[English] 日本語
 Yorodumi
Yorodumi- EMDB-18182: Closed conformation of the g-tubulin ring complex nucleating micr... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Closed conformation of the g-tubulin ring complex nucleating microtubules | |||||||||
 Map data Map data | Refined map of the closed gTuRC conformation | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Microtubule / cytoskeleton / g-tubulin ring complex / tubulin / STRUCTURAL PROTEIN | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.39 Å | |||||||||
 Authors Authors | Llorca O / Serna M | |||||||||
| Funding support |  Spain, European Union, 2 items Spain, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2024 Journal: Science / Year: 2024Title: Transition of human γ-tubulin ring complex into a closed conformation during microtubule nucleation. Authors: Cláudia Brito / Marina Serna / Pablo Guerra / Oscar Llorca / Thomas Surrey /  Abstract: Microtubules are essential for intracellular organization and chromosome segregation. They are nucleated by the γ-tubulin ring complex (γTuRC). However, isolated vertebrate γTuRC adopts an open ...Microtubules are essential for intracellular organization and chromosome segregation. They are nucleated by the γ-tubulin ring complex (γTuRC). However, isolated vertebrate γTuRC adopts an open conformation that deviates from the microtubule structure, raising the question of the nucleation mechanism. In this study, we determined cryo-electron microscopy structures of human γTuRC bound to a nascent microtubule. Structural changes of the complex into a closed conformation ensure that γTuRC templates the 13-protofilament microtubules that exist in human cells. Closure is mediated by a latch that interacts with incorporating tubulin, making it part of the closing mechanism. Further rearrangements involve all γTuRC subunits and the removal of the actin-containing luminal bridge. Our proposed mechanism of microtubule nucleation by human γTuRC relies on large-scale structural changes that are likely the target of regulation in cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18182.map.gz emd_18182.map.gz | 205 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18182-v30.xml emd-18182-v30.xml emd-18182.xml emd-18182.xml | 28.1 KB 28.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18182_fsc.xml emd_18182_fsc.xml | 14.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_18182.png emd_18182.png | 131.2 KB | ||
| Masks |  emd_18182_msk_1.map emd_18182_msk_1.map | 259.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18182.cif.gz emd-18182.cif.gz | 9 KB | ||
