[English] 日本語
 Yorodumi
Yorodumi- EMDB-1581: Structure and functional role of dynein's microtubule-binding domain -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1581 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure and functional role of dynein's microtubule-binding domain | |||||||||
 Map data Map data | This is a 3-D map of the SRS-MTBD construct | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Assembly 1 is a helical 15 protofilaments / component name / tubulin. Assembly 2 is a monomer of the SRS-MTBD construct | |||||||||
| Biological species |  | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 35.0 Å | |||||||||
 Authors Authors | Carter AP / Garbarino JE / Wilson-Kubalek EM / Shipley WE / Cho C / Milligan RA / Vale RD / Gibbons IR | |||||||||
 Citation Citation |  Journal: Science / Year: 2008 Journal: Science / Year: 2008Title: Structure and functional role of dynein's microtubule-binding domain. Authors: Andrew P Carter / Joan E Garbarino / Elizabeth M Wilson-Kubalek / Wesley E Shipley / Carol Cho / Ronald A Milligan / Ronald D Vale / I R Gibbons /  Abstract: Dynein motors move various cargos along microtubules within the cytoplasm and power the beating of cilia and flagella. An unusual feature of dynein is that its microtubule-binding domain (MTBD) is ...Dynein motors move various cargos along microtubules within the cytoplasm and power the beating of cilia and flagella. An unusual feature of dynein is that its microtubule-binding domain (MTBD) is separated from its ring-shaped AAA+ adenosine triphosphatase (ATPase) domain by a 15-nanometer coiled-coil stalk. We report the crystal structure of the mouse cytoplasmic dynein MTBD and a portion of the coiled coil, which supports a mechanism by which the ATPase domain and MTBD may communicate through a shift in the heptad registry of the coiled coil. Surprisingly, functional data suggest that the MTBD, and not the ATPase domain, is the main determinant of the direction of dynein motility. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1581.map.gz emd_1581.map.gz | 1.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1581-v30.xml emd-1581-v30.xml emd-1581.xml emd-1581.xml | 9.7 KB 9.7 KB | Display Display |  EMDB header EMDB header |
| Images |  1581.gif 1581.gif | 103.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1581 http://ftp.pdbj.org/pub/emdb/structures/EMD-1581 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1581 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1581 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1581.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1581.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is a 3-D map of the SRS-MTBD construct | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
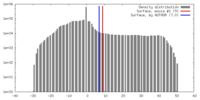
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Synthetic construct of dynein microtubule-binding domain (85-82) ...
| Entire | Name: Synthetic construct of dynein microtubule-binding domain (85-82) fused to seryl-tRNA synthase-monomer. Abbreviated name is SRS-MTBD-85-82 |
|---|---|
| Components |
|
-Supramolecule #1000: Synthetic construct of dynein microtubule-binding domain (85-82) ...
| Supramolecule | Name: Synthetic construct of dynein microtubule-binding domain (85-82) fused to seryl-tRNA synthase-monomer. Abbreviated name is SRS-MTBD-85-82 type: sample / ID: 1000 Details: The SRS-MTBD-85-82 construct has a 12 heptad long stalk, only the first 3 heptad repeats were visible in this map. No density was observed for the SRS. Oligomeric state: SRS-MTBD-85-82 monomers bound to 15 protofilaments helical microtubules Number unique components: 2 |
|---|
-Macromolecule #1: microtubule
| Macromolecule | Name: microtubule / type: protein_or_peptide / ID: 1 / Name.synonym: microtubule / Oligomeric state: monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 4 K / Instrument: OTHER / Details: Vitrification instrument: Vitrobot / Method: 1.5 sec blot |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Specialist optics | Energy filter - Name:  Field Emission Gun Field Emission Gun |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Sampling interval: 3.5 µm / Number real images: 10 / Average electron dose: 10 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal magnification: 29000 |
| Sample stage | Specimen holder: eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 35.0 Å / Resolution method: OTHER / Software - Name: phoelix |
|---|---|
| CTF correction | Details: each image |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Details | PDBEntryID_givenInChain. Protocol: Rigid body. The crystal structure was manually docked into the EM density using the chimera software package. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)