+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of ARMC8-specific nanobody bound to CTLH-SR4 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CTLH / E3 ubiquitin ligase / LIGASE | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
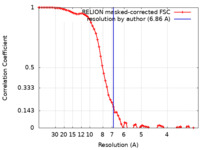
| Method | single particle reconstruction / cryo EM / Resolution: 6.86 Å | |||||||||
 Authors Authors | Chrustowicz J / Sherpa D | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2022 Journal: Elife / Year: 2022Title: Modular UBE2H-CTLH E2-E3 complexes regulate erythroid maturation. Authors: Dawafuti Sherpa / Judith Mueller / Özge Karayel / Peng Xu / Yu Yao / Jakub Chrustowicz / Karthik V Gottemukkala / Christine Baumann / Annette Gross / Oliver Czarnecki / Wei Zhang / Jun Gu / ...Authors: Dawafuti Sherpa / Judith Mueller / Özge Karayel / Peng Xu / Yu Yao / Jakub Chrustowicz / Karthik V Gottemukkala / Christine Baumann / Annette Gross / Oliver Czarnecki / Wei Zhang / Jun Gu / Johan Nilvebrant / Sachdev S Sidhu / Peter J Murray / Matthias Mann / Mitchell J Weiss / Brenda A Schulman / Arno F Alpi /     Abstract: The development of haematopoietic stem cells into mature erythrocytes - erythropoiesis - is a controlled process characterized by cellular reorganization and drastic reshaping of the proteome ...The development of haematopoietic stem cells into mature erythrocytes - erythropoiesis - is a controlled process characterized by cellular reorganization and drastic reshaping of the proteome landscape. Failure of ordered erythropoiesis is associated with anaemias and haematological malignancies. Although the ubiquitin system is a known crucial post-translational regulator in erythropoiesis, how the erythrocyte is reshaped by the ubiquitin system is poorly understood. By measuring the proteomic landscape of in vitro human erythropoiesis models, we found dynamic differential expression of subunits of the CTLH E3 ubiquitin ligase complex that formed maturation stage-dependent assemblies of topologically homologous RANBP9- and RANBP10-CTLH complexes. Moreover, protein abundance of CTLH's cognate E2 ubiquitin conjugating enzyme UBE2H increased during terminal differentiation, and UBE2H expression depended on catalytically active CTLH E3 complexes. CRISPR-Cas9-mediated inactivation of CTLH E3 assemblies or UBE2H in erythroid progenitors revealed defects, including spontaneous and accelerated erythroid maturation as well as inefficient enucleation. Thus, we propose that dynamic maturation stage-specific changes of UBE2H-CTLH E2-E3 modules control the orderly progression of human erythropoiesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16243.map.gz emd_16243.map.gz | 2.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16243-v30.xml emd-16243-v30.xml emd-16243.xml emd-16243.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16243_fsc.xml emd_16243_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_16243.png emd_16243.png | 37.8 KB | ||
| Masks |  emd_16243_msk_1.map emd_16243_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16243.cif.gz emd-16243.cif.gz | 4 KB | ||
| Others |  emd_16243_half_map_1.map.gz emd_16243_half_map_1.map.gz emd_16243_half_map_2.map.gz emd_16243_half_map_2.map.gz | 23.4 MB 23.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16243 http://ftp.pdbj.org/pub/emdb/structures/EMD-16243 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16243 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16243 | HTTPS FTP |
-Validation report
| Summary document |  emd_16243_validation.pdf.gz emd_16243_validation.pdf.gz | 604.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_16243_full_validation.pdf.gz emd_16243_full_validation.pdf.gz | 604.3 KB | Display | |
| Data in XML |  emd_16243_validation.xml.gz emd_16243_validation.xml.gz | 12.9 KB | Display | |
| Data in CIF |  emd_16243_validation.cif.gz emd_16243_validation.cif.gz | 17.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16243 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16243 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16243 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16243 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_16243.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16243.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.612 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16243_msk_1.map emd_16243_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_16243_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_16243_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CLTH-SR4 with ARMC8-specific nanobody
| Entire | Name: CLTH-SR4 with ARMC8-specific nanobody |
|---|---|
| Components |
|
-Supramolecule #1: CLTH-SR4 with ARMC8-specific nanobody
| Supramolecule | Name: CLTH-SR4 with ARMC8-specific nanobody / type: complex / ID: 1 / Parent: 0 Details: RANBP9, TWA1, ARMC8, RMND5A, MAEA, ARMC8-specific nanobody |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 69.6 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)