| Others |  emd_18182_additional_1.map.gz emd_18182_additional_1.map.gz emd_18182_half_map_1.map.gz emd_18182_half_map_1.map.gz emd_18182_half_map_2.map.gz emd_18182_half_map_2.map.gz | 223.7 MB 205.8 MB 205.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18182 http://ftp.pdbj.org/pub/emdb/structures/EMD-18182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18182 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_18182.map.gz / Format: CCP4 / Size: 259.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18182.map.gz / Format: CCP4 / Size: 259.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Refined map of the closed gTuRC conformation | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.36754 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18182_msk_1.map emd_18182_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
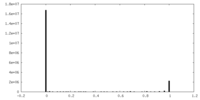
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Sharpened map of the closed gTuRC conformation
| File | emd_18182_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map of the closed gTuRC conformation | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half 2 map of the refined closed gTuRC conformation structure
| File | emd_18182_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half 2 map of the refined closed gTuRC conformation structure | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half 1 map of the refined closed gTuRC conformation structure
| File | emd_18182_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half 1 map of the refined closed gTuRC conformation structure | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : gamma-tubulin ring complex
| Entire | Name: gamma-tubulin ring complex |
|---|---|
| Components |
|
-Supramolecule #1: gamma-tubulin ring complex
| Supramolecule | Name: gamma-tubulin ring complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: GCP2
| Macromolecule | Name: GCP2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSEFRIHHDV NELLSLLRVH GGDGAEVYID LLQKNRTPYV TTTVSAHSAK VKIAEFSRTP EDFLKKYDE LKSKNTRNLD PLVYLLSKLT EDKETLQYLQ QNAKERAELA AAAVGSSTTS I NVPAAASK ISMQELEELR KQLGSVATGS TLQQSLELKR KMLRDKQNKK ...String: MSEFRIHHDV NELLSLLRVH GGDGAEVYID LLQKNRTPYV TTTVSAHSAK VKIAEFSRTP EDFLKKYDE LKSKNTRNLD PLVYLLSKLT EDKETLQYLQ QNAKERAELA AAAVGSSTTS I NVPAAASK ISMQELEELR KQLGSVATGS TLQQSLELKR KMLRDKQNKK NSGQHLPIFP AW VYERPAL IGDFLIGAGI STDTALPIGT LPLASQESAV VEDLLYVLVG VDGRYVSAQP LAG RQSRTF LVDPNLDLSI RELVHRILPV AASYSAVTRF IEEKSSFEYG QVNHALAAAM RTLV KEHLI LVSQLEQLHR QGLLSLQKLW FYIQPAMRTM DILASLATSV DKGECLGGST LSLLH DRSF SYTGDSQAQE LCLYLTKAAS APYFEVLEKW IYRGIIHDPY