+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the rod CNG channel bound to calmodulin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CNG channel / calmodulin / rod photoreceptor / vision / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnon-motile cilium membrane / intracellular cyclic nucleotide activated cation channel complex / intracellularly cGMP-activated cation channel activity / rod photoreceptor outer segment / intracellularly cAMP-activated cation channel activity / Inactivation, recovery and regulation of the phototransduction cascade / Activation of the phototransduction cascade / sensory perception of chemical stimulus / transmembrane transporter complex / retina homeostasis ...non-motile cilium membrane / intracellular cyclic nucleotide activated cation channel complex / intracellularly cGMP-activated cation channel activity / rod photoreceptor outer segment / intracellularly cAMP-activated cation channel activity / Inactivation, recovery and regulation of the phototransduction cascade / Activation of the phototransduction cascade / sensory perception of chemical stimulus / transmembrane transporter complex / retina homeostasis / photoreceptor outer segment membrane / sodium channel activity / sodium ion transport / cGMP binding / monoatomic cation transmembrane transport / molecular sequestering activity / photoreceptor outer segment / monoatomic cation transport / cAMP binding / visual perception / potassium ion transport / calcium channel activity / sensory perception of smell / calcium ion transport / molecular adaptor activity / positive regulation of gene expression / protein homodimerization activity / protein-containing complex / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
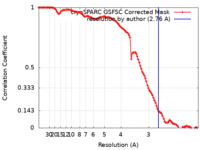
| Method | single particle reconstruction / cryo EM / Resolution: 2.76 Å | |||||||||
 Authors Authors | Marino J | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structural basis of calmodulin modulation of the rod cyclic nucleotide-gated channel. Authors: Diane C A Barret / Dina Schuster / Matthew J Rodrigues / Alexander Leitner / Paola Picotti / Gebhard F X Schertler / U Benjamin Kaupp / Volodymyr M Korkhov / Jacopo Marino /   Abstract: Calmodulin (CaM) regulates many ion channels to control calcium entry into cells, and mutations that alter this interaction are linked to fatal diseases. The structural basis of CaM regulation ...Calmodulin (CaM) regulates many ion channels to control calcium entry into cells, and mutations that alter this interaction are linked to fatal diseases. The structural basis of CaM regulation remains largely unexplored. In retinal photoreceptors, CaM binds to the CNGB subunit of cyclic nucleotide-gated (CNG) channels and, thereby, adjusts the channel's Cyclic guanosine monophosphate (cGMP) sensitivity in response to changes in ambient light conditions. Here, we provide the structural characterization for CaM regulation of a CNG channel by using a combination of single-particle cryo-electron microscopy and structural proteomics. CaM connects the CNGA and CNGB subunits, resulting in structural changes both in the cytosolic and transmembrane regions of the channel. Cross-linking and limited proteolysis-coupled mass spectrometry mapped the conformational changes induced by CaM in vitro and in the native membrane. We propose that CaM is a constitutive subunit of the rod channel to ensure high sensitivity in dim light. Our mass spectrometry-based approach is generally relevant for studying the effect of CaM on ion channels in tissues of medical interest, where only minute quantities are available. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16311.map.gz emd_16311.map.gz | 63.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16311-v30.xml emd-16311-v30.xml emd-16311.xml emd-16311.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16311_fsc.xml emd_16311_fsc.xml | 10.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_16311.png emd_16311.png | 53.4 KB | ||
| Masks |  emd_16311_msk_1.map emd_16311_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16311.cif.gz emd-16311.cif.gz | 6.7 KB | ||
| Others |  emd_16311_half_map_1.map.gz emd_16311_half_map_1.map.gz emd_16311_half_map_2.map.gz emd_16311_half_map_2.map.gz | 116.2 MB 116.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16311 http://ftp.pdbj.org/pub/emdb/structures/EMD-16311 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16311 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16311 | HTTPS FTP |
-Related structure data
