+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | BtuB3G3 bound to cyanocobalamin with disordered EL8 | |||||||||
 Map data Map data | Sharpened final map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cyanocobalamin / transporter / lipoprotein / complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsiderophore transmembrane transport / siderophore uptake transmembrane transporter activity / cell outer membrane Similarity search - Function | |||||||||
| Biological species |  Bacteroides thetaiotaomicron VPI-5482 (bacteria) Bacteroides thetaiotaomicron VPI-5482 (bacteria) | |||||||||
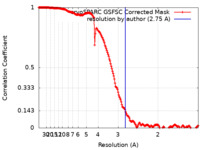
| Method | single particle reconstruction / cryo EM / Resolution: 2.75 Å | |||||||||
 Authors Authors | Silale A / Abellon-Ruiz J / van den Berg B | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: BtuB TonB-dependent transporters and BtuG surface lipoproteins form stable complexes for vitamin B uptake in gut Bacteroides. Authors: Javier Abellon-Ruiz / Kalyanashis Jana / Augustinas Silale / Andrew M Frey / Arnaud Baslé / Matthias Trost / Ulrich Kleinekathöfer / Bert van den Berg /   Abstract: Vitamin B (cobalamin) is required for most human gut microbes, many of which are dependent on scavenging to obtain this vitamin. Since bacterial densities in the gut are extremely high, competition ...Vitamin B (cobalamin) is required for most human gut microbes, many of which are dependent on scavenging to obtain this vitamin. Since bacterial densities in the gut are extremely high, competition for this keystone micronutrient is severe. Contrasting with Enterobacteria, members of the dominant genus Bacteroides often encode several BtuB vitamin B outer membrane transporters together with a conserved array of surface-exposed B-binding lipoproteins. Here we show that the BtuB transporters from Bacteroides thetaiotaomicron form stable, pedal bin-like complexes with surface-exposed BtuG lipoprotein lids, which bind B with high affinities. Closing of the BtuG lid following B capture causes destabilisation of the bound B by a conserved BtuB extracellular loop, causing translocation of the vitamin to BtuB and subsequent transport. We propose that TonB-dependent, lipoprotein-assisted small molecule uptake is a general feature of Bacteroides spp. that is important for the success of this genus in colonising the human gut. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17574.map.gz emd_17574.map.gz | 484 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17574-v30.xml emd-17574-v30.xml emd-17574.xml emd-17574.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17574_fsc.xml emd_17574_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_17574.png emd_17574.png | 72.6 KB | ||
| Masks |  emd_17574_msk_1.map emd_17574_msk_1.map | 512 MB |  Mask map Mask map | |
| Others |  emd_17574_half_map_1.map.gz emd_17574_half_map_1.map.gz emd_17574_half_map_2.map.gz emd_17574_half_map_2.map.gz | 474.6 MB 474.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17574 http://ftp.pdbj.org/pub/emdb/structures/EMD-17574 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17574 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17574 | HTTPS FTP |
-Related structure data
| Related structure data |  8p97MC  8blwC  8bmxC  8bmyC  8bmzC  8bn0C  8okvC  8p98C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17574.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17574.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened final map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.74 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17574_msk_1.map emd_17574_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_17574_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_17574_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of TonB-dependent transporter BtuB3 with surface lipoprot...
| Entire | Name: Complex of TonB-dependent transporter BtuB3 with surface lipoprotein BtuG3 bound to cyanocobalamin (state 2) |
|---|---|
| Components |
|
-Supramolecule #1: Complex of TonB-dependent transporter BtuB3 with surface lipoprot...
