[English] 日本語
 Yorodumi
Yorodumi- EMDB-15727: Poliovirus type 2 (strain MEF-1) virus-like particle in complex w... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Poliovirus type 2 (strain MEF-1) virus-like particle in complex with capsid binder compound 17 | |||||||||
 Map data Map data | Local resolution scaled final map from RELION. Sharpened with an ad-hoc B-factor of -15A^2. Contour level determined in UCSF ChimeraX. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Capsid protein / VIRUS LIKE PARTICLE / inhibitor / complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / receptor-mediated endocytosis of virus by host cell / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane ...symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / receptor-mediated endocytosis of virus by host cell / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / ribonucleoside triphosphate phosphatase activity / nucleoside-triphosphate phosphatase / channel activity / monoatomic ion transmembrane transport / DNA replication / RNA helicase activity / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / viral RNA genome replication / RNA-directed RNA polymerase activity / DNA-templated transcription / virion attachment to host cell / host cell nucleus / structural molecule activity / proteolysis / RNA binding / zinc ion binding / ATP binding Similarity search - Function | |||||||||
| Biological species |  Human poliovirus 2 Human poliovirus 2 | |||||||||
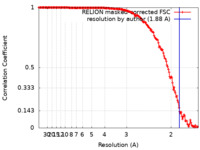
| Method | single particle reconstruction / cryo EM / Resolution: 1.88 Å | |||||||||
 Authors Authors | Bahar MW / Fry EE / Stuart DI | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2022 Journal: Commun Biol / Year: 2022Title: A conserved glutathione binding site in poliovirus is a target for antivirals and vaccine stabilisation. Authors: Mohammad W Bahar / Veronica Nasta / Helen Fox / Lee Sherry / Keith Grehan / Claudine Porta / Andrew J Macadam / Nicola J Stonehouse / David J Rowlands / Elizabeth E Fry / David I Stuart /   Abstract: Strategies to prevent the recurrence of poliovirus (PV) after eradication may utilise non-infectious, recombinant virus-like particle (VLP) vaccines. Despite clear advantages over inactivated or ...Strategies to prevent the recurrence of poliovirus (PV) after eradication may utilise non-infectious, recombinant virus-like particle (VLP) vaccines. Despite clear advantages over inactivated or attenuated virus vaccines, instability of VLPs can compromise their immunogenicity. Glutathione (GSH), an important cellular reducing agent, is a crucial co-factor for the morphogenesis of enteroviruses, including PV. We report cryo-EM structures of GSH bound to PV serotype 3 VLPs showing that it can enhance particle stability. GSH binds the positively charged pocket at the interprotomer interface shown recently to bind GSH in enterovirus F3 and putative antiviral benzene sulphonamide compounds in other enteroviruses. We show, using high-resolution cryo-EM, the binding of a benzene sulphonamide compound with a PV serotype 2 VLP, consistent with antiviral activity through over-stabilizing the interprotomer pocket, preventing the capsid rearrangements necessary for viral infection. Collectively, these results suggest GSH or an analogous tight-binding antiviral offers the potential for stabilizing VLP vaccines. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15727.map.gz emd_15727.map.gz | 210 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15727-v30.xml emd-15727-v30.xml emd-15727.xml emd-15727.xml | 25.9 KB 25.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15727_fsc.xml emd_15727_fsc.xml | 15.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_15727.png emd_15727.png | 98.8 KB | ||
| Filedesc metadata |  emd-15727.cif.gz emd-15727.cif.gz | 8.1 KB | ||
| Others |  emd_15727_half_map_1.map.gz emd_15727_half_map_1.map.gz emd_15727_half_map_2.map.gz emd_15727_half_map_2.map.gz | 276.4 MB 276.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15727 http://ftp.pdbj.org/pub/emdb/structures/EMD-15727 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15727 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15727 | HTTPS FTP |
-Related structure data
| Related structure data |  8ayzMC  8ayxC  8ayyC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15727.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15727.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local resolution scaled final map from RELION. Sharpened with an ad-hoc B-factor of -15A^2. Contour level determined in UCSF ChimeraX. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.829 Å | ||||||||||||||||||||||||||||||||||||
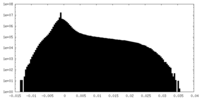
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map from final refinement in RELION. Contour...
