+ Open data
Open data
- Basic information
Basic information
| Entry | 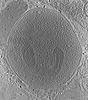 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
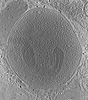
| Title | Stimulated rat synaptosome. | |||||||||
 Map data Map data | stimulated rat synaptosome | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | synapse / neurotransmission / EXOCYTOSIS | |||||||||
| Biological species |  | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Radecke J / Seeger R / Zuber B | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: EMBO Rep / Year: 2023 Journal: EMBO Rep / Year: 2023Title: Morphofunctional changes at the active zone during synaptic vesicle exocytosis. Authors: Julika Radecke / Raphaela Seeger / Anna Kádková / Ulrike Laugks / Amin Khosrozadeh / Kenneth N Goldie / Vladan Lučić / Jakob B Sørensen / Benoît Zuber /     Abstract: Synaptic vesicle (SV) fusion with the plasma membrane (PM) proceeds through intermediate steps that remain poorly resolved. The effect of persistent high or low exocytosis activity on intermediate ...Synaptic vesicle (SV) fusion with the plasma membrane (PM) proceeds through intermediate steps that remain poorly resolved. The effect of persistent high or low exocytosis activity on intermediate steps remains unknown. Using spray-mixing plunge-freezing cryo-electron tomography we observe events following synaptic stimulation at nanometer resolution in near-native samples. Our data suggest that during the stage that immediately follows stimulation, termed early fusion, PM and SV membrane curvature changes to establish a point contact. The next stage-late fusion-shows fusion pore opening and SV collapse. During early fusion, proximal tethered SVs form additional tethers with the PM and increase the inter-SV connector number. In the late-fusion stage, PM-proximal SVs lose their interconnections, allowing them to move toward the PM. Two SNAP-25 mutations, one arresting and one disinhibiting spontaneous release, cause connector loss. The disinhibiting mutation causes loss of membrane-proximal multiple-tethered SVs. Overall, tether formation and connector dissolution are triggered by stimulation and respond to spontaneous fusion rate manipulation. These morphological observations likely correspond to SV transition from one functional pool to another. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18744.map.gz emd_18744.map.gz | 94 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18744-v30.xml emd-18744-v30.xml emd-18744.xml emd-18744.xml | 10.2 KB 10.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_18744.png emd_18744.png | 232.3 KB | ||
| Filedesc metadata |  emd-18744.cif.gz emd-18744.cif.gz | 4.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18744 http://ftp.pdbj.org/pub/emdb/structures/EMD-18744 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18744 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18744 | HTTPS FTP |
-Validation report
| Summary document |  emd_18744_validation.pdf.gz emd_18744_validation.pdf.gz | 396.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18744_full_validation.pdf.gz emd_18744_full_validation.pdf.gz | 396.4 KB | Display | |
| Data in XML |  emd_18744_validation.xml.gz emd_18744_validation.xml.gz | 3.9 KB | Display | |
| Data in CIF |  emd_18744_validation.cif.gz emd_18744_validation.cif.gz | 4.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18744 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18744 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18744 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18744 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18744.map.gz / Format: CCP4 / Size: 126.8 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) Download / File: emd_18744.map.gz / Format: CCP4 / Size: 126.8 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | stimulated rat synaptosome | ||||||||||||||||||||
| Voxel size | X=Y=Z: 22.4 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Stimulated rat synaptosome
| Entire | Name: Stimulated rat synaptosome |
|---|---|
| Components |
|
-Supramolecule #1: Stimulated rat synaptosome
| Supramolecule | Name: Stimulated rat synaptosome / type: cell / ID: 1 / Parent: 0 Details: sprayed with KCl milliseconds before freezing in order to stimulate exocytosis. |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
| |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER | |||||||||||||||||||||||||||
| Sectioning | Other: NO SECTIONING | |||||||||||||||||||||||||||
| Fiducial marker | Manufacturer: AURION Immuno Gold Reagents & Accessories / Diameter: 10 nm |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 20 |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Average electron dose: 1.96 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 12.0 µm / Nominal defocus min: 8.0 µm |
| Sample stage | Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
- Image processing
Image processing
| Final reconstruction | Software - Name:  IMOD / Number images used: 101 IMOD / Number images used: 101 |
|---|
 Movie
Movie Controller
Controller