[English] 日本語
 Yorodumi
Yorodumi- EMDB-19195: Structure of the rabbit 80S ribosome stalled on a 2-TMD rhodopsin... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the rabbit 80S ribosome stalled on a 2-TMD rhodopsin intermediate in complex with Sec61-RAMP4 | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Ribosome / Membrane protein / Translocon / Transport | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationL13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / Major pathway of rRNA processing in the nucleolus and cytosol / Formation of a pool of free 40S subunits / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / muscle organ morphogenesis / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / membrane docking ...L13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / Major pathway of rRNA processing in the nucleolus and cytosol / Formation of a pool of free 40S subunits / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / muscle organ morphogenesis / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / membrane docking / endoplasmic reticulum Sec complex / pronephric nephron development / cotranslational protein targeting to membrane / Ssh1 translocon complex / Sec61 translocon complex / positive regulation of growth hormone secretion / protein insertion into ER membrane / protein-transporting ATPase activity / protein targeting to ER / positive regulation of organ growth / SRP-dependent cotranslational protein targeting to membrane, translocation / signal sequence receptor activity / post-translational protein targeting to membrane, translocation / insulin secretion / transmembrane protein transporter activity / ubiquitin ligase inhibitor activity / positive regulation of signal transduction by p53 class mediator / cytoplasmic microtubule / protein-RNA complex assembly / endoplasmic reticulum unfolded protein response / rough endoplasmic reticulum / MDM2/MDM4 family protein binding / guanyl-nucleotide exchange factor activity / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / post-embryonic development / positive regulation of translation / skeletal system development / phospholipid binding / glucose metabolic process / positive regulation of insulin secretion / antimicrobial humoral immune response mediated by antimicrobial peptide / heparin binding / large ribosomal subunit / ribosome binding / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / killing of cells of another organism / cytosolic large ribosomal subunit / defense response to Gram-negative bacterium / cytoplasmic translation / tRNA binding / postsynaptic density / rRNA binding / nuclear body / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / synapse / endoplasmic reticulum membrane / nucleolus / endoplasmic reticulum / RNA binding / zinc ion binding / membrane / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |      | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.69243 Å | ||||||||||||
 Authors Authors | Lewis AJO / Hegde RS | ||||||||||||
| Funding support |  United Kingdom, United Kingdom,  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation | Journal: Biochem J / Year: 2021 Title: New tools for automated cryo-EM single-particle analysis in RELION-4.0. Authors: Dari Kimanius / Liyi Dong / Grigory Sharov / Takanori Nakane / Sjors H W Scheres /  Abstract: We describe new tools for the processing of electron cryo-microscopy (cryo-EM) images in the fourth major release of the RELION software. In particular, we introduce VDAM, a variable-metric gradient ...We describe new tools for the processing of electron cryo-microscopy (cryo-EM) images in the fourth major release of the RELION software. In particular, we introduce VDAM, a variable-metric gradient descent algorithm with adaptive moments estimation, for image refinement; a convolutional neural network for unsupervised selection of 2D classes; and a flexible framework for the design and execution of multiple jobs in pre-defined workflows. In addition, we present a stand-alone utility called MDCatch that links the execution of jobs within this framework with metadata gathering during microscope data acquisition. The new tools are aimed at providing fast and robust procedures for unsupervised cryo-EM structure determination, with potential applications for on-the-fly processing and the development of flexible, high-throughput structure determination pipelines. We illustrate their potential on 12 publicly available cryo-EM data sets. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19195.map.gz emd_19195.map.gz | 263.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19195-v30.xml emd-19195-v30.xml emd-19195.xml emd-19195.xml | 73.1 KB 73.1 KB | Display Display |  EMDB header EMDB header |
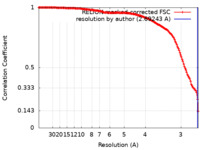
| FSC (resolution estimation) |  emd_19195_fsc.xml emd_19195_fsc.xml | 14.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_19195.png emd_19195.png | 141.5 KB | ||
| Filedesc metadata |  emd-19195.cif.gz emd-19195.cif.gz | 15.7 KB | ||
| Others |  emd_19195_half_map_1.map.gz emd_19195_half_map_1.map.gz emd_19195_half_map_2.map.gz emd_19195_half_map_2.map.gz | 224.8 MB 224.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19195 http://ftp.pdbj.org/pub/emdb/structures/EMD-19195 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19195 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19195 | HTTPS FTP |
-Related structure data
| Related structure data |  8rjbMC  8rjcC  8rjdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19195.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19195.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3398 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_19195_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
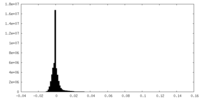
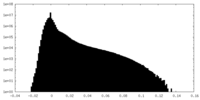
| Density Histograms |
-Half map: #2
| File | emd_19195_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
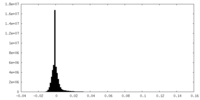
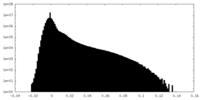
| Density Histograms |
- Sample components
Sample components
+Entire : 80S ribosome translating a stalled, two-TMD nascent chain (derive...
