+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | in situ Subtomogram average of Immature Rotavirus TLP spike | ||||||||||||||||||
 Map data Map data | main map | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Rotavirus / STA / VIRAL PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationviral intermediate capsid / host cell endoplasmic reticulum lumen / host cell rough endoplasmic reticulum / T=13 icosahedral viral capsid / permeabilization of host organelle membrane involved in viral entry into host cell / host cytoskeleton / viral outer capsid / host cell endoplasmic reticulum-Golgi intermediate compartment / host cell surface receptor binding / fusion of virus membrane with host plasma membrane ...viral intermediate capsid / host cell endoplasmic reticulum lumen / host cell rough endoplasmic reticulum / T=13 icosahedral viral capsid / permeabilization of host organelle membrane involved in viral entry into host cell / host cytoskeleton / viral outer capsid / host cell endoplasmic reticulum-Golgi intermediate compartment / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / structural molecule activity / metal ion binding / membrane Similarity search - Function | ||||||||||||||||||
| Biological species |  Rotavirus A Rotavirus A | ||||||||||||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 4.5 Å | ||||||||||||||||||
 Authors Authors | Shah PNM / Stuart DI | ||||||||||||||||||
| Funding support |  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Cell Host Microbe / Year: 2023 Journal: Cell Host Microbe / Year: 2023Title: Characterization of the rotavirus assembly pathway in situ using cryoelectron tomography. Authors: Pranav N M Shah / James B Gilchrist / Björn O Forsberg / Alister Burt / Andrew Howe / Shyamal Mosalaganti / William Wan / Julika Radecke / Yuriy Chaban / Geoff Sutton / David I Stuart / Mark Boyce /    Abstract: Rotavirus assembly is a complex process that involves the stepwise acquisition of protein layers in distinct intracellular locations to form the fully assembled particle. Understanding and ...Rotavirus assembly is a complex process that involves the stepwise acquisition of protein layers in distinct intracellular locations to form the fully assembled particle. Understanding and visualization of the assembly process has been hampered by the inaccessibility of unstable intermediates. We characterize the assembly pathway of group A rotaviruses observed in situ within cryo-preserved infected cells through the use of cryoelectron tomography of cellular lamellae. Our findings demonstrate that the viral polymerase VP1 recruits viral genomes during particle assembly, as revealed by infecting with a conditionally lethal mutant. Additionally, pharmacological inhibition to arrest the transiently enveloped stage uncovered a unique conformation of the VP4 spike. Subtomogram averaging provided atomic models of four intermediate states, including a pre-packaging single-layered intermediate, the double-layered particle, the transiently enveloped double-layered particle, and the fully assembled triple-layered virus particle. In summary, these complementary approaches enable us to elucidate the discrete steps involved in forming an intracellular rotavirus particle. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16774.map.gz emd_16774.map.gz | 7.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16774-v30.xml emd-16774-v30.xml emd-16774.xml emd-16774.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_16774.png emd_16774.png | 116 KB | ||
| Filedesc metadata |  emd-16774.cif.gz emd-16774.cif.gz | 6.7 KB | ||
| Others |  emd_16774_half_map_1.map.gz emd_16774_half_map_1.map.gz emd_16774_half_map_2.map.gz emd_16774_half_map_2.map.gz | 7.9 MB 7.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16774 http://ftp.pdbj.org/pub/emdb/structures/EMD-16774 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16774 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16774 | HTTPS FTP |
-Validation report
| Summary document |  emd_16774_validation.pdf.gz emd_16774_validation.pdf.gz | 882.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16774_full_validation.pdf.gz emd_16774_full_validation.pdf.gz | 881.7 KB | Display | |
| Data in XML |  emd_16774_validation.xml.gz emd_16774_validation.xml.gz | 9.3 KB | Display | |
| Data in CIF |  emd_16774_validation.cif.gz emd_16774_validation.cif.gz | 10.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16774 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16774 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16774 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16774 | HTTPS FTP |
-Related structure data
| Related structure data |  8coaMC  8bp8C  8co6C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16774.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16774.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.97 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half1
