[English] 日本語
 Yorodumi
Yorodumi- EMDB-17205: Human RAD51B-RAD51C-RAD51D-XRCC2 (BCDX2) complex, 3.4 A resolution -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human RAD51B-RAD51C-RAD51D-XRCC2 (BCDX2) complex, 3.4 A resolution | ||||||||||||||||||
 Map data Map data | (HANDEDNESS INVERTED) | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Complex / ssDNA-binding / ATPase / RAD51 / DNA BINDING PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationmeiotic DNA recombinase assembly / Rad51C-XRCC3 complex / Rad51B-Rad51C-Rad51D-XRCC2 complex / female meiosis sister chromatid cohesion / blastocyst growth / crossover junction DNA endonuclease activity / somite development / DNA strand invasion / gamma-tubulin binding / Impaired BRCA2 binding to PALB2 ...meiotic DNA recombinase assembly / Rad51C-XRCC3 complex / Rad51B-Rad51C-Rad51D-XRCC2 complex / female meiosis sister chromatid cohesion / blastocyst growth / crossover junction DNA endonuclease activity / somite development / DNA strand invasion / gamma-tubulin binding / Impaired BRCA2 binding to PALB2 / telomere maintenance via recombination / regulation of fibroblast apoptotic process / reciprocal meiotic recombination / centrosome cycle / positive regulation of neurogenesis / sister chromatid cohesion / ATP-dependent DNA damage sensor activity / Homologous DNA Pairing and Strand Exchange / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Resolution of D-loop Structures through Synthesis-Dependent Strand Annealing (SDSA) / Resolution of D-loop Structures through Holliday Junction Intermediates / microtubule organizing center / male meiosis I / Presynaptic phase of homologous DNA pairing and strand exchange / positive regulation of G2/M transition of mitotic cell cycle / response to X-ray / somitogenesis / ATP-dependent activity, acting on DNA / interstrand cross-link repair / neurogenesis / telomere maintenance / replication fork / response to gamma radiation / meiotic cell cycle / TP53 Regulates Transcription of DNA Repair Genes / double-strand break repair via homologous recombination / Meiotic recombination / HDR through Homologous Recombination (HRR) / multicellular organism growth / cell junction / mitotic cell cycle / single-stranded DNA binding / Factors involved in megakaryocyte development and platelet production / double-stranded DNA binding / DNA recombination / spermatogenesis / in utero embryonic development / negative regulation of neuron apoptotic process / chromosome, telomeric region / regulation of cell cycle / DNA repair / positive regulation of cell population proliferation / centrosome / perinuclear region of cytoplasm / mitochondrion / DNA binding / nucleoplasm / ATP binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | ||||||||||||||||||
 Authors Authors | Greenhough LA / Liang CC / West SC | ||||||||||||||||||
| Funding support |  United Kingdom, European Union, 5 items United Kingdom, European Union, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Structure and function of the RAD51B-RAD51C-RAD51D-XRCC2 tumour suppressor. Authors: Luke A Greenhough / Chih-Chao Liang / Ondrej Belan / Simone Kunzelmann / Sarah Maslen / Monica C Rodrigo-Brenni / Roopesh Anand / Mark Skehel / Simon J Boulton / Stephen C West /   Abstract: Homologous recombination is a fundamental process of life. It is required for the protection and restart of broken replication forks, the repair of chromosome breaks and the exchange of genetic ...Homologous recombination is a fundamental process of life. It is required for the protection and restart of broken replication forks, the repair of chromosome breaks and the exchange of genetic material during meiosis. Individuals with mutations in key recombination genes, such as BRCA2 (also known as FANCD1), or the RAD51 paralogues RAD51B, RAD51C (also known as FANCO), RAD51D, XRCC2 (also known as FANCU) and XRCC3, are predisposed to breast, ovarian and prostate cancers and the cancer-prone syndrome Fanconi anaemia. The BRCA2 tumour suppressor protein-the product of BRCA2-is well