SEFMVEEHEL RKERIQ EDY NDKYWDQRYT IVQQQIPSFL QKMADKILST GKYLNVVREC GHDVTCPVAK EIIYTLK ER AYVEQIEKAF NYASKVLLDF LMEEKELVAH LRSIKRYFLM DQGDFFVHFM DLAEEELR K PVEDITPPRL EALLELALRM STANTDPFKD DLKIDLMPHD LITQLLRVLA IETKQEKAM AHADPTELAL SGLEAFSFDY IVKWPLSLII NRKALTRYQM LFRHMFYCKH VERQLCSVWI SNKTAKQHS LHSAQWFAGA FTLRQRMLNF VQNIQYYMMF EVMEPTWHIL EKNLKSASNI D DVLGHHTG FLDTCLKDCM LTNPELLKVF SKLMSVCVMF TNCMQKFTQS MKLDGELGGQ TL EHSTVLG LPAGAEERAR KELARKHLAE HADTVQLVSG FEATINKFDK NFSAHLLDLL ARL SIYSTS DCEHGMASVI SRLDFNGFYT ERLERLSAER SQKATPQVPV LRGPPAPAPR VAVT AQ |
-Macromolecule #2: GCP3
| Macromolecule | Name: GCP3 / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MATPDQKSPN VLLQNLCCRI LGRSEADVAQ QFQYAVRVIG SNFAPTVERD EFLVAEKIKK ELIRQRREA DAALFSELHR KLHSQGVLKN KWSILYLLLS LSEDPRRQPS KVSSYATLFA Q ALPRDAHS TPYYYARPQT LPLSYQDRSA QSAQSSGSVG SSGISSIGLC ...String: MATPDQKSPN VLLQNLCCRI LGRSEADVAQ QFQYAVRVIG SNFAPTVERD EFLVAEKIKK ELIRQRREA DAALFSELHR KLHSQGVLKN KWSILYLLLS LSEDPRRQPS KVSSYATLFA Q ALPRDAHS TPYYYARPQT LPLSYQDRSA QSAQSSGSVG SSGISSIGLC ALSGPAPAPQ SL LPGQSNQ APGVGDCLRQ QLGSRLAWTL TANQPSSQAT TSKGVPSAVS RNMTRSRREG DTG GTMEIT EAALVRDILY VFQGIDGKNI KMNNTENCYK VEGKANLSRS LRDTAVRLSE LGWL HNKIR RYTDQRSLDR SFGLVGQSFC AALHQELREY YRLLSVLHSQ LQLEDDQGVN LGLES SLTL RRLLVWTYDP KIRLKTLAAL VDHCQGRKGG ELASAVHAYT KTGDPYMRSL VQHILS LVS HPVLSFLYRW IYDGELEDTY HEFFVASDPT VKTDRLWHDK YTLRKSMIPS FMTMDQS RK VLLIGKSINF LHQVCHDQTP TTKMIAVTKS AESPQDAADL FTDLENAFQG KIDAAYFE T SKYLLDVLNK KYSLLDHMQA MRRYLLLGQG DFIRHLMDLL KPELVRPATT LYQHNLTGI LETAVRATNA QFDSPEILRR LDVRLLEVSP GDTGWDVFSL DYHVDGPIAT VFTRECMSHY LRVFNFLWR AKRMEYILTD IRKGHMCNAK LLRNMPEFSG VLHQCHILAS EMVHFIHQMQ Y YITFEVLE CSWDELWNKV QQAQDLDHII AAHEVFLDTI ISRCLLDSDS RALLNQLRAV FD QIIELQN AQDAIYRAAL EELQRRLQFE EKKKQREIEG QWGVTAAEEE EENKRIGEFK ESI PKMCSQ LRILTHFYQG IVQQFLVLLT TSSDESLRFL SFRLDFNEHY KAREPRLRVS LGTR GRRSS HT |
-Macromolecule #3: GCP4
| Macromolecule | Name: GCP4 / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MIHELLLALS GYPGSIFTWN KRSGLQVSQD FPFLHPSETS VLNRLCRLGT DYIRFTEFIE QYTGHVQQQ DHHPSQQGQG GLHGIYLRAF CTGLDSVLQP YRQALLDLEQ EFLGDPHLSI S HVNYFLDQ FQLLFPSVMV VVEQIKSQKI HGCQILETVY KHSCGGLPPV ...String: MIHELLLALS GYPGSIFTWN KRSGLQVSQD FPFLHPSETS VLNRLCRLGT DYIRFTEFIE QYTGHVQQQ DHHPSQQGQG GLHGIYLRAF CTGLDSVLQP YRQALLDLEQ EFLGDPHLSI S HVNYFLDQ FQLLFPSVMV VVEQIKSQKI HGCQILETVY KHSCGGLPPV RSALEKILAV CH GVMYKQL SAWMLHGLLL DQHEEFFIKQ GPSSGNVSAQ PEEDEEDLGI GGLTGKQLRE LQD LRLIEE ENMLAPSLKQ FSLRVEILPS YIPVRVAEKI LFVGESVQMF ENQNVNLTRK GSIL KNQED TFAAELHRLK QQPLFSLVDF EQVVDRIRST VAEHLWKLMV EESDLLGQLK IIKDF YLLG RGELFQAFID TAQHMLKTPP TAVTEHDVNV AFQQSAHKVL LDDDNLLPLL HLTIEY HGK EHKADATQAR EGPSRETSPR EAPASGWAAL GLSYKVQWPL HILFTPAVLE KYNVVFK YL LSVRRVQAEL QHCWALQMQR KHLKSNQTDA IKWRLRNHMA FLVDNLQYYL QVDVLESQ F SQLLHQINST RDFESIRLAH DHFLSNLLAQ SFILLKPVFH CLNEILDLCH SFCSLVSQN LGPLDERGAA QLSILVKGFS RQSSLLFKIL SSVRNHQINS DLAQLLLRLD YNKYYTQAGG TLGSFGM |
