+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Early closed conformation of the g-tubulin ring complex | |||||||||
 Map data Map data | Refined map of the early closed gTuRC-MT conformation | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Microtubule / cytoskeleton / g-tubulin ring complex / tubulin / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmicrotubule nucleator activity / polar microtubule / gamma-tubulin complex / gamma-tubulin ring complex / mitotic spindle microtubule / meiotic spindle organization / microtubule nucleation / gamma-tubulin binding / non-motile cilium / pericentriolar material ...microtubule nucleator activity / polar microtubule / gamma-tubulin complex / gamma-tubulin ring complex / mitotic spindle microtubule / meiotic spindle organization / microtubule nucleation / gamma-tubulin binding / non-motile cilium / pericentriolar material / ciliary transition zone / cell leading edge / microtubule organizing center / mitotic sister chromatid segregation / single fertilization / cytoplasmic microtubule / spindle assembly / cytoplasmic microtubule organization / Loss of Nlp from mitotic centrosomes / Loss of proteins required for interphase microtubule organization from the centrosome / Recruitment of mitotic centrosome proteins and complexes / Recruitment of NuMA to mitotic centrosomes / Anchoring of the basal body to the plasma membrane / AURKA Activation by TPX2 / mitotic spindle organization / condensed nuclear chromosome / meiotic cell cycle / centriole / brain development / structural constituent of cytoskeleton / recycling endosome / microtubule cytoskeleton organization / spindle / neuron migration / spindle pole / apical part of cell / Regulation of PLK1 Activity at G2/M Transition / mitotic cell cycle / microtubule cytoskeleton / protein-containing complex assembly / microtubule binding / microtubule / neuron projection / cilium / ciliary basal body / centrosome / GTP binding / structural molecule activity / nucleoplasm / membrane / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
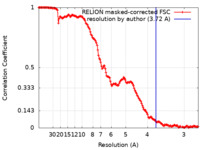
| Method | single particle reconstruction / cryo EM / Resolution: 3.72 Å | |||||||||
 Authors Authors | Llorca O / Serna M / Fernandez-Leiro R | |||||||||
| Funding support |  Spain, European Union, 2 items Spain, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2024 Journal: Science / Year: 2024Title: Transition of human γ-tubulin ring complex into a closed conformation during microtubule nucleation. Authors: Cláudia Brito / Marina Serna / Pablo Guerra / Oscar Llorca / Thomas Surrey /  Abstract: Microtubules are essential for intracellular organization