| Related structure data |  8bx7MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16311.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16311.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.02 Å | ||||||||||||||||||||||||||||||||||||
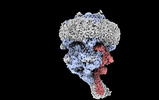
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16311_msk_1.map emd_16311_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_16311_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_16311_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Native CNG channel from retinal rods
| Entire | Name: Native CNG channel from retinal rods |
|---|---|
| Components |
|
-Supramolecule #1: Native CNG channel from retinal rods
| Supramolecule | Name: Native CNG channel from retinal rods / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 410 KDa |
-Macromolecule #1: Cyclic nucleotide-gated cation channel beta-1
| Macromolecule | Name: Cyclic nucleotide-gated cation channel beta-1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 155.228562 KDa |
| Sequence | String: MLGWVQRVLP QPPGTPQKTK QEEEGTEPEP ELEPKPETAP EETELEEVSL PPEEPCVGKE VAAVTLGPQG TQETALTPPT SLQAQVSVA PEAHSSPRGW VLTWLRKGVE KVVPQPAHSS RPSQNIAAGL ESPDQQAGAQ ILGQCGTGGS DEPSEPSRAE D PGPGPWLL ...String: MLGWVQRVLP QPPGTPQKTK QEEEGTEPEP ELEPKPETAP EETELEEVSL PPEEPCVGKE VAAVTLGPQG TQETALTPPT SLQAQVSVA PEAHSSPRGW VLTWLRKGVE KVVPQPAHSS RPSQNIAAGL ESPDQQAGAQ ILGQCGTGGS DEPSEPSRAE D PGPGPWLL RWFEQNLEKM LPQPPKISEG WRDEPTDAAL GPEPPGPALE IKPMLQAQES PSLPAPGPPE PEEEPIPEPQ PT IQASSLP PPQDSARLMA WILHRLEMAL PQPVIRGKGG EQESDAPVTC DVQTISILPG EQEESHLILE EVDPHWEEDE HQE GSTSTS PRTSEAAPAD EEKGKVVEQT PRELPRIQEE KEDEEEEKED GEEEEEEGRE KEEEEGEEKE EEEGREKEEE EGEK KEEEG REKEEEEGGE KEDEEGREKE EEEGRGKEEE EGGEKEEEEG RGKEEVEGRE EEEDEEEEQD HSVLLDSYLV PQSEE DRSE ESETQDQSEV GGAQAQGEVG GAQALSEESE TQDQSEVGGA QDQSEVGGAQ AQGEVGGAQE QDGVGGAQDQ STSHQE LQE EALADSSGVP ATEEHPELQV EDADADSRPL IAEENPPSPV QLPLSPAKSD TLAVPGSATG SLRKRLPSQD DEAEELK ML SPAASPVVAW SDPTSPQGTD DQDRATSTAS QNSAIINDRL QELVKLFKER TEKVKEKLID PDVTSDEESP KPSPAKKA P EPAPEVKPAE AGQVEEEHYC EMLCCKFKRR PWKKYQFPQS IDPLTNLMYI LWLFFVVLAW NWNCWLIPVR WAFPYQTPD NIHLWLLMDY LCDLIYLLDI TVFQMRLQFV RGGDIITDKK EMRNNYVKSQ RFKMDMLCLL PLDLLYLKFG VNPLLRLPRC LKYMAFFEF NNRLESILSK AYVYRVIRTT AYLLYSLHLN SCLYYWASAY EGLGSTHWVY DGVGNSYIRC YYWAVKTLIT I GGLPDPRT LFEIVFQGLN YFTGVFAFSV MIGQMRDVVG AATAGQTYYR SCMDSTVKYM NFYKIPRSVQ NRVKTWYEYT WH SQGMLDE SELMVQLPDK MRLDLAIDVN YSIVSKVALF QGCDRQMIFD MLKRLRSVVY LPNDYVCKKG EIGREMYIIQ AGQ VQVLGG PDGKSVLVTL KAGSVFGEIS LLAVGGGNRR TANVVAHGFT NLFILDKKDL NEILVHYPES QKLLRKKARR MLRN NNKPK EKSVLILPPR AGTPKLFNAA LAAAGKMGAK GGRGGRLALL RARLKELAAL EAAARQQQLL EQAKSSEDAA VGEEG SASP EQPPRPEPPA PEAPAPEPTA PEPLAPEAPA PEAPAPSSPP PASQERPEGD KDAARPEEHP VRIHVTLGPD PSEQIL LVE VPEKQEEKEK KEEETEEKEE GEEARKEKEE E UniProtKB: Cyclic nucleotide-gated channel beta-1 |
-Macromolecule #2: cGMP-gated cation channel alpha-1
| Macromolecule | Name: cGMP-gated cation channel alpha-1 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 79.712164 KDa |
| Sequence | String: MKKVIINTWH SFVNIPNVVG PDVEKEITRM ENGACSSFSG DDDDSASMFE ESETENPHAR DSFRSNTHGS GQPSQREQYL PGAIALFNV NNSSNKEQEP KEKKKKKKEK KSKPDDKNEN KKDPEKKKKK EKDKDKKKKE EKGKDKKEEE KKEVVVIDPS G NTYYNWLF ...String: MKKVIINTWH SFVNIPNVVG PDVEKEITRM ENGACSSFSG DDDDSASMFE ESETENPHAR DSFRSNTHGS GQPSQREQYL PGAIALFNV NNSSNKEQEP KEKKKKKKEK KSKPDDKNEN KKDPEKKKKK EKDKDKKKKE EKGKDKKEEE KKEVVVIDPS G NTYYNWLF CITLPVMYNW TMIIARACFD ELQSDYLEYW LAFDYLSDVV YLLDMFVRTR TGYLEQGLLV KEERKLIDKY KS TFQFKLD VLSVIPTDLL YIKFGWNYPE IRLNRLLRIS RMFEFFQRTE TRTNYPNIFR ISNLVMYIII IIHWNACVYF SIS KAIGFG NDTWVYPDVN DPDFGRLARK YVYSLYWSTL TLTTIGETPP PVRDSEYFFV VADFLIGVLI FATIVGNIGS MISN MNAAR AEFQARIDAI KQYMHFRNVS KDMEKRVIKW FDYLWTNKKT VDEREVLKYL PDKLRAEIAI NVHLDTLKKV RIFAD CEAG LLVELVLKLQ PQVYSPGDYI CKKGDIGREM YIIKEGKLAV VADDGITQFV VLSDGSYFGE ISILNIKGSK AGNRRT ANI KSIGYSDLFC LSKDDLMEAL TEYPDAKGML EEKGKQILMK DGLLDINIAN AGSDPKDLEE KVTRMESSVD LLQTRFA RI LAEYESMQQK LKQRLTKVEK FLKPLIDTEF SAIEGSGTES GPTDSTQD UniProtKB: Cyclic nucleotide-gated channel alpha-1 |
-Macromolecule #3: calmodulin-1
| Macromolecule | Name: calmodulin-1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.852545 KDa |
| Sequence | String: MADQLTEEQI AEFKEAFSLF DKDGDGTITT KELGTVMRSL GQNPTEAELQ DMINEVDADG NGTIDFPEFL TMMARKMKDT DSEEEIREA FRVFDKDGNG YISAAELRHV MTNLGEKLTD EEVDEMIREA DIDGDGQVNY EEFVQMMTAK UniProtKB: UNIPROTKB: A0A3Q7XW08 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 2 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 62.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)