| File | emd_15727_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map from final refinement in RELION. Contour level determined in UCSF ChimeraX. | ||||||||||||
| Projections & Slices |
| ||||||||||||
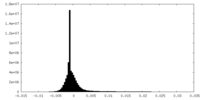
| Density Histograms |
-Half map: Half map from final refinement in RELION. Contour...
| File | emd_15727_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map from final refinement in RELION. Contour level determined in UCSF ChimeraX. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human poliovirus 2
| Entire | Name:  Human poliovirus 2 Human poliovirus 2 |
|---|---|
| Components |
|
-Supramolecule #1: Human poliovirus 2
| Supramolecule | Name: Human poliovirus 2 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: Recombinantly expressed virus-like particle of PV2 (strain MEF-1) NCBI-ID: 12083 / Sci species name: Human poliovirus 2 / Sci species strain: MEF-1 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  |
| Molecular weight | Theoretical: 5.81 MDa |
| Virus shell | Shell ID: 1 / Name: Virus shell 1 / Diameter: 310.0 Å / T number (triangulation number): 1 |
-Macromolecule #1: Capsid protein, VP1
| Macromolecule | Name: Capsid protein, VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 2 / Strain: MEF-1 Human poliovirus 2 / Strain: MEF-1 |
| Molecular weight | Theoretical: 33.111207 KDa |
| Recombinant expression | Organism:  Mesocricetus auratus (golden hamster) Mesocricetus auratus (golden hamster) |
| Sequence | String: GLGDLIEGVV EGVTRNALTP LTPANNLPDT QSSGPAHSKE TPALTAVETG ATNPLVPSDT VQTRHVIQKR TRSESTVESF FARGACVAI IEVDNDAPTK RASKLFSVWK ITYKDTVQLR RKLEFFTYSR FDMEFTFVVT SNYTDANNGH ALNQVYQIMY I PPGAPIPG ...String: GLGDLIEGVV EGVTRNALTP LTPANNLPDT QSSGPAHSKE TPALTAVETG ATNPLVPSDT VQTRHVIQKR TRSESTVESF FARGACVAI IEVDNDAPTK RASKLFSVWK ITYKDTVQLR RKLEFFTYSR FDMEFTFVVT SNYTDANNGH ALNQVYQIMY I PPGAPIPG KWNDYTWQTS SNPSVFYTYG APPARISVPY VGIANAYSHF YDGFAKVPLA GQASTEGDSL YGAASLNDFG SL AVRVVND HNPTKLTSKI RVYMKPKHVR VWCPRPPRAV PYYGPGVDYK DGLAPLPEKG LTTY UniProtKB: Genome polyprotein |
-Macromolecule #2: Capsid protein, VP0
| Macromolecule | Name: Capsid protein, VP0 / type: protein_or_peptide / ID: 2 / Details: Sequence is given for the VP0 polypeptide. / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 2 / Strain: MEF-1 Human poliovirus 2 / Strain: MEF-1 |
| Molecular weight | Theoretical: 37.456855 KDa |
| Recombinant expression | Organism:  Mesocricetus auratus (golden hamster) Mesocricetus auratus (golden hamster) |
| Sequence | String: MGAQVSSQKV GAHENSNRAY GGSTINYTTI NYYRDSASNA ASKQDFAQDP SKFTEPIKDV LIKTAPTLNS PNIEACGYSD RVMQLTLGN STITTQEAAN SVVAYGRWPE YIKDSEANPV DQPTEPDVAA CRFYTLDTVT WRKESRGWWW KLPDALKDMG L FGQNMFYH ...String: MGAQVSSQKV GAHENSNRAY GGSTINYTTI NYYRDSASNA ASKQDFAQDP SKFTEPIKDV LIKTAPTLNS PNIEACGYSD RVMQLTLGN STITTQEAAN SVVAYGRWPE YIKDSEANPV DQPTEPDVAA CRFYTLDTVT WRKESRGWWW KLPDALKDMG L FGQNMFYH YLGRAGYTVH VQCNASKFHQ GALGVFAVPE MCLAGDSTTH MFTKYENANP GEKGGEFKGS FTLDTNATNP AR NFCPVDY LFGSGVLAGN AFVYPHQIIN LRTNNCATLV LPYVNSLSID SMTKHNNWGI AILPLAPLDF ATESSTEIPI TLT IAPMCC EFNGLRNITV PRTQ UniProtKB: Genome polyprotein |
