[English] 日本語
 Yorodumi
Yorodumi- EMDB-19052: CryoEM structure of mTORC1 with a paediatric kidney cancer-associ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of mTORC1 with a paediatric kidney cancer-associated 1455-EWED-1458 duplication in mTOR, overall refinement | ||||||||||||
 Map data Map data | EM map of mTORC1 with the cancer-associated 1455-EWED-1458 duplication variant. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Target of Rapamycin / mTOR / Cancer-associated mutants / SIGNALING PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationActivation of the mRNA upon binding of the cap-binding complex and eIFs, and subsequent binding to 43S / positive regulation of SCF-dependent proteasomal ubiquitin-dependent catabolic process / RNA polymerase III type 2 promoter sequence-specific DNA binding / eukaryotic initiation factor 4E binding / RNA polymerase III type 1 promoter sequence-specific DNA binding / positive regulation of cytoplasmic translational initiation / regulation of locomotor rhythm / T-helper 1 cell lineage commitment / positive regulation of pentose-phosphate shunt / positive regulation of wound healing, spreading of epidermal cells ...Activation of the mRNA upon binding of the cap-binding complex and eIFs, and subsequent binding to 43S / positive regulation of SCF-dependent proteasomal ubiquitin-dependent catabolic process / RNA polymerase III type 2 promoter sequence-specific DNA binding / eukaryotic initiation factor 4E binding / RNA polymerase III type 1 promoter sequence-specific DNA binding / positive regulation of cytoplasmic translational initiation / regulation of locomotor rhythm / T-helper 1 cell lineage commitment / positive regulation of pentose-phosphate shunt / positive regulation of wound healing, spreading of epidermal cells / regulation of membrane permeability / TORC2 complex / cellular response to leucine starvation / TFIIIC-class transcription factor complex binding / heart valve morphogenesis / voluntary musculoskeletal movement / positive regulation of odontoblast differentiation / negative regulation of lysosome organization / TORC1 complex / calcineurin-NFAT signaling cascade / positive regulation of transcription of nucleolar large rRNA by RNA polymerase I / RNA polymerase III type 3 promoter sequence-specific DNA binding / positive regulation of keratinocyte migration / regulation of osteoclast differentiation / MTOR signalling / energy reserve metabolic process / regulation of lysosome organization / cellular response to L-leucine / Energy dependent regulation of mTOR by LKB1-AMPK / cellular response to nutrient / regulation of autophagosome assembly / Amino acids regulate mTORC1 / cellular response to methionine / negative regulation of cell size / positive regulation of osteoclast differentiation / TORC2 signaling / cellular response to osmotic stress / cell projection organization / anoikis / inositol hexakisphosphate binding / cardiac muscle cell development / negative regulation of calcineurin-NFAT signaling cascade / positive regulation of ubiquitin-dependent protein catabolic process / regulation of myelination / negative regulation of protein localization to nucleus / positive regulation of transcription by RNA polymerase III / positive regulation of ruffle assembly / regulation of cell size / positive regulation of myotube differentiation / negative regulation of macroautophagy / Macroautophagy / Constitutive Signaling by AKT1 E17K in Cancer / germ cell development / positive regulation of actin filament polymerization / oligodendrocyte differentiation / TORC1 signaling / positive regulation of oligodendrocyte differentiation / behavioral response to pain / response to amino acid / TOR signaling / mTORC1-mediated signalling / positive regulation of translational initiation / CD28 dependent PI3K/Akt signaling / HSF1-dependent transactivation / regulation of macroautophagy / positive regulation of TOR signaling / protein kinase activator activity / protein serine/threonine kinase inhibitor activity / enzyme-substrate adaptor activity / 'de novo' pyrimidine nucleobase biosynthetic process / positive regulation of epithelial to mesenchymal transition / social behavior / positive regulation of lipid biosynthetic process / positive regulation of G1/S transition of mitotic cell cycle / vascular endothelial cell response to laminar fluid shear stress / heart morphogenesis / regulation of cellular response to heat / positive regulation of lamellipodium assembly / neuronal action potential / T cell costimulation / phagocytic vesicle / positive regulation of stress fiber assembly / cardiac muscle contraction / positive regulation of endothelial cell proliferation / negative regulation of insulin receptor signaling pathway / translation initiation factor binding / 14-3-3 protein binding / cytoskeleton organization / translation repressor activity / endomembrane system / positive regulation of mitotic cell cycle / negative regulation of translational initiation / cellular response to nutrient levels / positive regulation of glycolytic process / Regulation of PTEN gene transcription / cellular response to amino acid starvation / cellular response to starvation / regulation of signal transduction by p53 class mediator / negative regulation of autophagy / VEGFR2 mediated vascular permeability Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | ||||||||||||
 Authors Authors | Anandapadamanaban M / Hay IM / Perisic O / Williams RL | ||||||||||||
| Funding support |  United Kingdom, European Union, 3 items United Kingdom, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: cryoEM structure of mTORC1 with the cancer-associated 1455-EWED-1458 duplication variant, overall refinement Authors: Anandapadamanaban M / Hay I / Perisic O / Williams R | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19052.map.gz emd_19052.map.gz | 479 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19052-v30.xml emd-19052-v30.xml emd-19052.xml emd-19052.xml | 24.4 KB 24.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19052_fsc.xml emd_19052_fsc.xml | 18.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_19052.png emd_19052.png | 113.4 KB | ||
| Masks |  emd_19052_msk_1.map emd_19052_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19052.cif.gz emd-19052.cif.gz | 9 KB | ||
| Others |  emd_19052_half_map_1.map.gz emd_19052_half_map_1.map.gz emd_19052_half_map_2.map.gz emd_19052_half_map_2.map.gz | 411.1 MB 410.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19052 http://ftp.pdbj.org/pub/emdb/structures/EMD-19052 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19052 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19052 | HTTPS FTP |
-Related structure data
| Related structure data |  8rchMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19052.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19052.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM map of mTORC1 with the cancer-associated 1455-EWED-1458 duplication variant. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19052_msk_1.map emd_19052_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
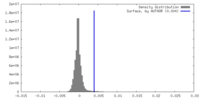
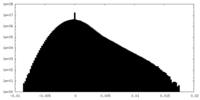
| Density Histograms |
-Half map: EM Half-1 map of mTORC1 with the cancer-associated...
| File | emd_19052_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM Half-1 map of mTORC1 with the cancer-associated 1455-EWED-1458 duplication variant. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: EM Half-2 map of mTORC1 with the cancer-associated...
| File | emd_19052_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM Half-2 map of mTORC1 with the cancer-associated 1455-EWED-1458 duplication variant. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : mTORC1 complex is a dimer of a heterotrimeric complex including m...
| Entire | Name: mTORC1 complex is a dimer of a heterotrimeric complex including mTOR, RAPTOR and mLST8 |
|---|---|
| Components |
|
-Supramolecule #1: mTORC1 complex is a dimer of a heterotrimeric complex including m...
