+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human apo TRPM8 in a closed state (composite map) | |||||||||
 Map data Map data | composite map created by combining consensus map with four copies of focused pre-MHR MHR1/2 map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | calcium ion channel / cold temperature sensor / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationthermoception / ligand-gated calcium channel activity / TRP channels / plasma membrane raft / response to cold / calcium ion transmembrane transport / calcium channel activity / intracellular calcium ion homeostasis / positive regulation of cold-induced thermogenesis / external side of plasma membrane ...thermoception / ligand-gated calcium channel activity / TRP channels / plasma membrane raft / response to cold / calcium ion transmembrane transport / calcium channel activity / intracellular calcium ion homeostasis / positive regulation of cold-induced thermogenesis / external side of plasma membrane / endoplasmic reticulum membrane / metal ion binding / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.65 Å | |||||||||
 Authors Authors | Palchevskyi S / Czarnocki-Cieciura M / Vistoli G / Gervasoni S / Nowak E / Beccari AR / Nowotny M / Talarico C | |||||||||
| Funding support |  Poland, 1 items Poland, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2023 Journal: Commun Biol / Year: 2023Title: Structure of human TRPM8 channel. Authors: Sergii Palchevskyi / Mariusz Czarnocki-Cieciura / Giulio Vistoli / Silvia Gervasoni / Elżbieta Nowak / Andrea R Beccari / Marcin Nowotny / Carmine Talarico /    Abstract: TRPM8 is a non-selective cation channel permeable to both monovalent and divalent cations that is activated by multiple factors, such as temperature, voltage, pressure, and changes in osmolality. It ...TRPM8 is a non-selective cation channel permeable to both monovalent and divalent cations that is activated by multiple factors, such as temperature, voltage, pressure, and changes in osmolality. It is a therapeutic target for anticancer drug development, and its modulators can be utilized for several pathological conditions. Here, we present a cryo-electron microscopy structure of a human TRPM8 channel in the closed state that was solved at 2.7 Å resolution. Our structure comprises the most complete model of the N-terminal pre-melastatin homology region. We also visualized several lipids that are bound by the protein and modeled how the human channel interacts with icilin. Analyses of pore helices in available TRPM structures showed that all these structures can be grouped into different closed, desensitized and open state conformations based on the register of the pore helix S6 which positions particular amino acid residues at the channel constriction. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15981.map.gz emd_15981.map.gz | 10.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15981-v30.xml emd-15981-v30.xml emd-15981.xml emd-15981.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15981.png emd_15981.png | 153.5 KB | ||
| Filedesc metadata |  emd-15981.cif.gz emd-15981.cif.gz | 7 KB | ||
| Others |  emd_15981_additional_1.map.gz emd_15981_additional_1.map.gz | 141.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15981 http://ftp.pdbj.org/pub/emdb/structures/EMD-15981 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15981 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15981 | HTTPS FTP |
-Related structure data
| Related structure data |  8bdcMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15981.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15981.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | composite map created by combining consensus map with four copies of focused pre-MHR MHR1/2 map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: raw consensus map
| File | emd_15981_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | raw consensus map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human TRPM8 in apo form
| Entire | Name: Human TRPM8 in apo form |
|---|---|
| Components |
|
-Supramolecule #1: Human TRPM8 in apo form
| Supramolecule | Name: Human TRPM8 in apo form / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 510 KDa |
-Macromolecule #1: Transient receptor potential cation channel subfamily M member 8
| Macromolecule | Name: Transient receptor potential cation channel subfamily M member 8 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 127.904719 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SNSFRAARLS MRNRRNDTLD STRTLYSSAS RSTDLSYSES DLVNFIQANF KKRECVFFTK DSKATENVCK CGYAQSQHME GTQINQSEK WNYKKHTKEF PTDAFGDIQF ETLGKKGKYI RLSCDTDAEI LYELLTQHWH LKTPNLVISV TGGAKNFALK P RMRKIFSR ...String: SNSFRAARLS MRNRRNDTLD STRTLYSSAS RSTDLSYSES DLVNFIQANF KKRECVFFTK DSKATENVCK CGYAQSQHME GTQINQSEK WNYKKHTKEF PTDAFGDIQF ETLGKKGKYI RLSCDTDAEI LYELLTQHWH LKTPNLVISV TGGAKNFALK P RMRKIFSR LIYIAQSKGA WILTGGTHYG LMKYIGEVVR DNTISRSSEE NIVAIGIAAW GMVSNRDTLI RNCDAEGYFL AQ YLMDDFT RDPLYILDNN HTHLLLVDNG CHGHPTVEAK LRNQLEKYIS ERTIQDSNYG GKIPIVCFAQ GGGKETLKAI NTS IKNKIP CVVVEGSGQI ADVIASLVEV EDALTSSAVK EKLVRFLPRT VSRLPEEETE SWIKWLKEIL ECSHLLTVIK MEEA GDEIV SNAISYALYK AFSTSEQDKD NWNGQLKLLL EWNQLDLAND EIFTNDRRWE SADLQEVMFT ALIKDRPKFV RLFLE NGLN LRKFLTHDVL TELFSNHFST LVYRNLQIAK NSYNDALLTF VWKLVANFRR GFRKEDRNGR DEMDIELHDV SPITRH PLQ ALFIWAILQN KKELSKVIWE QTRGCTLAAL GASKLLKTLA KVKNDINAAG ESEELANEYE TRAVELFTEC YSSDEDL AE QLLVYSCEAW GGSNCLELAV EATDQHFIAQ PGVQNFLSKQ WYGEISRDTK NWKIILCLFI IPLVGCGFVS FRKKPVDK H KKLLWYYVAF FTSPFVVFSW NVVFYIAFLL LFAYVLLMDF HSVPHPPELV LYSLVFVLFC DEVRQWYVNG VNYFTDLWN VMDTLGLFYF IAGIVFRLHS SNKSSLYSGR VIFCLDYIIF TLRLIHIFTV SRNLGPKIIM LQRMLIDVFF FLFLFAVWMV AFGVARQGI LRQNEQRWRW IFRSVIYEPY LAMFGQVPSD VDGTTYDFAH CTFTGNESKP LCVELDEHNL PRFPEWITIP L VCIYMLST NILLVNLLVA MFGYTVGTVQ ENNDQVWKFQ RYFLVQEYCS RLNIPFPFIV FAYFYMVVKK CFKCCCKEKN ME SSVCCFK NEDNETLAWE GVMKENYLVK INTKANDTSE EMRHRFRQLD TKLNDLKGLL KEIANKIK UniProtKB: Transient receptor potential cation channel subfamily M member 8 |
-Macromolecule #2: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 2 / Number of copies: 8 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #3: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 3 / Number of copies: 4 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Macromolecule #4: UNDECANE
| Macromolecule | Name: UNDECANE / type: ligand / ID: 4 / Number of copies: 4 / Formula: UND |
|---|---|
| Molecular weight | Theoretical: 156.308 Da |
| Chemical component information | 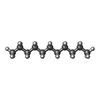 ChemComp-UND: |
-Macromolecule #5: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 5 / Number of copies: 8 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE Details: The grid was glow-discharged from both sides prior to vitrification |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | protein solubilized by LMNG |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 7918 / Average electron dose: 41.42 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8bdc: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)