and chromosome segregation. They are nucleated by the γ-tubulin ring complex (γTuRC). However, isolated vertebrate γTuRC adopts an open ...Microtubules are essential for intracellular organization and chromosome segregation. They are nucleated by the γ-tubulin ring complex (γTuRC). However, isolated vertebrate γTuRC adopts an open conformation that deviates from the microtubule structure, raising the question of the nucleation mechanism. In this study, we determined cryo-electron microscopy structures of human γTuRC bound to a nascent microtubule. Structural changes of the complex into a closed conformation ensure that γTuRC templates the 13-protofilament microtubules that exist in human cells. Closure is mediated by a latch that interacts with incorporating tubulin, making it part of the closing mechanism. Further rearrangements involve all γTuRC subunits and the removal of the actin-containing luminal bridge. Our proposed mechanism of microtubule nucleation by human γTuRC relies on large-scale structural changes that are likely the target of regulation in cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18181.map.gz emd_18181.map.gz | 242 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18181-v30.xml emd-18181-v30.xml emd-18181.xml emd-18181.xml | 46.9 KB 46.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18181_fsc.xml emd_18181_fsc.xml | 15.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_18181.png emd_18181.png | 131 KB | ||
| Masks |  emd_18181_msk_1.map emd_18181_msk_1.map | 303.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18181.cif.gz emd-18181.cif.gz | 10.2 KB | ||
| Others |  emd_18181_additional_1.map.gz emd_18181_additional_1.map.gz emd_18181_additional_2.map.gz emd_18181_additional_2.map.gz emd_18181_additional_3.map.gz emd_18181_additional_3.map.gz emd_18181_additional_4.map.gz emd_18181_additional_4.map.gz emd_18181_additional_5.map.gz emd_18181_additional_5.map.gz emd_18181_additional_6.map.gz emd_18181_additional_6.map.gz emd_18181_additional_7.map.gz emd_18181_additional_7.map.gz emd_18181_half_map_1.map.gz emd_18181_half_map_1.map.gz emd_18181_half_map_2.map.gz emd_18181_half_map_2.map.gz | 284 MB 269.7 MB 284 MB 8.9 MB 8.9 MB 8.9 MB 8.9 MB 242.5 MB 242.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18181 http://ftp.pdbj.org/pub/emdb/structures/EMD-18181 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18181 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18181 | HTTPS FTP |
-Related structure data