-Macromolecule #4: GCP5
| Macromolecule | Name: GCP5 / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MARHGPPWSR LDAQQERDVR ELVRGVAGLQ DEADPNFQLA LNFAWSNFRF HRFLDVNSHK IEKTIEGIY EKFVIHSDLS KAASWKRLTE EFLNAPLPSI KEIKTDAHYS ILSLLLCLSD S PSNSSYVE TPRNKEVEKK DDFDWGKYLM EDEEMDIGPY MDTPNWSEES ...String: MARHGPPWSR LDAQQERDVR ELVRGVAGLQ DEADPNFQLA LNFAWSNFRF HRFLDVNSHK IEKTIEGIY EKFVIHSDLS KAASWKRLTE EFLNAPLPSI KEIKTDAHYS ILSLLLCLSD S PSNSSYVE TPRNKEVEKK DDFDWGKYLM EDEEMDIGPY MDTPNWSEES EEENDQQPLS RE DSGIQVD RTPLEEQDQN RKLDPCISWK DEPDDRSWLE HHVVHQYWTA RPSQFPHSLH LHS NLAAVW DQHLYSSDPL YVPDDRVLVT ETQVIRETLW LLSGVKKLFI FQLIDGKVTV RNNI IVTHL THSCLRSVLE QIAAYGQVVF RLQEFIDEVM GHSSESMLPG SGSVPKKSTE APFRT YQAF MWALYKYFIS FKEELAEIEK CIINNDTTIT LAIVVDKLAP RLSQLKVLHK VFSTGV AEV PPDTRNVVRA SHLLNTLYKA ILEYDNVGEA SEQTVSLLFS LWVETVRPYL QTVDEWI VH GHLWDGAREF IIQRNKNVPV NHRDFWYATY TLYSVSEKTE NEEKMSDNAS ASSGSDQG P SSRQHTMVSF LKPVLKQIIM AGKSMQLLKN LQCAESTTCQ AGARDAERKS LYTLFLESV QSRLRHGEDS TPQVLTEQQA TKENLMKMQS IAESHLELDD VHDPLLAINF ARMYLEQSDF HEKFAGGDV CVDRSSESVT CQTFELTLRS CLYPHIDKQY LDCCGNLMQT LKKDYRLVEY L QAMRNFFL MEGGDTMYDF YTSIFDKIRE KETWQNVSFL NVQLQEAVGQ RYPEDSSRLS IS FENVDTA KKKLPVHILD GLTLSYKVPW PVDIVISLEC QKIYNQVFLL LLQIKWAKYS LDV LLFGEL VSTAEKPRLK EGLIHEQDTV AQFGPQKEPV RQQIHRMFLL RVKLMHFVNS LHNY IMTRI LHSTGLEFQH QVEEAKDLDQ LIKIHYRYLS TIHDRCLLRE KVSFVKEAIM KVLNL ALMF ADGWQAGLGT WRMESIEKME SDFKNCHMFL VTILNKAVCR GSFPHLESLA LSLMAG MEQ S |
-Macromolecule #5: GCP6
| Macromolecule | Name: GCP6 / type: protein_or_peptide / ID: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MASITQLFDD LCEALLPAAK THLGQRSVNR KRAKRSLKKV AYNALFTNLF QDETQQLQPD MSKLPARNK ILMLSFDLRV GGLGPKADRL EELVEELEAA PCCPLLEVGS VLDLLVQLAG S GPPQVLPR KRDYFLNNKH VGRNVPYSGY DCDDLSVFEM DVQSLISREE ...String: MASITQLFDD LCEALLPAAK THLGQRSVNR KRAKRSLKKV AYNALFTNLF QDETQQLQPD MSKLPARNK ILMLSFDLRV GGLGPKADRL EELVEELEAA PCCPLLEVGS VLDLLVQLAG S GPPQVLPR KRDYFLNNKH VGRNVPYSGY DCDDLSVFEM DVQSLISREE CLCHSMIQET LQ VMEAAPG TGLPTVGLFS FGDPCGDRFE RDTRVSLFGA LVHSRTYDMD VRLGLPPVPD NAD LSGLAI KVPPSVDQWE DEGFQSASNL TPDSQSEPSV TPDVDLWEAA LTYEASKRRC WERV GCPPG HREEPYLTEA GRDAFDKFCR LHQGELQLLA GGVLQAPQPV LVKECELVKD VLNVL IGVV SATFSLCQPA QAFVVKRGVH VSGASPESIS SLLSEVAEYG TCYTRLSHFS LQPVLD SLY SKGLVFQAFT SGLRRYLQYY RACVLSTPPT LSLLTIGFLF KKLGRQLRYL AELCGVG AV LPGTCGGGPR