+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | In vitro Nitrosopumilus maritimus S-layer with NH4Cl | |||||||||
 Map data Map data | PostProcessed map with B-factor sharpening | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Nmar_1547 S-layer / STRUCTURAL PROTEIN | |||||||||
| Function / homology | membrane / Uncharacterized protein Function and homology information Function and homology information | |||||||||
| Biological species |  Nitrosopumilus maritimus SCM1 (archaea) Nitrosopumilus maritimus SCM1 (archaea) | |||||||||
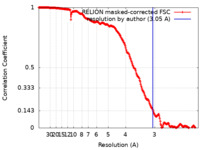
| Method | single particle reconstruction / cryo EM / Resolution: 3.05 Å | |||||||||
 Authors Authors | von Kuegelgen A / van Dorst S / Bharat TAM | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Membraneless channels sieve cations in ammonia-oxidizing marine archaea. Authors: Andriko von Kügelgen / C Keith Cassidy / Sofie van Dorst / Lennart L Pagani / Christopher Batters / Zephyr Ford / Jan Löwe / Vikram Alva / Phillip J Stansfeld / Tanmay A M Bharat /    Abstract: Nitrosopumilus maritimus is an ammonia-oxidizing archaeon that is crucial to the global nitrogen cycle. A critical step for nitrogen oxidation is the entrapment of ammonium ions from a dilute marine ...Nitrosopumilus maritimus is an ammonia-oxidizing archaeon that is crucial to the global nitrogen cycle. A critical step for nitrogen oxidation is the entrapment of ammonium ions from a dilute marine environment at the cell surface and their subsequent channelling to the cell membrane of N. maritimus. Here we elucidate the structure of the molecular machinery responsible for this process, comprising the surface layer (S-layer), using electron cryotomography and subtomogram averaging from cells. We supplemented our in situ structure of the ammonium-binding S-layer array with a single-particle electron cryomicroscopy structure, revealing detailed features of this immunoglobulin-rich and glycan-decorated S-layer. Biochemical analyses showed strong ammonium binding by the cell surface, which was lost after S-layer disassembly. Sensitive bioinformatic analyses identified similar S-layers in many ammonia-oxidizing archaea, with conserved sequence and structural characteristics. Moreover, molecular simulations and structure determination of ammonium-enriched specimens enabled us to examine the cation-binding properties of the S-layer, revealing how it concentrates ammonium ions on its cell-facing side, effectively acting as a multichannel sieve on the cell membrane. This in situ structural study illuminates the biogeochemically essential process of ammonium binding and channelling, common to many marine microorganisms that are fundamental to the nitrogen cycle. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16486.map.gz emd_16486.map.gz | 114 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16486-v30.xml emd-16486-v30.xml emd-16486.xml emd-16486.xml | 26.2 KB 26.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16486_fsc.xml emd_16486_fsc.xml | 22.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_16486.png emd_16486.png | 267.7 KB | ||
| Masks |  emd_16486_msk_1.map emd_16486_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16486.cif.gz emd-16486.cif.gz | 7.7 KB | ||