| Related structure data |  8q62MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18181.map.gz / Format: CCP4 / Size: 303.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18181.map.gz / Format: CCP4 / Size: 303.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Refined map of the early closed gTuRC-MT conformation | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.36754 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
+Mask #1
+Additional map: Refined and sharpened map of the early closed gTuRC-MT conformation
+Additional map: Refined and sharpened map of the early closed...
+Additional map: Refined and sharpened map of the early closed...
+Additional map: CryoDRGN volume 6
+Additional map: CryoDRGN volume 5
+Additional map: CryoDRGN volume 3
+Additional map: CryoDRGN volume 1
+Half map: Half map 1 of the refined map of...
+Half map: Half map 2 of the refined map of...
- Sample components
Sample components
-Entire : gamma-tubulin ring complex
| Entire | Name: gamma-tubulin ring complex |
|---|---|
| Components |
|
-Supramolecule #1: gamma-tubulin ring complex
| Supramolecule | Name: gamma-tubulin ring complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #2-#4, #1, #6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Tubulin gamma-1 chain
| Macromolecule | Name: Tubulin gamma-1 chain / type: protein_or_peptide / ID: 1 / Number of copies: 14 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 51.22777 KDa |
| Sequence | String: MPREIITLQL GQCGNQIGFE FWKQLCAEHG ISPEGIVEEF ATEGTDRKDV FFYQADDEHY IPRAVLLDLE PRVIHSILNS PYAKLYNPE NIYLSEHGGG AGNNWASGFS QGEKIHEDIF DIIDREADGS DSLEGFVLCH SIAGGTGSGL GSYLLERLND R YPKKLVQT ...String: MPREIITLQL GQCGNQIGFE FWKQLCAEHG ISPEGIVEEF ATEGTDRKDV FFYQADDEHY IPRAVLLDLE PRVIHSILNS PYAKLYNPE NIYLSEHGGG AGNNWASGFS QGEKIHEDIF DIIDREADGS DSLEGFVLCH SIAGGTGSGL GSYLLERLND R YPKKLVQT YSVFPNQDEM SDVVVQPYNS LLTLKRLTQN ADCVVVLDNT ALNRIATDRL HIQNPSFSQI NQLVSTIMSA ST TTLRYPG YMNNDLIGLI ASLIPTPRLH FLMTGYTPLT TDQSVASVRK TTVLDVMRRL LQPKNVMVST GRDRQTNHCY IAI LNIIQG EVDPTQVHKS LQRIRERKLA NFIPWGPASI QVALSRKSPY LPSAHRVSGL MMANHTSISS LFERTCRQYD KLRK REAFL EQFRKEDMFK DNFDEMDTSR EIVQQLIDEY HAATRPDYIS WGTQEQ UniProtKB: Tubulin gamma-1 chain |
-Macromolecule #2: Gamma-tubulin complex component 5
| Macromolecule | Name: Gamma-tubulin complex component 5 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 118.467547 KDa |
| Sequence | String: MARHGPPWSR LDAQQERDVR ELVRGVAGLQ DEADPNFQLA LNFAWSNFRF HRFLDVNSHK IEKTIEGIYE KFVIHSDLSK AASWKRLTE EFLNAPLPSI KEIKTDAHYS ILSLLLCLSD SPSNSSYVET PRNKEVEKKD DFDWGKYLME DEEMDIGPYM D TPNWSEES ...String: MARHGPPWSR LDAQQERDVR ELVRGVAGLQ DEADPNFQLA LNFAWSNFRF HRFLDVNSHK IEKTIEGIYE KFVIHSDLSK AASWKRLTE EFLNAPLPSI KEIKTDAHYS ILSLLLCLSD SPSNSSYVET PRNKEVEKKD DFDWGKYLME DEEMDIGPYM D TPNWSEES EEENDQQPLS REDSGIQVDR