+Supramolecule #1: 80S ribosome translating a stalled, two-TMD nascent chain (derive...
+Macromolecule #1: Protein transport protein Sec61 subunit alpha isoform 1
+Macromolecule #2: Protein transport protein Sec61 subunit beta
+Macromolecule #3: Protein transport protein Sec61 subunit gamma
+Macromolecule #4: Stress-associated endoplasmic reticulum protein
+Macromolecule #5: Ribosomal protein L8
+Macromolecule #6: Nascent chain
+Macromolecule #7: Large ribosomal subunit protein uL4
+Macromolecule #8: Ribosomal_L18_c domain-containing protein
+Macromolecule #9: 60S ribosomal protein L6
+Macromolecule #10: Ribosomal Protein uL30
+Macromolecule #11: 60S ribosomal protein L7a
+Macromolecule #12: 60S ribosomal protein L9
+Macromolecule #13: 60S ribosomal protein L10
+Macromolecule #14: 60S ribosomal protein L11
+Macromolecule #16: 60S ribosomal protein L13
+Macromolecule #17: 60S ribosomal protein L14
+Macromolecule #18: Ribosomal protein L15
+Macromolecule #19: 60S ribosomal protein L13
+Macromolecule #20: uL22
+Macromolecule #21: Ribosomal protein L26
+Macromolecule #22: Ribosomal protein L19
+Macromolecule #23: 60S ribosomal protein L18a
+Macromolecule #24: 60S ribosomal protein L21
+Macromolecule #25: Ribosomal protein L22
+Macromolecule #26: Ribosomal protein L23
+Macromolecule #27: Ribosomal protein L24
+Macromolecule #28: Ribosomal_L23eN domain-containing protein
+Macromolecule #29: Ribosomal protein L26
+Macromolecule #30: 60S ribosomal protein L27
+Macromolecule #31: 60S ribosomal protein L27a
+Macromolecule #32: 60S ribosomal protein L29
+Macromolecule #33: Large ribosomal subunit protein eL30
+Macromolecule #34: 60S ribosomal protein L31
+Macromolecule #35: Ribosomal protein L32
+Macromolecule #36: 60S ribosomal protein L35a
+Macromolecule #37: 60S ribosomal protein L34
+Macromolecule #38: 60S ribosomal protein L35
+Macromolecule #39: 60S ribosomal protein L36
+Macromolecule #40: Ribosomal protein L37
+Macromolecule #41: 60S ribosomal protein L38
+Macromolecule #42: 60S ribosomal protein L39-like
+Macromolecule #43: eL40
+Macromolecule #44: 60S ribosomal protein L41
+Macromolecule #45: 60S ribosomal protein L36a-like
+Macromolecule #46: 60S ribosomal protein L37a
+Macromolecule #48: 60S ribosomal protein L28
+Macromolecule #51: Ribosomal protein L3
+Macromolecule #15: 28S rRNA
+Macromolecule #47: P-site tRNA
+Macromolecule #49: 5S rRNA
+Macromolecule #50: 5.8S rRNA
+Macromolecule #52: MAGNESIUM ION
+Macromolecule #53: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 17540 / Average electron dose: 54.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.7 µm / Nominal defocus min: 1.9000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8rjb: |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)