| Others |  emd_16486_additional_1.map.gz emd_16486_additional_1.map.gz emd_16486_half_map_1.map.gz emd_16486_half_map_1.map.gz emd_16486_half_map_2.map.gz emd_16486_half_map_2.map.gz | 112 MB 113.9 MB 113.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16486 http://ftp.pdbj.org/pub/emdb/structures/EMD-16486 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16486 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16486 | HTTPS FTP |
-Validation report
| Summary document |  emd_16486_validation.pdf.gz emd_16486_validation.pdf.gz | 1.4 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16486_full_validation.pdf.gz emd_16486_full_validation.pdf.gz | 1.4 MB | Display | |
| Data in XML |  emd_16486_validation.xml.gz emd_16486_validation.xml.gz | 22.4 KB | Display | |
| Data in CIF |  emd_16486_validation.cif.gz emd_16486_validation.cif.gz | 30.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16486 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16486 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16486 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16486 | HTTPS FTP |
-Related structure data
| Related structure data |  8c8kC  8c8lC  8c8mC  8c8nC  8c8oC  8c8rC C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16486.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16486.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PostProcessed map with B-factor sharpening | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.092 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16486_msk_1.map emd_16486_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
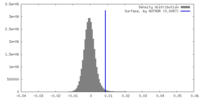
| Density Histograms |
-Additional map: Refine3D main map without B-factor sharpening
| File | emd_16486_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Refine3D main map without B-factor sharpening | ||||||||||||
| Projections & Slices |
| ||||||||||||
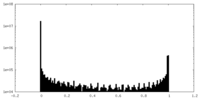
| Density Histograms |
-Half map: Half map 1
| File | emd_16486_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
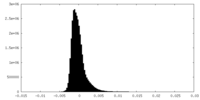
| Density Histograms |
-Half map: Half map 2
| File | emd_16486_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
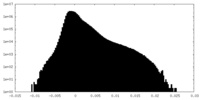
| Density Histograms |
- Sample components
Sample components
-Entire : Nitrosopumilus maritimus S-layer
| Entire | Name: Nitrosopumilus maritimus S-layer |
|---|---|
| Components |
|
-Supramolecule #1: Nitrosopumilus maritimus S-layer
| Supramolecule | Name: Nitrosopumilus maritimus S-layer / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all / Details: Nitrosopumilus maritimus S-layer C2 symmetrised |
|---|---|
| Source (natural) | Organism:  Nitrosopumilus maritimus SCM1 (archaea) / Strain: SCM1 / Location in cell: extracellular Nitrosopumilus maritimus SCM1 (archaea) / Strain: SCM1 / Location in cell: extracellular |
-Macromolecule #1: In vitro Nitrosopumilus maritimus S-layer with increased NH4Cl - ...
| Macromolecule | Name: In vitro Nitrosopumilus maritimus S-layer with increased NH4Cl - C2 symmetrised type: protein_or_peptide / ID: 1 / Details: NH4Cl (ammonium chloride) / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Nitrosopumilus maritimus SCM1 (archaea) / Strain: SCM1 Nitrosopumilus maritimus SCM1 (archaea) / Strain: SCM1 |