| Supramolecule | Name: mTORC1 complex is a dimer of a heterotrimeric complex including mTOR, RAPTOR and mLST8 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: This cancer-associated mTORC1 variant consists of mTOR 1455-EWED-1458 duplication, RAPTOR and mLST8 complex and its bound to a substrate 4EBP1. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 1.09 MDa |
-Macromolecule #1: Serine/threonine-protein kinase mTOR
| Macromolecule | Name: Serine/threonine-protein kinase mTOR / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: non-specific serine/threonine protein kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 289.257969 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLGTGPAAAT TAATTSSNVS VLQQFASGLK SRNEETRAKA AKELQHYVTM ELREMSQEES TRFYDQLNHH IFELVSSSDA NERKGGILA IASLIGVEGG NATRIGRFAN YLRNLLPSND PVVMEMASKA IGRLAMAGDT FTAEYVEFEV KRALEWLGAD R NEGRRHAA ...String: MLGTGPAAAT TAATTSSNVS VLQQFASGLK SRNEETRAKA AKELQHYVTM ELREMSQEES TRFYDQLNHH IFELVSSSDA NERKGGILA IASLIGVEGG NATRIGRFAN YLRNLLPSND PVVMEMASKA IGRLAMAGDT FTAEYVEFEV KRALEWLGAD R NEGRRHAA VLVLRELAIS VPTFFFQQVQ PFFDNIFVAV WDPKQAIREG AVAALRACLI LTTQREPKEM QKPQWYRHTF EE AEKGFDE TLAKEKGMNR DDRIHGALLI LNELVRISSM EGERLREEME EITQQQLVHD KYCKDLMGFG TKPRHITPFT SFQ AVQPQQ SNALVGLLGY SSHQGLMGFG TSPSPAKSTL VESRCCRDLM EEKFDQVCQW VLKCRNSKNS LIQMTILNLL PRLA AFRPS AFTDTQYLQD TMNHVLSCVK KEKERTAAFQ ALGLLSVAVR SEFKVYLPRV LDIIRAALPP KDFAHKRQKA MQVDA TVFT CISMLARAMG PGIQQDIKEL LEPMLAVGLS PALTAVLYDL SRQIPQLKKD IQDGLLKMLS LVLMHKPLRH PGMPKG LAH QLASPGLTTL PEASDVGSIT LALRTLGSFE FEGHSLTQFV RHCADHFLNS EHKEIRMEAA RTCSRLLTPS IHLISGH AH VVSQTAVQVV ADVLSKLLVV GITDPDPDIR YCVLASLDER FDAHLAQAEN LQALFVALND QVFEIRELAI CTVGRLSS M NPAFVMPFLR KMLIQILTEL EHSGIGRIKE QSARMLGHLV SNAPRLIRPY MEPILKALIL KLKDPDPDPN PGVINNVLA TIGELAQVSG LEMRKWVDEL FIIIMDMLQD SSLLAKRQVA LWTLGQLVAS TGYVVEPYRK YPTLLEVLLN FLKTEQNQGT RREAIRVLG LLGALDPYKH KVNIGMIDQS RDASAVSLSE SKSSQDSSDY STSEMLVNMG NLPLDEFYPA VSMVALMRIF R DQSLSHHH TMVVQAITFI FKSLGLKCVQ FLPQVMPTFL NVIRVCDGAI REFLFQQLGM LVSFVKSHIR PYMDEIVTLM RE FWVMNTS IQSTIILLIE QIVVALGGEF KLYLPQLIPH MLRVFMHDNS PGRIVSIKLL AAIQLFGANL DDYLHLLLPP IVK LFDAPE APLPSRKAAL ETVDRLTESL DFTDYASRII HPIVRTLDQS PELRSTAMDT LSSLVFQLGK KYQIFIPMVN KVLV RHRIN HQRYDVLICR IVKGYTLADE EEDPLIYQHR MLRSGQGDAL ASGPVETGPM KKLHVSTINL QKAWGAARRV SKDDW LEWL RRLSLELLKD SSSPSLRSCW ALAQAYNPMA RDLFNAAFVS CWSELNEDQQ DELIRSIELA LTSQDIAEVT QTLLNL AEF MEHSDKGPLP LRDDNGIVLL GERAAKCRAY AKALHYKELE FQKGPTPAIL ESLISINNKL QQPEAAAGVL EYAMKHF GE LEIQATWYEK LHEWEDALVA YDKKMDTNKD DPELMLGRMR CLEALGEWGQ LHQQCCEKWT LVNDETQAKM ARMAAAAA W GLGQWDSMEE