| Supramolecule | Name: Complex of TonB-dependent transporter BtuB3 with surface lipoprotein BtuG3 bound to cyanocobalamin (state 2) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Bacteroides thetaiotaomicron VPI-5482 (bacteria) Bacteroides thetaiotaomicron VPI-5482 (bacteria) |
| Molecular weight | Theoretical: 122 KDa |
-Macromolecule #1: TonB-dependent receptor
| Macromolecule | Name: TonB-dependent receptor / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Bacteroides thetaiotaomicron VPI-5482 (bacteria) Bacteroides thetaiotaomicron VPI-5482 (bacteria) |
| Molecular weight | Theoretical: 78.890609 KDa |
| Recombinant expression | Organism:  Bacteroides thetaiotaomicron VPI-5482 (bacteria) Bacteroides thetaiotaomicron VPI-5482 (bacteria) |
| Sequence | String: MSGQQHSSDN KNWAEKGIVI REVEILGKRP MKDIGVQQTR FDSLVLKENI ALSMADILTF NSSIFVKSYG RATLSTVSFR GTSASHTQV TWNGMRINNP MLGMTDFSMI PSYFIDDASL LHGTSSVNMA GGGLGGLVKL STVPAHQEGF GMQYVQGIGS F STFDEFLQ ...String: MSGQQHSSDN KNWAEKGIVI REVEILGKRP MKDIGVQQTR FDSLVLKENI ALSMADILTF NSSIFVKSYG RATLSTVSFR GTSASHTQV TWNGMRINNP MLGMTDFSMI PSYFIDDASL LHGTSSVNMA GGGLGGLVKL STVPAHQEGF GMQYVQGIGS F STFDEFLQ LKYGDKHWQI STRAVYQSSP NDYKYRNHDK KENIYDDKYN IIEQYYPIER NRSGAYKDLH ILQEVYYNTG KG DRFGLNA WYTDSNRELA LLTTDQGDLM DFENRQREHT LRSVLSWDHT RENWKVSARG GYVHTWLAYD YKRDLGNGIM ATM TRSRSK VNTFYGQLDG EYFFSDKLLL TAGVSAHQHL VNSLDKDLNK DDNKNDKYGQ GRKNDSIVYF DKGRIELSGN VSLK WQPVN RLGMSLVLRG EMFGTKWAPV IPAFFVDYVL SKRGNIMAKA SITRNYRFPT LNDLYFLPGG NPALNNESGF TYETG LSFS VDKDNVYTLS GSASWFDQHI NDWIIWLPTS KGFYSPVNLK KVHAYGVEVQ ADYAVAIDKA WKLGLNGTFA WTPSIN EGE PTSKADQSVG KQLPYIPEYS ATLSGRLTYR SWGLLYKWCY YSERYTMTSN AVSYTGHLPP YLMSNVTLEK GFSLRWA DL SLKGTVNNLF DEEYLSVLSR PMPGINFEFF IGITPKWGKK KKGGGHHHHH H UniProtKB: TonB-dependent receptor |
-Macromolecule #2: Putative surface layer protein
| Macromolecule | Name: Putative surface layer protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Bacteroides thetaiotaomicron VPI-5482 (bacteria) Bacteroides thetaiotaomicron VPI-5482 (bacteria) |
| Molecular weight | Theoretical: 41.165793 KDa |
| Sequence | String: MKRILLSVLF IVFCLTAFVG CMKWDYGKME PFRATGDGLF IMNEGNFQYG NATLSYYDPE TKKVENEIFY RANAMKLGDV AQSMIVRDT IGWVVVNNSH VIFAISTNTF KEVGRITGLT SPRYIHFISD EKAYITQIWD YRIFIVNPKT YQITGYIECP D MTMETGST ...String: MKRILLSVLF IVFCLTAFVG CMKWDYGKME PFRATGDGLF IMNEGNFQYG NATLSYYDPE TKKVENEIFY RANAMKLGDV AQSMIVRDT IGWVVVNNSH VIFAISTNTF KEVGRITGLT SPRYIHFISD EKAYITQIWD YRIFIVNPKT YQITGYIECP D MTMETGST EQMVQYGKYV YVNCWSYQNR ILKIDTTTDK VVDQLTVGIQ PTSLVMDKNF KMWTITDGGY KGSPYGYEEP SL YRIDAET FKIEKQFKFQ LGDAPSEVQL NGAGDELYWI NKDIWRMSVD EERVPVRPFL KYRDTKYYGL TVSPKNGDVY VAD AIDYQQ QGMIYRYTED GELVDEFYVG IIPGAFCWK UniProtKB: Surface layer protein |
-Macromolecule #3: CYANOCOBALAMIN
| Macromolecule | Name: CYANOCOBALAMIN / type: ligand / ID: 3 / Number of copies: 1 / Formula: CNC |
|---|---|
| Molecular weight | Theoretical: 1.356373 KDa |
| Chemical component information |  ChemComp-CNC: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | |||||||||
| Details | The protein complex was purified in lauryl maltose neopentyl glycol (LMNG). Prior to making grids, the sample was subjected to size exclusion chromatography without LMNG in the mobile phase to separate protein-LMNG complexes from free LMNG. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Number real images: 9826 / Average electron dose: 35.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)