[English] 日本語
 Yorodumi
Yorodumi- EMDB-19173: CryoEM structure of human rho1 GABAA receptor in complex with pre... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of human rho1 GABAA receptor in complex with pregnenolone sulfate | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationGABA receptor activation / GABA-gated chloride ion channel activity / GABA-A receptor complex / GABA-A receptor activity / gamma-aminobutyric acid signaling pathway / synaptic transmission, GABAergic / intracellular vesicle / chloride channel complex / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / chloride transmembrane transport ...GABA receptor activation / GABA-gated chloride ion channel activity / GABA-A receptor complex / GABA-A receptor activity / gamma-aminobutyric acid signaling pathway / synaptic transmission, GABAergic / intracellular vesicle / chloride channel complex / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / chloride transmembrane transport / modulation of chemical synaptic transmission / GABA-ergic synapse / presynaptic membrane / chemical synaptic transmission / postsynaptic membrane / protein domain specific binding / protein-containing complex binding / glutamatergic synapse / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.21 Å | |||||||||
 Authors Authors | Chen F / John C / Rebecca JH / Erik L | |||||||||
| Funding support |  Sweden, 1 items Sweden, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Divergent mechanisms of steroid inhibition in the human rho1 GABAA receptor Authors: Chen F / John C / Rebecca JH / Erik L | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19173.map.gz emd_19173.map.gz | 266.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19173-v30.xml emd-19173-v30.xml emd-19173.xml emd-19173.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19173_fsc.xml emd_19173_fsc.xml | 13.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_19173.png emd_19173.png | 173.3 KB | ||
| Filedesc metadata |  emd-19173.cif.gz emd-19173.cif.gz | 6.3 KB | ||
| Others |  emd_19173_half_map_1.map.gz emd_19173_half_map_1.map.gz emd_19173_half_map_2.map.gz emd_19173_half_map_2.map.gz | 262.6 MB 262.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19173 http://ftp.pdbj.org/pub/emdb/structures/EMD-19173 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19173 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19173 | HTTPS FTP |
-Related structure data
| Related structure data |  8rh9MC  8rh4C  8rh7C  8rh8C  8rhgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19173.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19173.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.6725 Å | ||||||||||||||||||||||||||||||||||||
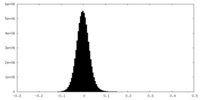
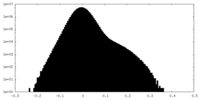
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_19173_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
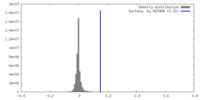
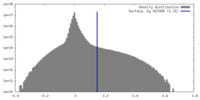
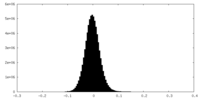
| Density Histograms |
-Half map: #1
| File | emd_19173_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : human rho1 GABAA receptor in complex with estradiol
+Supramolecule #1: human rho1 GABAA receptor in complex with estradiol
+Macromolecule #1: Gamma-aminobutyric acid receptor subunit rho-1
+Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
+Macromolecule #3: N-OCTANE
+Macromolecule #4: DECANE
+Macromolecule #5: DODECANE
+Macromolecule #6: HEXANE
+Macromolecule #7: Pregnenolone sulfate
+Macromolecule #8: CHLORIDE ION
+Macromolecule #9: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)