TPLEEQDQNR KLDPCISWKD EPDDRSWLEH HVVHQYWTAR PSQFPHSLHL HS NLAAVWD QHLYSSDPLY VPDDRVLVTE TQVIRETLWL LSGVKKLFIF QLIDGKVTVR NNIIVTHLTH SCLRSVLEQI AAY GQVVFR LQEFIDEVMG HSSESMLPGS GSVPKKSTEA PFRTYQAFMW ALYKYFISFK EELAEIEKCI INNDTTITLA IVVD KLAPR LSQLKVLHKV FSTGVAEVPP DTRNVVRASH LLNTLYKAIL EYDNVGEASE QTVSLLFSLW VETVRPYLQT VDEWI VHGH LWDGAREFII QRNKNVPVNH RDFWYATYTL YSVSEKTENE EKMSDNASAS SGSDQGPSSR QHTMVSFLKP VLKQII MAG KSMQLLKNLQ CAESTTCQAG ARDAERKSLY TLFLESVQSR LRHGEDSTPQ VLTEQQATKE NLMKMQSIAE SHLELDD VH DPLLAINFAR MYLEQSDFHE KFAGGDVCVD RSSESVTCQT FELTLRSCLY PHIDKQYLDC CGNLMQTLKK DYRLVEYL Q AMRNFFLMEG GDTMYDFYTS IFDKIREKET WQNVSFLNVQ LQEAVGQRYP EDSSRLSISF ENVDTAKKKL PVHILDGLT LSYKVPWPVD IVISLECQKI YNQVFLLLLQ IKWAKYSLDV LLFGELVSTA EKPRLKEGLI HEQDTVAQFG PQKEPVRQQI HRMFLLRVK LMHFVNSLHN YIMTRILHST GLEFQHQVEE AKDLDQLIKI HYRYLSTIHD RCLLREKVSF VKEAIMKVLN L ALMFADGW QAGLGTWRME SIEKMESDFK NCHMFLVTIL NKAVCRGSFP HLESLALSLM AGMEQS UniProtKB: Gamma-tubulin complex component 5 |
-Macromolecule #3: Gamma-tubulin complex component 4
| Macromolecule | Name: Gamma-tubulin complex component 4 / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 76.179969 KDa |
| Sequence | String: MIHELLLALS GYPGSIFTWN KRSGLQVSQD FPFLHPSETS VLNRLCRLGT DYIRFTEFIE QYTGHVQQQD HHPSQQGQGG LHGIYLRAF CTGLDSVLQP YRQALLDLEQ EFLGDPHLSI SHVNYFLDQF QLLFPSVMVV VEQIKSQKIH GCQILETVYK H SCGGLPPV ...String: MIHELLLALS GYPGSIFTWN KRSGLQVSQD FPFLHPSETS VLNRLCRLGT DYIRFTEFIE QYTGHVQQQD HHPSQQGQGG LHGIYLRAF CTGLDSVLQP YRQALLDLEQ EFLGDPHLSI SHVNYFLDQF QLLFPSVMVV VEQIKSQKIH GCQILETVYK H SCGGLPPV RSALEKILAV CHGVMYKQLS AWMLHGLLLD QHEEFFIKQG PSSGNVSAQP EEDEEDLGIG GLTGKQLREL QD LRLIEEE NMLAPSLKQF SLRVEILPSY IPVRVAEKIL FVGESVQMFE NQNVNLTRKG SILKNQEDTF AAELHRLKQQ PLF SLVDFE QVVDRIRSTV AEHLWKLMVE ESDLLGQLKI IKDFYLLGRG ELFQAFIDTA QHMLKTPPTA VTEHDVNVAF QQSA HKVLL DDDNLLPLLH LTIEYHGKEH KADATQAREG PSRETSPREA PASGWAALGL SYKVQWPLHI LFTPAVLEKY NVVFK YLLS VRRVQAELQH CWALQMQRKH LKSNQTDAIK WRLRNHMAFL VDNLQYYLQV DVLESQFSQL LHQINSTRDF ESIRLA HDH FLSNLLAQSF ILLKPVFHCL NEILDLCHSF CSLVSQNLGP LDERGAAQLS ILVKGFSRQS SLLFKILSSV RNHQINS DL AQLLLRLDYN KYYTQAGGTL GSFGM UniProtKB: Gamma-tubulin complex component 4 |
-Macromolecule #4: Gamma-tubulin complex component 3
| Macromolecule | Name: Gamma-tubulin complex component 3 / type: protein_or_peptide / ID: 4 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 103.710102 KDa |
