[English] 日本語
 Yorodumi
Yorodumi- EMDB-17670: Portal from the Borrelia bacteriophage BB1 procapsid with bound s... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Portal from the Borrelia bacteriophage BB1 procapsid with bound scaffold protein | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Bacteriophage / portal / DNA packaging / scaffold / VIRAL PROTEIN | ||||||||||||
| Function / homology | Uncharacterised conserved protein UCP020357 / Protein of unknown function DUF1357 / Protein of unknown function (DUF1357) / Protein of unknown function DUF1073 / Phage portal protein / Uncharacterized protein / Phage portal protein Function and homology information Function and homology information | ||||||||||||
| Biological species |  Borreliella burgdorferi B31 (bacteria) Borreliella burgdorferi B31 (bacteria) | ||||||||||||
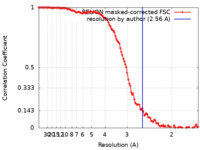
| Method | single particle reconstruction / cryo EM / Resolution: 2.56 Å | ||||||||||||
 Authors Authors | Rumnieks J / Fuzik T / Tars K | ||||||||||||
| Funding support | European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2023 Journal: J Mol Biol / Year: 2023Title: Structure of the Borrelia Bacteriophage φBB1 Procapsid. Authors: Jānis Rūmnieks / Tibor Füzik / Kaspars Tārs /   Abstract: Bacteriophages of Borrelia burgdorferi are a biologically important but under-investigated feature of the Lyme disease-causing spirochete. No virulent borrelial viruses have been identified, but all ...Bacteriophages of Borrelia burgdorferi are a biologically important but under-investigated feature of the Lyme disease-causing spirochete. No virulent borrelial viruses have been identified, but all B. burgdorferi isolates carry a prophage φBB1 as resident circular plasmids. Like its host, the φBB1 phage is quite distinctive and shares little sequence similarity with other known bacteriophages. We expressed φBB1 head morphogenesis proteins in Escherichia coli which resulted in assembly of homogeneous prolate procapsid structures and used cryo-electron microscopy to determine the three-dimensional structure of these particles. The φBB1 procapsids consist of 415 copies of the major capsid protein and an equal combined number of three homologous capsid decoration proteins that form trimeric knobs on the outside of the particle. One of the end vertices of the particle is occupied by a portal assembled from twelve copies of the portal protein. The φBB1 scaffolding protein is entirely α-helical and has an elongated shape with a small globular domain in the middle. Within the tubular section of the procapsid, the internal scaffold is built of stacked rings, each composed of 32 scaffolding protein molecules, which run in opposite directions from both caps with a heterogeneous part in the middle. Inside the portal-containing cap, the scaffold is organized asymmetrically with ten scaffolding protein molecules bound to the portal. The φBB1 procapsid structure provides better insight into the vast structural diversity of bacteriophages and presents clues of how elongated bacteriophage particles might be assembled. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17670.map.gz emd_17670.map.gz | 30.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17670-v30.xml emd-17670-v30.xml emd-17670.xml emd-17670.xml | 18.9 KB 18.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17670_fsc.xml emd_17670_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_17670.png emd_17670.png | 164.6 KB | ||
| Masks |  emd_17670_msk_1.map emd_17670_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17670.cif.gz emd-17670.cif.gz | 6.1 KB | ||
