+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20238 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Dimeric structure of ACAT1 | |||||||||
 Map data Map data | Dimeric structure of ACAT1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | MBOAT / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationsterol O-acyltransferase / sterol O-acyltransferase activity / cholesterol O-acyltransferase activity / macrophage derived foam cell differentiation / cholesterol storage / positive regulation of amyloid precursor protein biosynthetic process / very-low-density lipoprotein particle assembly / fatty-acyl-CoA binding / low-density lipoprotein particle clearance / LDL clearance ...sterol O-acyltransferase / sterol O-acyltransferase activity / cholesterol O-acyltransferase activity / macrophage derived foam cell differentiation / cholesterol storage / positive regulation of amyloid precursor protein biosynthetic process / very-low-density lipoprotein particle assembly / fatty-acyl-CoA binding / low-density lipoprotein particle clearance / LDL clearance / cholesterol efflux / cholesterol binding / cholesterol metabolic process / cholesterol homeostasis / endoplasmic reticulum membrane / endoplasmic reticulum / identical protein binding / membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Yan N / Qian HW | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Structural basis for catalysis and substrate specificity of human ACAT1. Authors: Hongwu Qian / Xin Zhao / Renhong Yan / Xia Yao / Shuai Gao / Xue Sun / Ximing Du / Hongyuan Yang / Catherine C L Wong / Nieng Yan /    Abstract: As members of the membrane-bound O-acyltransferase (MBOAT) enzyme family, acyl-coenzyme A:cholesterol acyltransferases (ACATs) catalyse the transfer of an acyl group from acyl-coenzyme A to ...As members of the membrane-bound O-acyltransferase (MBOAT) enzyme family, acyl-coenzyme A:cholesterol acyltransferases (ACATs) catalyse the transfer of an acyl group from acyl-coenzyme A to cholesterol to generate cholesteryl ester, the primary form in which cholesterol is stored in cells and transported in plasma. ACATs have gained attention as potential drug targets for the treatment of diseases such as atherosclerosis, Alzheimer's disease and cancer. Here we present the cryo-electron microscopy structure of human ACAT1 as a dimer of dimers. Each protomer consists of nine transmembrane segments, which enclose a cytosolic tunnel and a transmembrane tunnel that converge at the predicted catalytic site. Evidence from structure-guided mutational analyses suggests that acyl-coenzyme A enters the active site through the cytosolic tunnel, whereas cholesterol may enter from the side through the transmembrane tunnel. This structural and biochemical characterization helps to rationalize the preference of ACAT1 for unsaturated acyl chains, and provides insight into the catalytic mechanism of enzymes within the MBOAT family. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20238.map.gz emd_20238.map.gz | 77.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20238-v30.xml emd-20238-v30.xml emd-20238.xml emd-20238.xml | 13.4 KB 13.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20238.png emd_20238.png | 134.7 KB | ||
| Filedesc metadata |  emd-20238.cif.gz emd-20238.cif.gz | 6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20238 http://ftp.pdbj.org/pub/emdb/structures/EMD-20238 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20238 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20238 | HTTPS FTP |
-Validation report
| Summary document |  emd_20238_validation.pdf.gz emd_20238_validation.pdf.gz | 496.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_20238_full_validation.pdf.gz emd_20238_full_validation.pdf.gz | 496.4 KB | Display | |
| Data in XML |  emd_20238_validation.xml.gz emd_20238_validation.xml.gz | 6.1 KB | Display | |
| Data in CIF |  emd_20238_validation.cif.gz emd_20238_validation.cif.gz | 7.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20238 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20238 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20238 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20238 | HTTPS FTP |
-Related structure data
| Related structure data |  6p2jMC  6p2pC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20238.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20238.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Dimeric structure of ACAT1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.114 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
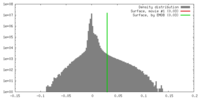
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ACAT1
| Entire | Name: ACAT1 |
|---|---|
| Components |
|
-Supramolecule #1: ACAT1
| Supramolecule | Name: ACAT1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Sterol O-acyltransferase 1
| Macromolecule | Name: Sterol O-acyltransferase 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: sterol O-acyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 69.745375 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADYKDDDDK SGPDEVDASG RMVGEEKMSL RNRLSKSREN PEEDEDQRNP AKESLETPSN GRIDIKQLIA KKIKLTAEAE ELKPFFMKE VGSHFDDFVT NLIEKSASLD NGGCALTTFS VLEGEKNNHR AKDLRAPPEQ GKIFIARRSL LDELLEVDHI R TIYHMFIA ...String: MADYKDDDDK SGPDEVDASG RMVGEEKMSL RNRLSKSREN PEEDEDQRNP AKESLETPSN GRIDIKQLIA KKIKLTAEAE ELKPFFMKE VGSHFDDFVT NLIEKSASLD NGGCALTTFS VLEGEKNNHR AKDLRAPPEQ GKIFIARRSL LDELLEVDHI R TIYHMFIA LLILFILSTL VVDYIDEGRL VLEFSLLSYA FGKFPTVVWT WWIMFLSTFS VPYFLFQHWA TGYSKSSHPL IR SLFHGFL FMIFQIGVLG FGPTYVVLAY TLPPASRFII IFEQIRFVMK AHSFVRENVP RVLNSAKEKS STVPIPTVNQ YLY FLFAPT LIYRDSYPRN PTVRWGYVAM KFAQVFGCFF YVYYIFERLC APLFRNIKQE PFSARVLVLC VFNSILPGVL ILFL TFFAF LHCWLNAFAE MLRFGDRMFY KDWWNSTSYS NYYRTWNVVV HDWLYYYAYK DFLWFFSKRF KSAAMLAVFA VSAVV HEYA LAVCLSFFYP VLFVLFMFFG MAFNFIVNDS RKKPIWNVLM WTSLFLGNGV LLCFYSQEWY ARQHCPLKNP TFLDYV RPR SWTCRYVFLE GSDEVDAVEG SHHHHHHHHH H UniProtKB: Sterol O-acyltransferase 1 |
-Macromolecule #2: S-{(3R,5R,9R)-1-[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydrox...
| Macromolecule | Name: S-{(3R,5R,9R)-1-[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)tetrahydrofuran-2-yl]-3,5,9-trihydroxy-8,8-dimethyl-3,5-dioxido-10,14-dioxo-2,4,6-trioxa-11,15-diaza- ...Name: S-{(3R,5R,9R)-1-[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)tetrahydrofuran-2-yl]-3,5,9-trihydroxy-8,8-dimethyl-3,5-dioxido-10,14-dioxo-2,4,6-trioxa-11,15-diaza-3lambda~5~,5lambda~5~-diphosphaheptadecan-17-yl} (9Z)-octadec-9-enethioate (non-preferred name) type: ligand / ID: 2 / Number of copies: 2 / Formula: 3VV |
|---|---|
| Molecular weight | Theoretical: 1.03198 KDa |
| Chemical component information |  ChemComp-3VV: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)