| File | emd_16774_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
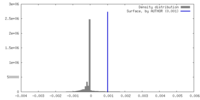
| Density Histograms |
-Half map: half2
| File | emd_16774_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
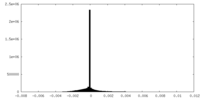
| Density Histograms |
- Sample components
Sample components
-Entire : Rotavirus A
| Entire | Name:  Rotavirus A Rotavirus A |
|---|---|
| Components |
|
-Supramolecule #1: Rotavirus A
| Supramolecule | Name: Rotavirus A / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 28875 / Sci species name: Rotavirus A / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: Outer capsid protein VP4
| Macromolecule | Name: Outer capsid protein VP4 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Rotavirus A Rotavirus A |
| Molecular weight | Theoretical: 86.767953 KDa |
| Recombinant expression | Organism:  Chlorocebus aethiops aethiops (mammal) Chlorocebus aethiops aethiops (mammal) |
| Sequence | String: MASLIYRQLL TNSYTVDLSD EIQEIGSTKS QNVTINPGPF AQTGYAPVNW GPGEINDSTT VGPLLDGPYQ PTTFNPPVDY WMLLAPTTP GVIVEGTNNT DRWLATILIE PNVQSENRTY TIFGIQEQLT VSNTSQDQWK FIDVVKTTAN GSIGQYGPLL S SPKLYAVM ...String: MASLIYRQLL TNSYTVDLSD EIQEIGSTKS QNVTINPGPF AQTGYAPVNW GPGEINDSTT VGPLLDGPYQ PTTFNPPVDY WMLLAPTTP GVIVEGTNNT DRWLATILIE PNVQSENRTY TIFGIQEQLT VSNTSQDQWK FIDVVKTTAN GSIGQYGPLL S SPKLYAVM KHNEKLYTYE GQTPNATTAH YSTTNYDSVN MTAFCDFYII PRSEESKCTE YINNGLPPIQ NTRNVVPLSL TA RDVIHYR AQANEDIVIS KTSLWKEMQY NRDITIRFKF ANTIIKSGGL GYKWSEISFK PANYQYTYTR DGEEVTAHTT CSV NGMNDF SFNGGYLPTD FVVSKFEVIK ENSYVYIDYW DDSQAFRNVV YVRSLAANLN SVMCTGGSYN FSLPVGQWPV LTGG AVSLH SAGVTLSTQF TDFVSLNSLR FRFRLAVEEP HFKLTRTRLD RLYGLPAADP NNGKEYYEIA GRFSLISLVP SNDDY QTPI ANSVTVRQDL ERQLGELREE FNALSQEIAM SQLIDLALLP LDMFSMFSGI KSTIDAAKSM ATNVMKKFKK SGLANS VST LTDSLSDAAS SISRGSSIRS IGSSASAWTD VSTQITDISS SVSSVSTQTS TISRRLRLKE MATQTEGMNF DDISAAV LK TKIDKSTQIS PNTIPDIVTE ASEKFIPNRA YRVINNDDVF EAGIDGKFFA YKVDTFEEIP FDVQKFADLV TDSPVISA I IDFKTLKNLN DNYGISRQQA FNLLRSDPRV LREFINQDNP IIRNRIEQLI MQCRL UniProtKB: Outer capsid protein VP4 |
-Macromolecule #2: Outer capsid glycoprotein VP7
| Macromolecule | Name: Outer capsid glycoprotein VP7 / type: protein_or_peptide / ID: 2 / Number of copies: 13 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Rotavirus A Rotavirus A |
| Molecular weight | Theoretical: 37.230664 KDa |
| Recombinant expression | Organism:  Chlorocebus aethiops aethiops (mammal) Chlorocebus aethiops aethiops (mammal) |
| Sequence | String: MYGIEYTTVL TFLISIILLN YILKSLTRIM DFIIYRFLFI IVILSPFLRA QNYGINLPIT GSMDIAYANS TQEEPFLTST LCLYYPTEA ATEINDNSWK DTLSQLFLTK GWPTGSVYFK EYTNIASFSV DPQLYCDYNV VLMKYDATLQ LDMSELADLI L NEWLCNPM ...String: MYGIEYTTVL TFLISIILLN YILKSLTRIM DFIIYRFLFI IVILSPFLRA QNYGINLPIT GSMDIAYANS TQEEPFLTST LCLYYPTEA ATEINDNSWK DTLSQLFLTK GWPTGSVYFK EYTNIASFSV DPQLYCDYNV VLMKYDATLQ LDMSELADLI L NEWLCNPM DITLYYYQQT DEANKWISMG SSCTIKVCPL NTQTLGIGCL TTDATTFEEV ATAEKLVITD VVDGVNHKLD VT TATCTIR NCKKLGPREN VAVIQVGGSD ILDITADPTT APQTERMMRI NWKKWWQVFY TVVDYVDQII QVMSKRSRSL NSA AFYYRV UniProtKB: Outer capsid glycoprotein VP7 |
-Macromolecule #3: Intermediate capsid protein VP6
| Macromolecule | Name: Intermediate capsid protein VP6 / type: protein_or_peptide / ID: 3 / Number of copies: 13 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Rotavirus A Rotavirus A |
| Molecular weight | Theoretical: 44.910738 KDa |
| Recombinant expression | Organism:  Chlorocebus aethiops aethiops (mammal) Chlorocebus aethiops aethiops (mammal) |
| Sequence | String: MDVLYSLSKT LKDARDKIVE GTLYSNVSDL IQQFNQMIIT MNGNEFQTGG IGNLPIRNWN FNFGLLGTTL LNLDANYVET ARNTIDYFV DFVDNVCMDE MVRESQRNGI APQSDSLRKL SAIKFKRINF DNSSEYIENW NLQNRRQRTG FTFHKPNIFP Y SASFTLNR ...String: MDVLYSLSKT LKDARDKIVE GTLYSNVSDL IQQFNQMIIT MNGNEFQTGG IGNLPIRNWN FNFGLLGTTL LNLDANYVET ARNTIDYFV DFVDNVCMDE MVRESQRNGI APQSDSLRKL SAIKFKRINF DNSSEYIENW NLQNRRQRTG FTFHKPNIFP Y SASFTLNR SQPAHDNLMG TMWLNAGSEI QVAGFDYSCA INAPANIQQF EHIVPLRRVL TTATITLLPD AERFSFPRVI NS ADGATTW FFNPVILRPN NVEVEFLLNG QIINTYQARF GTIVARNFDT IRLSFQLMRP PNMTPAVAVL FPNAQPFEHH ATV GLTLRI ESAVCESVLA DASETLLANV TSVRQEYAIP VGPVFPPGMN WTDLITNYSP SREDNLQRVF TVASIRSMLI K UniProtKB: Intermediate capsid protein VP6 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE |
| Details | MA104 cells infected with Rotavirus |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Detector mode: COUNTING / Average electron dose: 2.19 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 4.5 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: Warp (ver. 1.0.9) / Number subtomograms used: 16740 |
|---|---|
| Extraction | Number tomograms: 85 / Number images used: 16740 / Software - Name: crYOLO |
| Final 3D classification | Software - Name: RELION (ver. 3.0) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.0) |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8coa: |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)