[English] 日本語
 Yorodumi
Yorodumi- EMDB-16090: Structure of a yeast 80S ribosome-bound N-Acetyltransferase B complex -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a yeast 80S ribosome-bound N-Acetyltransferase B complex | |||||||||
 Map data Map data | Homogenous Refinement of a class of 80S yeast ribosomes with two copies of the NatB complex. Half map. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | translation / n-acetylation / nascent chain / co-translational modification / NatB / RIBOSOME | |||||||||
| Function / homology |  Function and homology information Function and homology informationN-terminal methionine Nalpha-acetyltransferase NatB / NatB complex / mitochondrion inheritance / protein N-terminal-methionine acetyltransferase activity / protein-N-terminal amino-acid acetyltransferase activity / acetyltransferase activator activity / pre-mRNA 5'-splice site binding / cytosolic large ribosomal subunit assembly / response to cycloheximide / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) ...N-terminal methionine Nalpha-acetyltransferase NatB / NatB complex / mitochondrion inheritance / protein N-terminal-methionine acetyltransferase activity / protein-N-terminal amino-acid acetyltransferase activity / acetyltransferase activator activity / pre-mRNA 5'-splice site binding / cytosolic large ribosomal subunit assembly / response to cycloheximide / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / negative regulation of mRNA splicing, via spliceosome / preribosome, large subunit precursor / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / L13a-mediated translational silencing of Ceruloplasmin expression / translational elongation / ribosomal large subunit export from nucleus / translational termination / regulation of translational fidelity / protein-RNA complex assembly / maturation of LSU-rRNA / ERAD pathway / cytoskeleton organization / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / regulation of actin cytoskeleton organization / macroautophagy / translational initiation / maintenance of translational fidelity / modification-dependent protein catabolic process / protein tag activity / rRNA processing / ribosome biogenesis / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / negative regulation of translation / rRNA binding / structural constituent of ribosome / protein ubiquitination / ribosome / translation / response to antibiotic / mRNA binding / ubiquitin protein ligase binding / nucleolus / RNA binding / zinc ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Knorr AG / Mackens-Kiani T / Musial J / Berninghausen O / Becker T / Beatrix B / Beckmann R | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Biol / Year: 2023 Journal: PLoS Biol / Year: 2023Title: The dynamic architecture of Map1- and NatB-ribosome complexes coordinates the sequential modifications of nascent polypeptide chains. Authors: Alexandra G Knorr / Timur Mackens-Kiani / Joanna Musial / Otto Berninghausen / Thomas Becker / Birgitta Beatrix / Roland Beckmann /  Abstract: Cotranslational modification of the nascent polypeptide chain is one of the first events during the birth of a new protein. In eukaryotes, methionine aminopeptidases (MetAPs) cleave off the starter ...Cotranslational modification of the nascent polypeptide chain is one of the first events during the birth of a new protein. In eukaryotes, methionine aminopeptidases (MetAPs) cleave off the starter methionine, whereas N-acetyl-transferases (NATs) catalyze N-terminal acetylation. MetAPs and NATs compete with other cotranslationally acting chaperones, such as ribosome-associated complex (RAC), protein targeting and translocation factors (SRP and Sec61) for binding sites at the ribosomal tunnel exit. Yet, whereas well-resolved structures for ribosome-bound RAC, SRP and Sec61, are available, structural information on the mode of ribosome interaction of eukaryotic MetAPs or of the five cotranslationally active NATs is only available for NatA. Here, we present cryo-EM structures of yeast Map1 and NatB bound to ribosome-nascent chain complexes. Map1 is mainly associated with the dynamic rRNA expansion segment ES27a, thereby kept at an ideal position below the tunnel exit to act on the emerging substrate nascent chain. For NatB, we observe two copies of the NatB complex. NatB-1 binds directly below the tunnel exit, again involving ES27a, and NatB-2 is located below the second universal adapter site (eL31 and uL22). The binding mode of the two NatB complexes on the ribosome differs but overlaps with that of NatA and Map1, implying that NatB binds exclusively to the tunnel exit. We further observe that ES27a adopts distinct conformations when bound to NatA, NatB, or Map1, together suggesting a contribution to the coordination of a sequential activity of these factors on the emerging nascent chain at the ribosomal exit tunnel. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16090.map.gz emd_16090.map.gz | 122.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16090-v30.xml emd-16090-v30.xml emd-16090.xml emd-16090.xml | 65.4 KB 65.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16090_fsc.xml emd_16090_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_16090.png emd_16090.png | 130.8 KB | ||
| Filedesc metadata |  emd-16090.cif.gz emd-16090.cif.gz | 13.5 KB | ||
| Others |  emd_16090_half_map_1.map.gz emd_16090_half_map_1.map.gz emd_16090_half_map_2.map.gz emd_16090_half_map_2.map.gz | 226.8 MB 226.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16090 http://ftp.pdbj.org/pub/emdb/structures/EMD-16090 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16090 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16090 | HTTPS FTP |
-Related structure data
| Related structure data |  8bjqMC  8bipC  8bqdC  8bqxC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16090.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16090.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Homogenous Refinement of a class of 80S yeast ribosomes with two copies of the NatB complex. Half map. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.045 Å | ||||||||||||||||||||||||||||||||||||
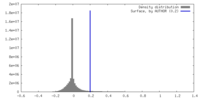
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Homogenous Refinement of a class of 80S yeast...
