+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human 20S proteasome assembly structure 1 | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Complex / HYDROLASE | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcerebellar granule cell precursor proliferation / purine ribonucleoside triphosphate binding / Antigen processing: Ub, ATP-independent proteasomal degradation / sperm glycocalyx / proteasome core complex assembly / Regulation of ornithine decarboxylase (ODC) / Proteasome assembly / proteasome core complex / perinuclear theca / Cross-presentation of soluble exogenous antigens (endosomes) ...cerebellar granule cell precursor proliferation / purine ribonucleoside triphosphate binding / Antigen processing: Ub, ATP-independent proteasomal degradation / sperm glycocalyx / proteasome core complex assembly / Regulation of ornithine decarboxylase (ODC) / Proteasome assembly / proteasome core complex / perinuclear theca / Cross-presentation of soluble exogenous antigens (endosomes) / mitotic spindle assembly checkpoint signaling / Somitogenesis / proteasome binding / myofibril / sperm head-tail coupling apparatus / protein folding chaperone complex / immune system process / proteasome endopeptidase complex / chaperone-mediated protein complex assembly / NF-kappaB binding / proteasome core complex, beta-subunit complex / threonine-type endopeptidase activity / proteasome assembly / proteasome core complex, alpha-subunit complex / : / ciliary tip / proteasome complex / Regulation of activated PAK-2p34 by proteasome mediated degradation / sarcomere / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / Asymmetric localization of PCP proteins / Ubiquitin-dependent degradation of Cyclin D / SCF-beta-TrCP mediated degradation of Emi1 / NIK-->noncanonical NF-kB signaling / AUF1 (hnRNP D0) binds and destabilizes mRNA / TNFR2 non-canonical NF-kB pathway / centriole / sperm end piece / negative regulation of inflammatory response to antigenic stimulus / P-body / Assembly of the pre-replicative complex / Vpu mediated degradation of CD4 / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / lipopolysaccharide binding / Dectin-1 mediated noncanonical NF-kB signaling / Degradation of DVL / Degradation of AXIN / Degradation of CRY and PER proteins / Hh mutants are degraded by ERAD / Activation of NF-kappaB in B cells / G2/M Checkpoints / Degradation of GLI1 by the proteasome / Hedgehog ligand biogenesis / Regulation of RUNX3 expression and activity / Autodegradation of the E3 ubiquitin ligase COP1 / Defective CFTR causes cystic fibrosis / GSK3B and BTRC:CUL1-mediated-degradation of NFE2L2 / Negative regulation of NOTCH4 signaling / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / Hedgehog 'on' state / Vif-mediated degradation of APOBEC3G / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / Degradation of GLI2 by the proteasome / GLI3 is processed to GLI3R by the proteasome / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / MAPK6/MAPK4 signaling / Degradation of CDH1 / Degradation of beta-catenin by the destruction complex / Oxygen-dependent proline hydroxylation of Hypoxia-inducible Factor Alpha / CDK-mediated phosphorylation and removal of Cdc6 / ABC-family protein mediated transport / CLEC7A (Dectin-1) signaling / SCF(Skp2)-mediated degradation of p27/p21 / FCERI mediated NF-kB activation / response to virus / Regulation of expression of SLITs and ROBOs / Regulation of PTEN stability and activity / nuclear matrix / Interleukin-1 signaling / Orc1 removal from chromatin / Regulation of RAS by GAPs / Regulation of RUNX2 expression and activity / The role of GTSE1 in G2/M progression after G2 checkpoint / Separation of Sister Chromatids / UCH proteinases / KEAP1-NFE2L2 pathway / Downstream TCR signaling / Antigen processing: Ubiquitination & Proteasome degradation / sperm principal piece / RUNX1 regulates transcription of genes involved in differentiation of HSCs / ER-Phagosome pathway / Neddylation / regulation of inflammatory response / sperm midpiece / secretory granule lumen / endopeptidase activity / molecular adaptor activity / ficolin-1-rich granule lumen / proteasome-mediated ubiquitin-dependent protein catabolic process Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
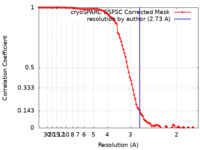
| Method | single particle reconstruction / cryo EM / Resolution: 2.73 Å | |||||||||||||||