YTCMIPRDTH DGAFYRAVLA LHQDLFSLAQ QCIDKARDLL DAELTAMAGE SYSRAYGAMV SCHMLSELE EVIQYKLVPE RREIIRQIWW ERLQGCQRIV EDWQKILMVR SLVVSPHEDM RTWLKYASLC GKSGRLALAH KTLVLLLGVD PSRQLDHPL PTVHPQVTYA YMKNMWKSAR KIDAFQHMQH FVQTMQQQAQ HAIATEDQQH KQELHKLMAR CFLKLGEWQL N LQGINEST IPKVLQYYSA ATEHDRSWYK AWHAWAVMNF EAVLHYKHQN QARDEKKKLR HASGANITNA TTAATTAATA TT TASTEGS NSESEAESTE NSPTPSPLQK KVTEDLSKTL LMYTVPAVQG FFRSISLSRG NNLQDTLRVL TLWFDYGHWP DVN EALVEG VKAIQIDTWL QVIPQLIARI DTPRPLVGRL IHQLLTDIGR YHPQALIYPL TVASKSTTTA RHNAANKILK NMCE HSNTL VQQAMMVSEE LIRVAILWHE MWHEGLEEAS RLYFGERNVK GMFEVLEPLH AMMERGPQTL KETSFNQAYG RDLME AQEW CRKYMKSGNV KDLTQAWDLY YHVFRRISKQ LPQLTSLELQ YVSPKLLMCR DLELAVPGTY DPNQPIIRIQ SIAPSL QVI TSKQRPRKLT LMGSNGHEFV FLLKGHEDLR QDERVMQLFG LVNTLLANDP TSLRKNLSIQ RYAVIPLSTN SGLIGWV PH CDTLHALIRD YREKKKILLN IEHRIMLRMA PDYDHLTLMQ KVEVFEHAVN NTAGDDLAKL LWLKSPSSEV WFDRRTNY T RSLAVMSMVG YILGLGDRHP SNLMLDRLSG KILHIDFGDC FEVAMTREKF PEKIPFRLTR MLTNAMEVTG LDGNYRITC HTVMEVLREH KDSVMAVLEA FVYDPLLNWR LMDTNTKGNK RSRTRTDSYS AGQSVEILDG VELGEPAHKK TGTTVPESIH SFIGDGLVK PEALNKKAIQ IINRVRDKLT GRDFSHDDTL DVPTQVELLI KQATSHENLC QCYIGWCPFW UniProtKB: Serine/threonine-protein kinase mTOR |
-Macromolecule #2: Target of rapamycin complex subunit LST8
| Macromolecule | Name: Target of rapamycin complex subunit LST8 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 35.91009 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MNTSPGTVGS DPVILATAGY DHTVRFWQAH SGICTRTVQH QDSQVNALEV TPDRSMIAAA GYQHIRMYDL NSNNPNPIIS YDGVNKNIA SVGFHEDGRW MYTGGEDCTA RIWDLRSRNL QCQRIFQVNA PINCVCLHPN QAELIVGDQS GAIHIWDLKT D HNEQLIPE ...String: MNTSPGTVGS DPVILATAGY DHTVRFWQAH SGICTRTVQH QDSQVNALEV TPDRSMIAAA GYQHIRMYDL NSNNPNPIIS YDGVNKNIA SVGFHEDGRW MYTGGEDCTA RIWDLRSRNL QCQRIFQVNA PINCVCLHPN QAELIVGDQS GAIHIWDLKT D HNEQLIPE PEVSITSAHI DPDASYMAAV NSTGNCYVWN LTGGIGDEVT QLIPKTKIPA HTRYALQCRF SPDSTLLATC SA DQTCKIW RTSNFSLMTE LSIKSGNPGE SSRGWMWGCA FSGDSQYIVT ASSDNLARLW CVETGEIKRE YGGHQKAVVC LAF NDSVLG UniProtKB: Target of rapamycin complex subunit LST8 |
-Macromolecule #3: Regulatory-associated protein of mTOR
| Macromolecule | Name: Regulatory-associated protein of mTOR / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 149.200016 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MESEMLQSPL LGLGEEDEAD LTDWNLPLAF MKKRHCEKIE GSKSLAQSWR MKDRMKTVSV ALVLCLNVGV DPPDVVKTTP CARLECWID PLSMGPQKAL ETIGANLQKQ YENWQPRARY KQSLDPTVDE VKKLCTSLRR NAKEERVLFH YNGHGVPRPT V NGEVWVFN ...String: MESEMLQSPL LGLGEEDEAD LTDWNLPLAF MKKRHCEKIE GSKSLAQSWR MKDRMKTVSV ALVLCLNVGV DPPDVVKTTP CARLECWID PLSMGPQKAL ETIGANLQKQ YENWQPRARY KQSLDPTVDE