+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
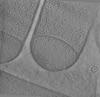
| Title | Tomograms of spoIVB Bacillus subtilis sporangia | |||||||||||||||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||||||||
 Keywords Keywords | Sporulation / Bacillus subtilis / coat / macromolecular assemblies / STRUCTURAL PROTEIN | |||||||||||||||||||||||||||||||||
| Biological species |  | |||||||||||||||||||||||||||||||||
| Method | electron tomography / cryo EM | |||||||||||||||||||||||||||||||||
 Authors Authors | Bauda E / Gallet B / Moravcova J / Effantin G / Chan H / Novacek J / Jouneau PH / Rodrigues CDA / Schoehn G / Moriscot C / Morlot C | |||||||||||||||||||||||||||||||||
| Funding support |  France, European Union, France, European Union,  Australia, 10 items Australia, 10 items
| |||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Ultrastructure of macromolecular assemblies contributing to bacterial spore resistance revealed by in situ cryo-electron tomography. Authors: Elda Bauda / Benoit Gallet / Jana Moravcova / Gregory Effantin / Helena Chan / Jiri Novacek / Pierre-Henri Jouneau / Christopher D A Rodrigues / Guy Schoehn / Christine Moriscot / Cecile Morlot /     Abstract: Bacterial spores owe their incredible resistance capacities to molecular structures that protect the cell content from external aggressions. Among the determinants of resistance are the quaternary ...Bacterial spores owe their incredible resistance capacities to molecular structures that protect the cell content from external aggressions. Among the determinants of resistance are the quaternary structure of the chromosome and an extracellular shell made of proteinaceous layers (the coat), the assembly of which remains poorly understood. Here, in situ cryo-electron tomography on lamellae generated by cryo-focused ion beam micromachining provides insights into the ultrastructural organization of Bacillus subtilis sporangia. The reconstructed tomograms reveal that early during sporulation, the chromosome in the forespore adopts a toroidal structure harboring 5.5-nm thick fibers. At the same stage, coat proteins at the surface of the forespore form a stack of amorphous or structured layers with distinct electron density, dimensions and organization. By analyzing mutant strains using cryo-electron tomography and transmission electron microscopy on resin sections, we distinguish seven nascent coat regions with different molecular properties, and propose a model for the contribution of coat morphogenetic proteins. | |||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19405.map.gz emd_19405.map.gz | 3.1 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19405-v30.xml emd-19405-v30.xml emd-19405.xml emd-19405.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_19405.png emd_19405.png | 188.4 KB | ||
| Filedesc metadata |  emd-19405.cif.gz emd-19405.cif.gz | 4.4 KB | ||
| Others |  emd_19405_additional_1.map.gz emd_19405_additional_1.map.gz emd_19405_additional_2.map.gz emd_19405_additional_2.map.gz emd_19405_additional_3.map.gz emd_19405_additional_3.map.gz emd_19405_additional_4.map.gz emd_19405_additional_4.map.gz emd_19405_additional_5.map.gz emd_19405_additional_5.map.gz emd_19405_additional_6.map.gz emd_19405_additional_6.map.gz | 3.7 GB 3.1 GB 5.6 GB 3.7 GB 3.2 GB 3.7 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19405 http://ftp.pdbj.org/pub/emdb/structures/EMD-19405 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19405 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19405 | HTTPS FTP |
-Validation report
| Summary document |  emd_19405_validation.pdf.gz emd_19405_validation.pdf.gz | 444.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19405_full_validation.pdf.gz emd_19405_full_validation.pdf.gz | 443.9 KB | Display | |
| Data in XML |  emd_19405_validation.xml.gz emd_19405_validation.xml.gz | 4.6 KB | Display | |
| Data in CIF |  emd_19405_validation.cif.gz emd_19405_validation.cif.gz | 5.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19405 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19405 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19405 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19405 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19405.map.gz / Format: CCP4 / Size: 3.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19405.map.gz / Format: CCP4 / Size: 3.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 8.812 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #6
| File | emd_19405_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
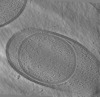
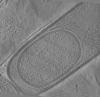
| Projections & Slices |
| ||||||||||||
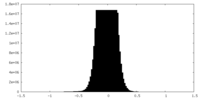
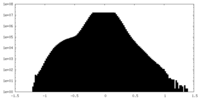
| Density Histograms |
-Additional map: #5
| File | emd_19405_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #4
| File | emd_19405_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #3
| File | emd_19405_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #2
| File | emd_19405_additional_5.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_19405_additional_6.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Sporulating cells of Bacillus subtilis strain 168 deleted from th...
| Entire | Name: Sporulating cells of Bacillus subtilis strain 168 deleted from the spoIVB gene |
|---|---|
| Components |
|
-Supramolecule #1: Sporulating cells of Bacillus subtilis strain 168 deleted from th...
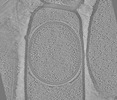
| Supramolecule | Name: Sporulating cells of Bacillus subtilis strain 168 deleted from the spoIVB gene type: cell / ID: 1 / Parent: 0 Details: View of a developing spore within a Bacillus subtilis mother cell |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
| Cryo protectant | 5% glycerol |
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 / Focused ion beam - Current: 1 / Focused ion beam - Duration: 20 / Focused ion beam - Temperature: 83 K / Focused ion beam - Initial thickness: 4000 / Focused ion beam - Final thickness: 150 Focused ion beam - Details: The value given for _em_focused_ion_beam.instrument is Versa 3D Dual Beam. This is not in a list of allowed values {'DB235', 'OTHER'} so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 85.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 20.0 µm / Nominal defocus min: 10.0 µm / Nominal magnification: 33000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: BACK PROJECTION / Number images used: 41 |
|---|
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X