| Sequence | String: MATPDQKSPN VLLQNLCCRI LGRSEADVAQ QFQYAVRVIG SNFAPTVERD EFLVAEKIKK ELIRQRREAD AALFSELHRK LHSQGVLKN KWSILYLLLS LSEDPRRQPS KVSSYATLFA QALPRDAHST PYYYARPQTL PLSYQDRSAQ SAQSSGSVGS S GISSIGLC ...String: MATPDQKSPN VLLQNLCCRI LGRSEADVAQ QFQYAVRVIG SNFAPTVERD EFLVAEKIKK ELIRQRREAD AALFSELHRK LHSQGVLKN KWSILYLLLS LSEDPRRQPS KVSSYATLFA QALPRDAHST PYYYARPQTL PLSYQDRSAQ SAQSSGSVGS S GISSIGLC ALSGPAPAPQ SLLPGQSNQA PGVGDCLRQQ LGSRLAWTLT ANQPSSQATT SKGVPSAVSR NMTRSRREGD TG GTMEITE AALVRDILYV FQGIDGKNIK MNNTENCYKV EGKANLSRSL RDTAVRLSEL GWLHNKIRRY TDQRSLDRSF GLV GQSFCA ALHQELREYY RLLSVLHSQL QLEDDQGVNL GLESSLTLRR LLVWTYDPKI RLKTLAALVD HCQGRKGGEL ASAV HAYTK TGDPYMRSLV QHILSLVSHP VLSFLYRWIY DGELEDTYHE FFVASDPTVK TDRLWHDKYT LRKSMIPSFM TMDQS RKVL LIGKSINFLH QVCHDQTPTT KMIAVTKSAE SPQDAADLFT DLENAFQGKI DAAYFETSKY LLDVLNKKYS LLDHMQ AMR RYLLLGQGDF IRHLMDLLKP ELVRPATTLY QHNLTGILET AVRATNAQFD SPEILRRLDV RLLEVSPGDT GWDVFSL DY HVDGPIATVF TRECMSHYLR VFNFLWRAKR MEYILTDIRK GHMCNAKLLR NMPEFSGVLH QCHILASEMV HFIHQMQY Y ITFEVLECSW DELWNKVQQA QDLDHIIAAH EVFLDTIISR CLLDSDSRAL LNQLRAVFDQ IIELQNAQDA IYRAALEEL QRRLQFEEKK KQREIEGQWG VTAAEEEEEN KRIGEFKESI PKMCSQLRIL THFYQGIVQQ FLVLLTTSSD ESLRFLSFRL DFNEHYKAR EPRLRVSLGT RGRRSSHT UniProtKB: Gamma-tubulin complex component 3 |
-Macromolecule #5: Gamma-tubulin complex component 2
| Macromolecule | Name: Gamma-tubulin complex component 2 / type: protein_or_peptide / ID: 5 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 102.666953 KDa |
| Sequence | String: MSEFRIHHDV NELLSLLRVH GGDGAEVYID LLQKNRTPYV TTTVSAHSAK VKIAEFSRTP EDFLKKYDEL KSKNTRNLDP LVYLLSKLT EDKETLQYLQ QNAKERAELA AAAVGSSTTS INVPAAASKI SMQELEELRK QLGSVATGST LQQSLELKRK M LRDKQNKK ...String: MSEFRIHHDV NELLSLLRVH GGDGAEVYID LLQKNRTPYV TTTVSAHSAK VKIAEFSRTP EDFLKKYDEL KSKNTRNLDP LVYLLSKLT EDKETLQYLQ QNAKERAELA AAAVGSSTTS INVPAAASKI SMQELEELRK QLGSVATGST LQQSLELKRK M LRDKQNKK NSGQHLPIFP AWVYERPALI GDFLIGAGIS TDTALPIGTL PLASQESAVV EDLLYVLVGV DGRYVSAQPL AG RQSRTFL VDPNLDLSIR ELVHRILPVA ASYSAVTRFI EEKSSFEYGQ VNHALAAAMR TLVKEHLILV SQLEQLHRQG LLS LQKLWF YIQPAMRTMD ILASLATSVD KGECLGGSTL SLLHDRSFSY TGDSQAQELC LYLTKAASAP YFEVLEKWIY RGII HDPYS EFMVEEHELR KERIQEDYND KYWDQRYTIV QQQIPSFLQK MADKILSTGK YLNVVRECGH DVTCPVAKEI IYTLK ERAY VEQIEKAFNY ASKVLLDFLM EEKELVAHLR SIKRYFLMDQ GDFFVHFMDL AEEELRKPVE DITPPRLEAL LELALR MST ANTDPFKDDL KIDLMPHDLI TQLLRVLAIE TKQEKAMAHA DPTELALSGL EAFSFDYIVK WPLSLIINRK ALTRYQM LF RHMFYCKHVE RQLCSVWISN KTAKQHSLHS AQWFAGAFTL RQRMLNFVQN IQYYMMFEVM EPTWHILEKN LKSASNID D VLGHHTGFLD TCLKDCMLTN PELLKVFSKL MSVCVMFTNC MQKFTQSMKL DGELGGQTLE HSTVLGLPAG AEERARKEL ARKHLAEHAD TVQLVSGFEA TINKFDKNFS AHLLDLLARL SIYSTSDCEH GMASVISRLD FNGFYTERLE RLSAERSQKA TPQVPVLRG PPAPAPRVAV TAQ UniProtKB: Gamma-tubulin complex component 2 |
-Macromolecule #6: Gamma-tubulin complex component 6
| Macromolecule | Name: Gamma-tubulin complex component 6 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 200.733641 KDa |