| Sequence | String: MNNEIGRKIT SLTLMTIMVA GGLTFAIPGV MPEAMAANAN LFVSAENSQF DNYMSGPQVI EVVVIDSDIN DTDEAKGEPD VTVNGKVLRM VQAVDGNWYG YFADRDQAQI ADSTATTADS GLDFGVFCAS SSGTAALGFS TTETDGIAIP ITIANATATG NGTQTGSSSG ...String: MNNEIGRKIT SLTLMTIMVA GGLTFAIPGV MPEAMAANAN LFVSAENSQF DNYMSGPQVI EVVVIDSDIN DTDEAKGEPD VTVNGKVLRM VQAVDGNWYG YFADRDQAQI ADSTATTADS GLDFGVFCAS SSGTAALGFS TTETDGIAIP ITIANATATG NGTQTGSSSG GAITTTCAAN TLDASTANGT INVVREAKDP VAASGSVSVG QIGLKNGTAN SGPNWPFIQL YELNPTGNVV VQYNKGGGVQ STTLTFDTVD QFAELSLDRT VFPRVSQVHA TITDLWLNID PTDEDSWTFA TNTKNTTSSF NVDTFYQVFD ENGASGGSAL TLRTTLSSLM CEDNCVLTLD VDAQSSGTPV VTIQDNGDSI LTQLNASSNT NANNASAFGI STETAKLGTG SIPVTITEQG PNSGVFGTYD ESDKSVLKIT DNAKRGTSAS LDYNETPQTI LVGFSFASID IQPVTDEWTS GQEIPVVIVD ADQNKNSRAD EDLDLNNPDV TLIPALRTGD PFTIDEGGTP SLIFTNGTNG DDSIFDTGAI NNTSAGQVGN FTLNINVTRF SSATNITSTE SIDTFSKRLI SAQTANSSAN FDVDFAIIDL GSATLETLKE TVVDEDNTAV GFNFFNYDVR SLGADTVSIA LLNTTGNILP WVNNDTRNVD KNNAILLVSN STNSQAYVDL TNAVSDAVYG STNTDSNVNI GFAMYFTGVG DLAAKEVIVM DFFSFGFTDD GVQSSERFAN QIIRIEAEET GDNTSTFEGS LEYVMVNQIN IQDAGTFSGI TPIADDPSFI VIEDLTDEDA PRVNYNDLGA DGVTTPVSDQ EEAPSHSGVV SLNADSYKIA DTVVITVEDL DLNVDSDLID IFTVVSDNSK ATDDAVGSAT TQSLSFGELG RLLDVTFDDV IWSTPDGANN TATGNDSDTC STELSNAGIT DTGLGATGFT LVETGAATGV FVGDFQIPSF WCRVSDTTTT PYTYAGDEET TTGLDIEVNY VDFRDASGEI VEVGDSAGVR ANTGSVSLDR TVYPVPFGTI ADSSKAANAA PNGRSVFPIH ATGITSTIDS TEELPTGDLT IHVRINDPDF DENPAGEDAM DQDNALKISV IRGSDSVVLG YAGASERTGK IDVGGNNGTI SNIRSFGEMD EIAPDAGIFE LDVNIKFTDG PASAQCNSHD TLYTALDGTT GKADTNRFDD GAASGQEYCI LQGDILQVEY TDPADASGDA NTVTDSATFD LRNGVLQSDK SVYIIGSDMI LTLIEPDFDL DNDSAETYDL DLIEWDSDAA TTTMGNKGVT GAAAAFDPEP TDFRETGDST GIFQIVIEIP ESLSNDKLER GEEIILEYTD WGPSGSDYVG DEDEDVNLTI YTSNFGATVE LDQKVYSWTD KVYITIVAPD HNFDSDLVDE IGETDSDPIK VSTRGFDLDN YKLVETGTDT GIFTGEVILT GFTAHDADGD GNTGDATGTT SGSGPTDGLL ATDDDDGLTV SFEFSEDETI VGSALIRWNI GEVQWLEASY PASGTGVVRV IDPDMNLDPE AVDNFEVDVW SDSDAGGIDL TVTETNEATG IFEGTVFFTT LDESSGHRLR VSEGDTVTAE YEDNTLPDPY TTADELDITA TSLIGTVVPP LERAPAANLR TVDAFGNSLD SVSVDQQVQI SADLANGQDR EQSFAYLVQI QDANGVTVSL AWITGSLSSG QSFSPALSWI PTEAGTYTAT AFVWESVDNP TALSPPVSTT VNVS UniProtKB: Uncharacterized protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
Details: 50 mM HEPES/NaOH pH=7.5, 500 mM NaCl, 50 mM MgCl2, 10 mM CaCl2, 2.5 mM NHCl4 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER/RHODIUM / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: AIR / Details: 15 mA | ||||||||||||||||||
| Vitrification | Cryogen name: NITROGEN / Chamber humidity: 100 % / Chamber temperature: 283.15 K / Instrument: FEI VITROBOT MARK IV Details: absorption for 60 sec and blotted for 5 sec with blot force -10. | ||||||||||||||||||
| Details | In vitro isolate S-layer cell envelopes |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 70.0 K / Max: 70.0 K |
| Specialist optics | Spherical aberration corrector: not used / Chromatic aberration corrector: not used / Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 3 / Number real images: 9435 / Average exposure time: 4.2 sec. / Average electron dose: 49.863 e/Å2 Details: collected over two sessions, one session with the stage tilted by 30 deg |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 5.0 µm / Calibrated defocus min: 2.0 µm / Calibrated magnification: 81000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: AB INITIO MODEL / Overall B value: 52.5 / Target criteria: Best Fit |
|---|
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)