[English] 日本語
 Yorodumi
Yorodumi- EMDB-15943: CryoEM reconstruction of GroEL-GroES-ADP.AlF3-Rubisco, class I. -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM reconstruction of GroEL-GroES-ADP.AlF3-Rubisco, class I. | |||||||||
 Map data Map data | CryoEM reconstruction of GroEL-GroES-Rubisco. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GroEL / GroES / Chaperone | |||||||||
| Biological species |  | |||||||||
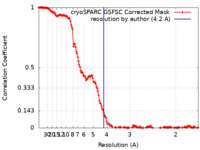
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Gardner S / Saibil HR | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structural basis of substrate progression through the bacterial chaperonin cycle. Authors: Scott Gardner / Michele C Darrow / Natalya Lukoyanova / Konstantinos Thalassinos / Helen R Saibil /  Abstract: The bacterial chaperonin GroEL-GroES promotes protein folding through ATP-regulated cycles of substrate protein binding, encapsulation, and release. Here, we have used cryoEM to determine structures ...The bacterial chaperonin GroEL-GroES promotes protein folding through ATP-regulated cycles of substrate protein binding, encapsulation, and release. Here, we have used cryoEM to determine structures of GroEL, GroEL-ADP·BeF, and GroEL-ADP·AlF-GroES all complexed with the model substrate Rubisco. Our structures provide a series of snapshots that show how the conformation and interactions of non-native Rubisco change as it proceeds through the GroEL-GroES reaction cycle. We observe specific charged and hydrophobic GroEL residues forming strong initial contacts with non-native Rubisco. Binding of ATP or ADP·BeF to GroEL-Rubisco results in the formation of an intermediate GroEL complex displaying striking asymmetry in the ATP/ADP·BeF-bound ring. In this ring, four GroEL subunits bind Rubisco and the other three are in the GroES-accepting conformation, suggesting how GroEL can recruit GroES without releasing bound substrate. Our cryoEM structures of stalled GroEL-ADP·AlF-Rubisco-GroES complexes show Rubisco folding intermediates interacting with GroEL-GroES via different sets of residues. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15943.map.gz emd_15943.map.gz | 20.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15943-v30.xml emd-15943-v30.xml emd-15943.xml emd-15943.xml | 14.3 KB 14.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15943_fsc.xml emd_15943_fsc.xml | 14.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_15943.png emd_15943.png | 96.5 KB | ||
| Masks |  emd_15943_msk_1.map emd_15943_msk_1.map | 343 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15943.cif.gz emd-15943.cif.gz | 4 KB | ||
| Others |  emd_15943_half_map_1.map.gz emd_15943_half_map_1.map.gz emd_15943_half_map_2.map.gz emd_15943_half_map_2.map.gz | 318.7 MB 318.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15943 http://ftp.pdbj.org/pub/emdb/structures/EMD-15943 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15943 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15943 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15943.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15943.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM reconstruction of GroEL-GroES-Rubisco. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.84938 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15943_msk_1.map emd_15943_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: CryoEM reconstruction of GroEL-GroES-Rubisco.
| File | emd_15943_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM reconstruction of GroEL-GroES-Rubisco. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: CryoEM reconstruction of GroEL-GroES-Rubisco.
| File | emd_15943_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM reconstruction of GroEL-GroES-Rubisco. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GroEL-GroES-ADP.AlF3-Rubisco
| Entire | Name: GroEL-GroES-ADP.AlF3-Rubisco |
|---|---|
| Components |
|
-Supramolecule #1: GroEL-GroES-ADP.AlF3-Rubisco
| Supramolecule | Name: GroEL-GroES-ADP.AlF3-Rubisco / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.7 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 293 K / Instrument: SPOTITON Details: The grid was prepared using a chameleon (SPT Labtech).. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 2 / Average electron dose: 40.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.4000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)