[English] 日本語
 Yorodumi
Yorodumi- EMDB-16901: CryoEM structure of 20S Trichomonas vaginalis proteasome in compl... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of 20S Trichomonas vaginalis proteasome in complex with proteasome inhibitor Salinosporamid A | |||||||||
 Map data Map data | 2.89A CryoEM map of Tv20S proteasome with covalent inhibitor Salinosporamide A (Marizomib) bound to beta1 and beta2 subunits. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Proteasome / 20S / Trichomonas vaginalis / covalently bound Salinosporamide A / Marizomib / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationproteasome endopeptidase complex / proteasome core complex, beta-subunit complex / threonine-type endopeptidase activity / proteasome core complex, alpha-subunit complex / endopeptidase activity / proteasome-mediated ubiquitin-dependent protein catabolic process / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Trichomonas vaginalis G3 (eukaryote) Trichomonas vaginalis G3 (eukaryote) | |||||||||
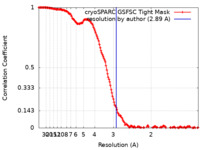
| Method | single particle reconstruction / cryo EM / Resolution: 2.89 Å | |||||||||
 Authors Authors | Silhan J / Fajtova P / Boura E | |||||||||
| Funding support | European Union, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural elucidation of recombinant Trichomonas vaginalis 20S proteasome bound to covalent inhibitors. Authors: Jan Silhan / Pavla Fajtova / Jitka Bartosova / Brianna M Hurysz / Jehad Almaliti / Yukiko Miyamoto / Lars Eckmann / William H Gerwick / Anthony J O'Donoghue / Evzen Boura /   Abstract: The proteasome is a proteolytic enzyme complex essential for protein homeostasis in mammalian cells and protozoan parasites like Trichomonas vaginalis (Tv), the cause of the most common, non-viral ...The proteasome is a proteolytic enzyme complex essential for protein homeostasis in mammalian cells and protozoan parasites like Trichomonas vaginalis (Tv), the cause of the most common, non-viral sexually transmitted disease. Tv and other protozoan 20S proteasomes have been validated as druggable targets for antimicrobials. However, low yields and purity of the native proteasome have hindered studies of the Tv 20S proteasome (Tv20S). We address this challenge by creating a recombinant protozoan proteasome by expressing all seven α and seven β subunits of Tv20S alongside the Ump-1 chaperone in insect cells. The recombinant Tv20S displays biochemical equivalence to its native counterpart, confirmed by various assays. Notably, the marizomib (MZB) inhibits all catalytic subunits of Tv20S, while the peptide inhibitor carmaphycin-17 (CP-17) specifically targets β2 and β5. Cryo-electron microscopy (cryo-EM) unveils the structures of Tv20S bound to MZB and CP-17 at 2.8 Å. These findings explain MZB's low specificity for Tv20S compared to the human proteasome and demonstrate CP-17's higher specificity. Overall, these data provide a structure-based strategy for the development of specific Tv20S inhibitors to treat trichomoniasis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16901.map.gz emd_16901.map.gz | 306.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16901-v30.xml emd-16901-v30.xml emd-16901.xml emd-16901.xml | 34.4 KB 34.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16901_fsc.xml emd_16901_fsc.xml | 16.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_16901.png emd_16901.png | 170.4 KB | ||
| Filedesc metadata |  emd-16901.cif.gz emd-16901.cif.gz | 9.1 KB | ||
| Others |  emd_16901_half_map_1.map.gz emd_16901_half_map_1.map.gz emd_16901_half_map_2.map.gz emd_16901_half_map_2.map.gz | 300.9 MB 300.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16901 http://ftp.pdbj.org/pub/emdb/structures/EMD-16901 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16901 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16901 | HTTPS FTP |
-Related structure data
| Related structure data |  8oixMC  8p0tC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16901.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16901.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
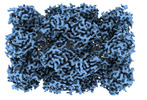
| Annotation | 2.89A CryoEM map of Tv20S proteasome with covalent inhibitor Salinosporamide A (Marizomib) bound to beta1 and beta2 subunits. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.7 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: 2.89A CryoEM map of Tv20S proteasome with covalent...
| File | emd_16901_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 2.89A CryoEM map of Tv20S proteasome with covalent inhibitor Salinosporamide A (Marizomib) bound to beta1 and beta2 subunits. | ||||||||||||
| Projections & Slices |
| ||||||||||||
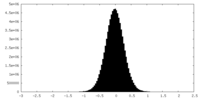
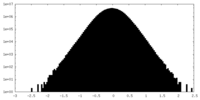
| Density Histograms |
-Half map: 2.89A CryoEM map of Tv20S proteasome with covalent...
| File | emd_16901_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 2.89A CryoEM map of Tv20S proteasome with covalent inhibitor Salinosporamide A (Marizomib) bound to beta1 and beta2 subunits. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Ternary complex of 20S proteasome from Trichomonas vaginalis
+Supramolecule #1: Ternary complex of 20S proteasome from Trichomonas vaginalis
+Macromolecule #1: Family T1, proteasome alpha subunit, threonine peptidase
+Macromolecule #2: Family T1, proteasome alpha subunit, threonine peptidase
+Macromolecule #3: Proteasome subunit alpha type
+Macromolecule #4: Proteasome subunit alpha type
+Macromolecule #5: Proteasome subunit alpha type
+Macromolecule #6: Family T1, proteasome alpha subunit, threonine peptidase
+Macromolecule #7: Family T1, proteasome alpha subunit, threonine peptidase
+Macromolecule #8: Proteasome subunit beta
+Macromolecule #9: proteasome endopeptidase complex
+Macromolecule #10: Proteasome subunit beta
+Macromolecule #11: Proteasome subunit beta
+Macromolecule #12: Proteasome subunit beta
+Macromolecule #13: Proteasome subunit beta
+Macromolecule #14: Family T1, proteasome beta subunit, threonine peptidase
+Macromolecule #15: (3AR,6R,6AS)-6-((S)-((S)-CYCLOHEX-2-ENYL)(HYDROXY)METHYL)-6A-METH...
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.015 kPa / Details: 15 mA | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 7983 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.6 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)