGCNNDSL VQQIFNAAQL DNYTRIGFAP SSWIDDYFDW VKPQSSCCRV D NITDQFCN ASVVDPACVR CRPLTPEGKQ RPQGGDFMRF LPMFLSDNPN PKCGKGGHAA YSSAVNILLG HGTRVGATYF MT YHTVLQT SADFIDALKK ARLIASNVTE TMGINGSAYR VFPYSVFYVF YEQYLTIIDD TIFNLGVSLG AIFLVTMVLL GCE LWSAVI MCATIAMVLV NMFGVMWLWG ISLNAVSLVN LVMSCGISVE FCSHITRAFT VSMKGSRVER AEEALAHMGS SVFS GITLT KFGGIVVLAF AKSQIFQIFY FRMYLAMVLL GATHGLIFLP VLLSYIGPSV NKAKSCATEE RYKGTERERL LNFWS HPQF EK UniProtKB: NPC intracellular cholesterol transporter 1 |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 8 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: 4-(3-bromo-4-{4-[4-({(2R,4S)-2-(2,4-dichlorophenyl)-2-[(1H-1,2,4-...
| Macromolecule | Name: 4-(3-bromo-4-{4-[4-({(2R,4S)-2-(2,4-dichlorophenyl)-2-[(1H-1,2,4-triazol-1-yl)methyl]-1,3-dioxolan-4-yl}methoxy)phenyl]piperazin-1-yl}phenyl)-2-[(2S)-butan-2-yl]-2,4-dihydro-3H-1,2,4-triazol-3-one type: ligand / ID: 5 / Number of copies: 1 / Formula: QDG |
|---|---|
| Molecular weight | Theoretical: 784.529 Da |
| Chemical component information | 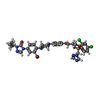 ChemComp-QDG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 80.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.02 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 209612 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