| Sequence | String: MASITQLFDD LCEALLPAAK THLGQRSVNR KRAKRSLKKV AYNALFTNLF QDETQQLQPD MSKLPARNKI LMLSFDLRVG GLGPKADRL EELVEELEAA PCCPLLEVGS VLDLLVQLAG SGPPQVLPRK RDYFLNNKHV GRNVPYSGYD CDDLSVFEMD V QSLISREE ...String: MASITQLFDD LCEALLPAAK THLGQRSVNR KRAKRSLKKV AYNALFTNLF QDETQQLQPD MSKLPARNKI LMLSFDLRVG GLGPKADRL EELVEELEAA PCCPLLEVGS VLDLLVQLAG SGPPQVLPRK RDYFLNNKHV GRNVPYSGYD CDDLSVFEMD V QSLISREE CLCHSMIQET LQVMEAAPGT GLPTVGLFSF GDPCGDRFER DTRVSLFGAL VHSRTYDMDV RLGLPPVPDN AD LSGLAIK VPPSVDQWED EGFQSASNLT PDSQSEPSVT PDVDLWEAAL TYEASKRRCW ERVGCPPGHR EEPYLTEAGR DAF DKFCRL HQGELQLLAG GVLQAPQPVL VKECELVKDV LNVLIGVVSA TFSLCQPAQA FVVKRGVHVS GASPESISSL LSEV AEYGT CYTRLSHFSL QPVLDSLYSK GLVFQAFTSG LRRYLQYYRA CVLSTPPTLS LLTIGFLFKK LGRQLRYLAE LCGVG AVLP GTCGGGPRAA FPTGVKLLSY LYQEALHNCS NEHYPVLLSL LKTSCEPYTR FIHDWVYSGV FRDAYGEFMI QVNHEY LSF RDKLYWTHGY VLISKEVEDC VPVFLKHIAH DIYVCGKTIN LLKLCCPRHY LCWSDVPVPR ISVIFSLEEL KEIEKDC AV YVGRMERVAR HSSVSKEEKE LRMEIAKQEL IAHAREAASR VLSALSDRQM SERMALDARK REQFQRLKEQ FVKDQERR Q AARQEELDDD FSYARELRDR ERRLKSLEEE LERKARQALV DHYSKLSAEA ARREQKALWR IQRHRLESAR LRFLLEDEK HIQEMLKAVS EAHQPQEPPD VLLSVHPQVT SPGPEHPEGG QGCDSGSAEQ HSPAWDGWNR PGLLTPQPLK PLAVGAGGRG LQQAEGARP FSDSLSIGDF LPVGPGAEPS VQTGMVPLLE VALQTINLDL PPSAPGEAPA AASTQPSRPQ EYDFSTVLRP A VATSPAPG PLQAAECSLG SSGLQLWEDS CGKMDACGSA SRETLLPSHP PRRAALEEGS SQPTERLFGQ VSGGGLPTGD YA SEIAPTR PRWNTHGHVS DASIRVGENV SDVAPTQPRW NTHGHVSNAS ISLGESVSDV APTRPRWNIH GHVSNASIRV GEN VSDVAP TRPRWNTHGH VSNASIRVGE NVSDVAPTRP RWNTHGHVSD ASISLGESVS DMAPARPRWN THGHVSDASI SLGE SVSDM APTRPRWNTH GHVSDTSIRV GENVSDVAPI RSRCNTHGHV SDASISLGEP VSDVVSTRPR WNTHVPIPPP HMVLG ALSP EAEPNTPRPQ QSPPGHTSQS ALSLGAQSTV LDCGPRLPVE VGPSLSSPSS GCGEGSISVG ENVSDVAPTQ PWWPNT PGD SVSEELGPGR SGDTEDLSPN WPLNSQEDTA AQSSPGRGEE AEASAAEAQG GEQAYLAGLA GQYHLERYPD SYESMSE PP IAHLLRPVLP RAFAFPVDPQ VQSAADETAV QLSELLTLPV LMKRSITAPL AAHISLVNKA AVDYFFVELH LEAHYEAL R HFLLMEDGEF AQSLSDLLFE KLGAGQTPGE LLNPLVLNSV LSKALQCSLH GDTPHASNLS LALKYLPEVF APNAPDVLS CLELRYKVDW PLNIVITEGC VSKYSGVFSF LLQLKLMMWA LKDVCFHLKR TALLSHMAGS VQFRQLQLFK HEMQHFVKVI QGYIANQIL HVTWCEFRAR LATVGDLEEI QRAHAEYLHK AVFRGLLTEK AAPVMNVIHS IFSLVLKFRS QLISQAWGPP G GPRGAEHP NFALMQQSYN TFKYYSHFLF KVVTKLVNRG YQPHLEDFLL RINFNNYYQD A UniProtKB: Gamma-tubulin complex component 6 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.9 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Pretreatment - Atmosphere: AIR / Details: 15 mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 310 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 46276 / Average exposure time: 1.3 sec. / Average electron dose: 46.276 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)