| File | emd_16090_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Homogenous Refinement of a class of 80S yeast ribosomes with two copies of the NatB complex. Half map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
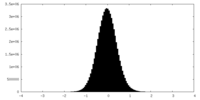
| Density Histograms |
-Half map: Homogenous Refinement of a class of 80S yeast...
| File | emd_16090_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Homogenous Refinement of a class of 80S yeast ribosomes with two copies of the NatB complex. Half map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
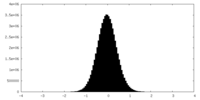
| Density Histograms |
- Sample components
Sample components
+Entire : Large ribosomal subunit of a yeast 80S ribosome with NatB-complex
+Supramolecule #1: Large ribosomal subunit of a yeast 80S ribosome with NatB-complex
+Macromolecule #1: N-terminal acetyltransferase B complex catalytic subunit NAT3
+Macromolecule #2: N-terminal acetyltransferase B complex subunit MDM20
+Macromolecule #5: 60S ribosomal protein L2-A
+Macromolecule #6: 60S ribosomal protein L3
+Macromolecule #7: 60S ribosomal protein L4-A
+Macromolecule #8: 60S ribosomal protein L5
+Macromolecule #9: 60S ribosomal protein L6-B
+Macromolecule #10: 60S ribosomal protein L7-A
+Macromolecule #11: 60S ribosomal protein L8-A
+Macromolecule #12: 60S ribosomal protein L9-A
+Macromolecule #13: 60S ribosomal protein L10
+Macromolecule #14: 60S ribosomal protein L11-B
+Macromolecule #15: 60S ribosomal protein L13-A
+Macromolecule #16: 60S ribosomal protein L14-A
+Macromolecule #17: 60S ribosomal protein L15-A
+Macromolecule #18: 60S ribosomal protein L16-A
+Macromolecule #19: 60S ribosomal protein L17-A
+Macromolecule #20: 60S ribosomal protein L18-A
+Macromolecule #21: 60S ribosomal protein L19-A
+Macromolecule #22: 60S ribosomal protein L20-A
+Macromolecule #23: 60S ribosomal protein L21-A
+Macromolecule #24: 60S ribosomal protein L22-A
+Macromolecule #25: 60S ribosomal protein L23-A
+Macromolecule #26: 60S ribosomal protein L24-A
+Macromolecule #27: 60S ribosomal protein L25
+Macromolecule #28: 60S ribosomal protein L26-A
+Macromolecule #29: 60S ribosomal protein L27-A
+Macromolecule #30: 60S ribosomal protein L28
+Macromolecule #31: 60S ribosomal protein L29
+Macromolecule #32: 60S ribosomal protein L30
+Macromolecule #33: 60S ribosomal protein L31-A
+Macromolecule #34: 60S ribosomal protein L32
+Macromolecule #35: 60S ribosomal protein L33-A
+Macromolecule #36: 60S ribosomal protein L34-A
+Macromolecule #37: 60S ribosomal protein L35-A
+Macromolecule #38: 60S ribosomal protein L36-A
+Macromolecule #39: 60S ribosomal protein L37-A
+Macromolecule #40: 60S ribosomal protein L38
+Macromolecule #41: 60S ribosomal protein L39
+Macromolecule #42: 60S ribosomal protein L40-A
+Macromolecule #43: 60S ribosomal protein L41-A
+Macromolecule #44: 60S ribosomal protein L42-A
+Macromolecule #45: 60S ribosomal protein L43-A
+Macromolecule #3: 5S rRNA
+Macromolecule #4: 5.8S rRNA
+Macromolecule #46: 25S rRNA
+Macromolecule #47: MAGNESIUM ION
+Macromolecule #48: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 45.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)