 Authors Authors | Schulman BA / Hanna JW / Harper JW / Adolf F / Du J / Rawson SD / Walsh Jr RM / Goodall EA | |||||||||||||||
| Funding support |  Germany, Germany,  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation | Journal: bioRxiv / Year: 2024 Title: Visualizing chaperone-mediated multistep assembly of the human 20S proteasome. Authors: Frank Adolf / Jiale Du / Ellen A Goodall / Richard M Walsh / Shaun Rawson / Susanne von Gronau / J Wade Harper / John Hanna / Brenda A Schulman /   Abstract: Dedicated assembly factors orchestrate stepwise production of many molecular machines, including the 28-subunit proteasome core particle (CP) that mediates protein degradation. Here, we report cryo- ...Dedicated assembly factors orchestrate stepwise production of many molecular machines, including the 28-subunit proteasome core particle (CP) that mediates protein degradation. Here, we report cryo-EM reconstructions of seven recombinant human subcomplexes that visualize all five chaperones and the three active site propeptides across a wide swath of the assembly pathway. Comparison of these chaperone-bound intermediates and a matching mature CP reveals molecular mechanisms determining the order of successive subunit additions, and how proteasome subcomplexes and assembly factors structurally adapt upon progressive subunit incorporation to stabilize intermediates, facilitate the formation of subsequent intermediates, and ultimately rearrange to coordinate proteolytic activation with gated access to active sites. The structural findings reported here explain many previous biochemical and genetic observations. This work establishes a methodologic approach for structural analysis of multiprotein complex assembly intermediates, illuminates specific functions of assembly factors, and reveals conceptual principles underlying human proteasome biogenesis. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18755.map.gz emd_18755.map.gz | 53.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18755-v30.xml emd-18755-v30.xml emd-18755.xml emd-18755.xml | 32.5 KB 32.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18755_fsc.xml emd_18755_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_18755.png emd_18755.png | 49.4 KB | ||
| Masks |  emd_18755_msk_1.map emd_18755_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18755.cif.gz emd-18755.cif.gz | 8 KB | ||
| Others |  emd_18755_additional_1.map.gz emd_18755_additional_1.map.gz emd_18755_half_map_1.map.gz emd_18755_half_map_1.map.gz emd_18755_half_map_2.map.gz emd_18755_half_map_2.map.gz | 88 MB 95.8 MB 95.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18755 http://ftp.pdbj.org/pub/emdb/structures/EMD-18755 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18755 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18755 | HTTPS FTP |
-Related structure data
| Related structure data |  8qyjMC  8qylC  8qymC  8qynC  8qyoC  8qysC  8qz9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18755.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18755.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8512 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18755_msk_1.map emd_18755_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
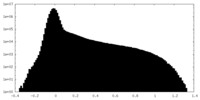
| Density Histograms |
-Additional map: #1
| File | emd_18755_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_18755_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_18755_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Human 20S proteasome assembly intermediate map 1
+Supramolecule #1: Human 20S proteasome assembly intermediate map 1
+Macromolecule #1: Proteasome subunit alpha type-2
+Macromolecule #2: Proteasome subunit alpha type-4
+Macromolecule #3: Proteasome subunit alpha type-7
+Macromolecule #4: Proteasome subunit alpha type-5
+Macromolecule #5: Proteasome subunit alpha type-1
+Macromolecule #6: Proteasome subunit alpha type-3
+Macromolecule #7: Proteasome subunit alpha type-6
+Macromolecule #8: Proteasome maturation protein
+Macromolecule #9: Proteasome assembly chaperone 1
+Macromolecule #10: Proteasome assembly chaperone 2
+Macromolecule #11: Proteasome subunit beta type-7
+Macromolecule #12: Proteasome assembly chaperone 3
+Macromolecule #13: Proteasome assembly chaperone 4
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 66.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)