| Others |  emd_17670_half_map_1.map.gz emd_17670_half_map_1.map.gz emd_17670_half_map_2.map.gz emd_17670_half_map_2.map.gz | 93.5 MB 93.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17670 http://ftp.pdbj.org/pub/emdb/structures/EMD-17670 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17670 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17670 | HTTPS FTP |
-Related structure data
| Related structure data |  8phpMC  8phoC  8phqC  8phrC  8phsC  8phtC  8phuC  8pkhC  8qo0C  8qo1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17670.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17670.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8336 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
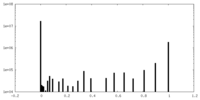
-Mask #1
| File |  emd_17670_msk_1.map emd_17670_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
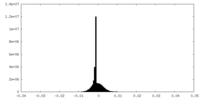
-Half map: #2
| File | emd_17670_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
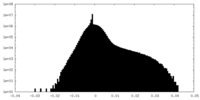
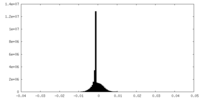
-Half map: #1
| File | emd_17670_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Portal with bound scaffold protein
| Entire | Name: Portal with bound scaffold protein |
|---|---|
| Components |
|
-Supramolecule #1: Portal with bound scaffold protein
| Supramolecule | Name: Portal with bound scaffold protein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Borreliella burgdorferi B31 (bacteria) Borreliella burgdorferi B31 (bacteria) |
-Macromolecule #1: Phage portal protein
| Macromolecule | Name: Phage portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Borreliella burgdorferi B31 (bacteria) Borreliella burgdorferi B31 (bacteria) |
| Molecular weight | Theoretical: 47.095508 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MCDLRKTKLI DKISSLELYK YSIFFRNYIE NVAEDCLKNG LILESAAHNV SEVELARLKV QLKNALLNCI ISYRFHGIGY VLVKTKDTL IDLEQPVNIE LPIGFEYLDY EYVRDLGVDF DHITYKVKSN NKNNSLDAVK IHKSRLIIYE NFDYILKRYV P CYTESFLL ...String: MCDLRKTKLI DKISSLELYK YSIFFRNYIE NVAEDCLKNG LILESAAHNV SEVELARLKV QLKNALLNCI ISYRFHGIGY VLVKTKDTL IDLEQPVNIE LPIGFEYLDY EYVRDLGVDF DHITYKVKSN NKNNSLDAVK IHKSRLIIYE NFDYILKRYV P CYTESFLL DIYLFEKIYV EIERRIENHN FLFYKDESLV QLQDALSSAT TSLSALTQSN NDRGSGILSS FLRKQNSNNH SK DISNLRN LNDSLSQELA RLKSNLNNEG MFYTATPSAS LEVIKYDLSY LKEALALIKA KIGADTKEPL TRSFNEQAKG LGN DGKGDR SNYYDFLKGV QEQVENSCNL KLTKYFGLDM KFNSLIMLSE EQKVERDIKL IELYSKYNQL IQSSSFNNEE LAML KEKLF SF UniProtKB: Phage portal protein |
-Macromolecule #2: Scaffold protein
| Macromolecule | Name: Scaffold protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Borreliella burgdorferi B31 (bacteria) Borreliella burgdorferi B31 (bacteria) |
| Molecular weight | Theoretical: 26.710363 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTEKEEKEDL QAQDKEEQQI KADTKVISVQ EFEEYMRFKE QANSKSKETS RDLSINERIT KELAEVEERE RIEKQLLLEA ERINEIDTL AKAHLSNHFN KEVLLAKGYT LKDIMQAQRR ELVRKFVPIE QIKAIAKVSD ISHIDGEILE QLVSLAKVNI K LRKNASSS ...String: MTEKEEKEDL QAQDKEEQQI KADTKVISVQ EFEEYMRFKE QANSKSKETS RDLSINERIT KELAEVEERE RIEKQLLLEA ERINEIDTL AKAHLSNHFN KEVLLAKGYT LKDIMQAQRR ELVRKFVPIE QIKAIAKVSD ISHIDGEILE QLVSLAKVNI K LRKNASSS SSSVDSIKGN IAIKSEERAS LLDSNFVPIN FTEFVQAISN TYKQRRIQFY ENLKRHKRTS IA UniProtKB: Uncharacterized protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - #0 - Film type ID: 1 / Support film - #0 - Material: CARBON / Support film - #0 - topology: HOLEY / Support film - #1 - Film type ID: 2 / Support film - #1 - Material: GRAPHENE / Support film - #1 - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Output model |  PDB-8php: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)