VKKLCTSLRR NAKEERVLFH YNGHGVPRPT V NGEVWVFN KNYTQYIPLS IYDLQTWMGS PSIFVYDCSN AGLIVKSFKQ FALQREQELE VAAINPNHPL AQMPLPPSMK NC IQLAACE ATELLPMIPD LPADLFTSCL TTPIKIALRW FCMQKCVSLV PGVTLDLIEK IPGRLNDRRT PLGELNWIFT AIT DTIAWN VLPRDLFQKL FRQDLLVASL FRNFLLAERI MRSYNCTPVS SPRLPPTYMH AMWQAWDLAV DICLSQLPTI IEEG TAFRH SPFFAEQLTA FQVWLTMGVE NRNPPEQLPI VLQVLLSQVH RLRALDLLGR FLDLGPWAVS LALSVGIFPY VLKLL QSSA RELRPLLVFI WAKILAVDSS CQADLVKDNG HKYFLSVLAD PYMPAEHRTM TAFILAVIVN SYHTGQEACL QGNLIA ICL EQLNDPHPLL RQWVAICLGR IWQNFDSARW CGVRDSAHEK LYSLLSDPIP EVRCAAVFAL GTFVGNSAER TDHSTTI DH NVAMMLAQLV SDGSPMVRKE LVVALSHLVV QYESNFCTVA LQFIEEEKNY ALPSPATTEG GSLTPVRDSP CTPRLRSV S SYGNIRAVAT ARSLNKSLQN LSLTEESGGA VAFSPGNLST SSSASSTLGS PENEEHILSF ETIDKMRRAS SYSSLNSLI GVSFNSVYTQ IWRVLLHLAA DPYPEVSDVA MKVLNSIAYK ATVNARPQRV LDTSSLTQSA PASPTNKGVH IHQAGGSPPA SSTSSSSLT NDVAKQPVSR DLPSGRPGTT GPAGAQYTPH SHQFPRTRKM FDKGPEQTAD DADDAAGHKS FISATVQTGF C DWSARYFA QPVMKIPEEH DLESQIRKER EWRFLRNSRV RRQAQQVIQK GITRLDDQIF LNRNPGVPSV VKFHPFTPCI AV ADKDSIC FWDWEKGEKL DYFHNGNPRY TRVTAMEYLN GQDCSLLLTA TDDGAIRVWK NFADLEKNPE MVTAWQGLSD MLP TTRGAG MVVDWEQETG LLMSSGDVRI VRIWDTDREM KVQDIPTGAD SCVTSLSCDS HRSLIVAGLG DGSIRVYDRR MALS ECRVM TYREHTAWVV KASLQKRPDG HIVSVSVNGD VRIFDPRMPE SVNVLQIVKG LTALDIHPQA DLIACGSVNQ FTAIY NSSG ELINNIKYYD GFMGQRVGAI SCLAFHPHWP HLAVGSNDYY ISVYSVEKRV R UniProtKB: Regulatory-associated protein of mTOR |
-Macromolecule #4: Eukaryotic translation initiation factor 4E-binding protein 1
| Macromolecule | Name: Eukaryotic translation initiation factor 4E-binding protein 1 type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.592968 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSGGSSCSQT PSRAIPATRR VVLGDGVQLP PGDYSTTPGG TLFSTTPGGT RIIYDRKFLM ECRNSPVTKT PPRDLPTIPG VTSPSSDEP PMEASQSHLR NSPEDKRAGG EESQFEMDI UniProtKB: Eukaryotic translation initiation factor 4E-binding protein 1 |
-Macromolecule #5: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER / type: ligand / ID: 5 / Number of copies: 2 / Formula: ANP |
|---|---|
| Molecular weight | Theoretical: 506.196 Da |
| Chemical component information |  ChemComp-ANP: |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 4 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Initial local fitting was done in Coot and Chimera and then subjected to ROSETTA for model rebuilding. Finally, in Phenix real-space refine was performed for the final model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 122.3 / Target criteria: cross-correlation coefficient |
| Output model |  PDB-8rch: |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)