AAFPTGVKLL SYLYQEALHN CSNEHYPVLL SLLKTSCEPY TRFIHDWV Y SGVFRDAYGE FMIQVNHEYL SFRDKLYWTH GYVLISKEVE DCVPVFLKHI AHDIYVCGK TINLLKLCCP RHYLCWSDVP VPRISVIFSL EELKEIEKDC AVYVGRMERV ARHSSVSKEE KELRMEIAK QELIAHAREA ASRVLSALSD RQMSERMALD ARKREQFQRL KEQFVKDQER R QAARQEEL DDDFSYAREL RDRERRLKSL EEELERKARQ ALVDHYSKLS AEAARREQKA LW RIQRHRL ESARLRFLLE DEKHIQEMLK AVSEAHQPQE PPDVLLSVHP QVTSPGPEHP EGG QGCDSG SAEQHSPAWD GWNRPGLLTP QPLKPLAVGA GGRGLQQAEG ARPFSDSLSI GDFL PVGPG AEPSVQTGMV PLLEVALQTI NLDLPPSAPG EAPAAASTQP SRPQEYDFST VLRPA VATS PAPGPLQAAE CSLGSSGLQL WEDSCGKMDA CGSASRETLL PSHPPRRAAL EEGSSQ PTE RLFGQVSGGG LPTGDYASEI APTRPRWNTH GHVSDASIRV GENVSDVAPT QPRWNTH GH VSNASISLGE SVSDVAPTRP RWNIHGHVSN ASIRVGENVS DVAPTRPRWN THGHVSNA S IRVGENVSDV APTRPRWNTH GHVSDASISL GESVSDMAPA RPRWNTHGHV SDASISLGE SVSDMAPTRP RWNTHGHVSD TSIRVGENVS DVAPIRSRCN THGHVSDASI SLGEPVSDVV STRPRWNTH VPIPPPHMVL GALSPEAEPN TPRPQQSPPG HTSQSALSLG AQSTVLDCGP R LPVEVGPS LSSPSSGCGE GSISVGENVS DVAPTQPWWP NTPGDSVSEE LGPGRSGDTE DL SPNWPLN SQEDTAAQSS PGRGEEAEAS AAEAQGGEQA YLAGLAGQYH LERYPDSYES MSE PPIAHL LRPVLPRAFA FPVDPQVQSA ADETAVQLSE LLTLPVLMKR SITAPLAAHI SLVN KAAVD YFFVELHLEA HYEALRHFLL MEDGEFAQSL SDLLFEKLGA GQTPGELLNP LVLNS VLSK ALQCSLHGDT PHASNLSLAL KYLPEVFAPN APDVLSCLEL RYKVDWPLNI VITEGC VSK YSGVFSFLLQ LKLMMWALKD VCFHLKRTAL LSHMAGSVQF RQLQLFKHEM QHFVKVI QG YIANQILHVT WCEFRARLAT VGDLEEIQRA HAEYLHKAVF RGLLTEKAAP VMNVIHSI F SLVLKFRSQL ISQAWGPPGG PRGAEHPNFA LMQQSYNTFK YYSHFLFKVV TKLVNRGYQ PHLEDFLLRI NFNNYYQDA |
-Macromolecule #6: gamma-tubulin
| Macromolecule | Name: gamma-tubulin / type: protein_or_peptide / ID: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPREIITLQL GQCGNQIGFE FWKQLCAEHG ISPEGIVEEF ATEGTDRKDV FFYQADDEHY IPRAVLLDL EPRVIHSILN SPYAKLYNPE NIYLSEHGGG AGNNWASGFS QGEKIHEDIF D IIDREADG SDSLEGFVLC HSIAGGTGSG LGSYLLERLN DRYPKKLVQT ...String: MPREIITLQL GQCGNQIGFE FWKQLCAEHG ISPEGIVEEF ATEGTDRKDV FFYQADDEHY IPRAVLLDL EPRVIHSILN SPYAKLYNPE NIYLSEHGGG AGNNWASGFS QGEKIHEDIF D IIDREADG SDSLEGFVLC HSIAGGTGSG LGSYLLERLN DRYPKKLVQT YSVFPNQDEM SD VVVQPYN SLLTLKRLTQ NADCVVVLDN TALNRIATDR LHIQNPSFSQ INQLVSTIMS AST TTLRYP GYMNNDLIGL IASLIPTPRL HFLMTGYTPL TTDQSVASVR KTTVLDVMRR LLQP KNVMV STGRDRQTNH CYIAILNIIQ GEVDPTQVHK SLQRIRERKL ANFIPWGPAS IQVAL SRKS PYLPSAHRVS GLMMANHTSI SSLFERTCRQ YDKLRKREAF LEQFRKEDMF KDNFDE MDT SREIVQQLID EYHAATRPDY ISWGTQEQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.9 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 310 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 46276 / Average exposure time: 1.3 sec. / Average electron dose: 46.276 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)