characterized, but the cellular functions of the RAD51 paralogues remain unclear. Genetic knockouts display growth defects, reduced RAD51 focus formation, spontaneous chromosome abnormalities, sensitivity to PARP inhibitors and replication fork defects, but the precise molecular roles of RAD51 paralogues in fork stability, DNA repair and cancer avoidance remain unknown. Here we used cryo-electron microscopy, AlphaFold2 modelling and structural proteomics to determine the structure of the RAD51B-RAD51C-RAD51D-XRCC2 complex (BCDX2), revealing that RAD51C-RAD51D-XRCC2 mimics three RAD51 protomers aligned within a nucleoprotein filament, whereas RAD51B is highly dynamic. Biochemical and single-molecule analyses showed that BCDX2 stimulates the nucleation and extension of RAD51 filaments-which are essential for recombinational DNA repair-in reactions that depend on the coupled ATPase activities of RAD51B and RAD51C. Our studies demonstrate that BCDX2 orchestrates RAD51 assembly on single stranded DNA for replication fork protection and double strand break repair, in reactions that are critical for tumour avoidance. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17205.map.gz emd_17205.map.gz | 4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17205-v30.xml emd-17205-v30.xml emd-17205.xml emd-17205.xml | 25.6 KB 25.6 KB | Display Display |  EMDB header EMDB header |
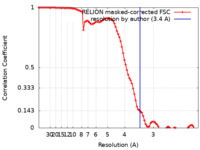
| FSC (resolution estimation) |  emd_17205_fsc.xml emd_17205_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_17205.png emd_17205.png | 61.3 KB | ||
| Filedesc metadata |  emd-17205.cif.gz emd-17205.cif.gz | 7.7 KB | ||
| Others |  emd_17205_half_map_1.map.gz emd_17205_half_map_1.map.gz emd_17205_half_map_2.map.gz emd_17205_half_map_2.map.gz | 50 MB 50 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17205 http://ftp.pdbj.org/pub/emdb/structures/EMD-17205 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17205 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17205 | HTTPS FTP |
-Related structure data
| Related structure data |  8ouyMC  8ouzC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17205.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17205.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | (HANDEDNESS INVERTED) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Corrected handedness
| File | emd_17205_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Corrected handedness | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Corrected handedness
| File | emd_17205_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Corrected handedness | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : RAD51B-RAD51C-RAD51D-XRCC2 (BCDX2)
| Entire | Name: RAD51B-RAD51C-RAD51D-XRCC2 (BCDX2) |
|---|---|
| Components |
|
-Supramolecule #1: RAD51B-RAD51C-RAD51D-XRCC2 (BCDX2)
| Supramolecule | Name: RAD51B-RAD51C-RAD51D-XRCC2 (BCDX2) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: Recombinant BCDX2 complexes purified from Sf9 insect cells, crosslinked with 0.005% glutaraldehyde (RT, 10 minutes) in the presence of 30 nt ssDNA, and vitrified in the presence of ADP.AlFx. ...Details: Recombinant BCDX2 complexes purified from Sf9 insect cells, crosslinked with 0.005% glutaraldehyde (RT, 10 minutes) in the presence of 30 nt ssDNA, and vitrified in the presence of ADP.AlFx. Map was enhanced using DeepEMhancer |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: DNA repair protein RAD51 homolog 2
| Macromolecule | Name: DNA repair protein RAD51 homolog 2 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.296984 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSKKLKRVG LSQELCDRLS RHQILTCQDF LCLSPLELMK VTGLSYRGVH ELLCMVSRAC APKMQTAYGI KAQRSADFSP AFLSTTLSA LDEALHGGVA CGSLTEITGP PGCGKTQFCI MMSILATLPT NMGGLEGAVV YIDTESAFSA ERLVEIAESR F PRYFNTEE ...String: MGSKKLKRVG LSQELCDRLS RHQILTCQDF LCLSPLELMK VTGLSYRGVH ELLCMVSRAC APKMQTAYGI KAQRSADFSP AFLSTTLSA LDEALHGGVA CGSLTEITGP PGCGKTQFCI MMSILATLPT NMGGLEGAVV YIDTESAFSA ERLVEIAESR F PRYFNTEE KLLLTSSKVH LYRELTCDEV LQRIESLEEE IISKGIKLVI LDSVASVVRK EFDAQLQGNL KERNKFLARE AS SLKYLAE EFSIPVILTN QITTHLSGAL ASQADLVSPA DDLSLSEGTS GSSCVIAALG NTWSHSVNTR LILQYLDSER RQI LIAKSP LAPFTSFVYT IKEEGLVLQA YGNS UniProtKB: DNA repair protein RAD51 homolog 2 |
-Macromolecule #2: DNA repair protein RAD51 homolog 3
| Macromolecule | Name: DNA repair protein RAD51 homolog 3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.244609 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRGKTFRFEM QRDLVSFPLS PAVRVKLVSA GFQTAEELLE VKPSELSKEV GISKAEALET LQIIRRECLT NKPRYAGTSE SHKKCTALE LLEQEHTQGF IITFCSALDD ILGGGVPLMK TTEICGAPGV GKTQLCMQLA VDVQIPECFG GVAGEAVFID T EGSFMVDR ...String: MRGKTFRFEM QRDLVSFPLS PAVRVKLVSA GFQTAEELLE VKPSELSKEV GISKAEALET LQIIRRECLT NKPRYAGTSE SHKKCTALE LLEQEHTQGF IITFCSALDD ILGGGVPLMK TTEICGAPGV GKTQLCMQLA VDVQIPECFG GVAGEAVFID T EGSFMVDR VVDLATACIQ HLQLIAEKHK GEEHRKALED FTLDNILSHI YYFRCRDYTE LLAQVYLLPD FLSEHSKVRL VI VDGIAFP FRHDLDDLSL RTRLLNGLAQ QMISLANNHR LAVILTNQMT TKIDRNQALL VPALGESWGH AATIRLIFHW DRK QRLATL YKSPSQKECT VLFQIKPQGF RDTVVTSACS LQTEGSLSTR KRSRDPEEEL UniProtKB: DNA repair protein RAD51 homolog 3 |
-Macromolecule #3: DNA repair protein RAD51 homolog 4
| Macromolecule | Name: DNA repair protein RAD51 homolog 4 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 35.084254 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGVLRVGLCP GLTEEMIQLL RSHRIKTVVD LVSADLEEVA QKCGLSYKAL VALRRVLLAQ FSAFPVNGAD LYEELKTSTA ILSTGIGSL DKLLDAGLYT GEVTEIVGGP GSGKTQVCLC MAANVAHGLQ QNVLYVDSNG GLTASRLLQL LQAKTQDEEE Q AEALRRIQ ...String: MGVLRVGLCP GLTEEMIQLL RSHRIKTVVD LVSADLEEVA QKCGLSYKAL VALRRVLLAQ FSAFPVNGAD LYEELKTSTA ILSTGIGSL DKLLDAGLYT GEVTEIVGGP GSGKTQVCLC MAANVAHGLQ QNVLYVDSNG GLTASRLLQL LQAKTQDEEE Q AEALRRIQ VVHAFDIFQM LDVLQELRGT VAQQVTGSSG TVKVVVVDSV TAVVSPLLGG QQREGLALMM QLARELKTLA RD LGMAVVV TNHITRDRDS GRLKPALGRS WSFVPSTRIL LDTIEGAGAS GGRRMACLAK SSRQPTGFQE MVDIGTWGTS EQS ATLQGD QT UniProtKB: DNA repair protein RAD51 homolog 4 |
-Macromolecule #4: DNA repair protein XRCC2
| Macromolecule | Name: DNA repair protein XRCC2 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 31.995525 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MCSAFHRAES GTELLARLEG RSSLKEIEPN LFADEDSPVH GDILEFHGPE GTGKTEMLYH LTARCILPKS EGGLEVEVLF IDTDYHFDM LRLVTILEHR LSQSSEEIIK YCLGRFFLVY CSSSTHLLLT LYSLESMFCS HPSLCLLILD SLSAFYWIDR V NGGESVNL ...String: MCSAFHRAES GTELLARLEG RSSLKEIEPN LFADEDSPVH GDILEFHGPE GTGKTEMLYH LTARCILPKS EGGLEVEVLF IDTDYHFDM LRLVTILEHR LSQSSEEIIK YCLGRFFLVY CSSSTHLLLT LYSLESMFCS HPSLCLLILD SLSAFYWIDR V NGGESVNL QESTLRKCSQ CLEKLVNDYR LVLFATTQTI MQKASSSSEE PSHASRRLCD VDIDYRPYLC KAWQQLVKHR MF FSKQDDS QSSNQFSLVS RCLKSNSLKK HFFIIGESGV EFC UniProtKB: DNA repair protein XRCC2 |
-Macromolecule #5: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 1 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 3 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #7: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 7 / Number of copies: 2 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.25 mg/mL | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 25 mM HEPES pH 7.5, 100 mM NaCl, 2.5 mM MgCl2, 0.5 mM ADP.BeFx (0.5 mM ADP, 0.5 mM BeSO4, 10 mM NaF), 0.25 mM TCEP, 0.0005% Tween20 | |||||||||||||||||||||||||||
| Grid | Model: UltrAuFoil R2/2 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Details: 25 mA | |||||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number grids imaged: 1 / Number real images: 24449 / Average exposure time: 9.0 sec. / Average electron dose: 47.0 e/Å2 Details: 12,555 movies were collected with zero tilt, and a further 11,894 movies with a 20 degree tilt |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Details | AlphaFold2 monomer predictions of RAD51B, RAD51C, RAD51D and XRCC2 were docked into the EM density using Dock and Rebuild in Phenix. ATP and Mg2+ were added, and the coordinates were iteratively real space refined using Phenix and manually modified in COOT |
| Refinement | Space: REAL |
| Output model |  PDB-8ouy: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)