-Macromolecule #3: Capsid protein, VP3
| Macromolecule | Name: Capsid protein, VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 2 / Strain: MEF-1 Human poliovirus 2 / Strain: MEF-1 |
| Molecular weight | Theoretical: 26.472293 KDa |
| Recombinant expression | Organism:  Mesocricetus auratus (golden hamster) Mesocricetus auratus (golden hamster) |
| Sequence | String: GLPVLNTPGS NQYLTADNYQ SPCAIPEFDV TPPIDIPGEV RNMMELAEID TMIPLNLTNQ RKNTMDMYRV ELNDAAHSDT PILCLSLSP ASDPRLAHTM LGEILNYYTH WAGSLKFTFL FCGSMMATGK LLVSYAPPGA EAPKSRKEAM LGTHVIWDIG L QSSCTMVV ...String: GLPVLNTPGS NQYLTADNYQ SPCAIPEFDV TPPIDIPGEV RNMMELAEID TMIPLNLTNQ RKNTMDMYRV ELNDAAHSDT PILCLSLSP ASDPRLAHTM LGEILNYYTH WAGSLKFTFL FCGSMMATGK LLVSYAPPGA EAPKSRKEAM LGTHVIWDIG L QSSCTMVV PWISNTTYRQ TINDSFTEGG YISMFYQTRV VVPLSTPRKM DILGFVSACN DFSVRLLRDT THISQEAMPQ UniProtKB: Genome polyprotein |
-Macromolecule #4: SPHINGOSINE
| Macromolecule | Name: SPHINGOSINE / type: ligand / ID: 4 / Number of copies: 1 / Formula: SPH |
|---|---|
| Molecular weight | Theoretical: 299.492 Da |
| Chemical component information |  ChemComp-SPH: |
-Macromolecule #5: 4-[[4-[1,3-bis(oxidanylidene)isoindol-2-yl]phenyl]sulfonylamino]b...
| Macromolecule | Name: 4-[[4-[1,3-bis(oxidanylidene)isoindol-2-yl]phenyl]sulfonylamino]benzoic acid type: ligand / ID: 5 / Number of copies: 1 / Formula: FHK |
|---|---|
| Molecular weight | Theoretical: 422.411 Da |
| Chemical component information | 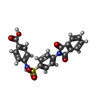 ChemComp-FHK: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 262 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.75 mg/mL | ||||||
|---|---|---|---|---|---|---|---|
| Buffer | pH: 7 / Component:
Details: 1 x DPBS was made from tissue culture grade Dulbecco's Phosphate Buffered Saline (Sigma-Aldrich). 20 mM EDTA was prepared from fresh stocks of buffered EDTA. | ||||||
| Grid | Model: EMS Lacey Carbon / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: LACEY / Support film - Film thickness: 3 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR Details: The specific type of grid used was Ultra-thin carbon support film, 3nm - on lacey carbon AGS187-4 from Agar Scientific. | ||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: Single blotting with 3.5ul of sample. Four second blot time and -17 blot force on the vitrobot.. | ||||||
| Details | Recombinant VLP of PV2 mixed with capsid inhibitor compound 17 at a molar ratio of 1:2500. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 4750 / Average exposure time: 1.7 sec. / Average electron dose: 34.68 e/Å2 Details: Images were collected in electron counting mode with a calibrated pixel size of 0.829 A/pixel. Super-resolution pixel size 0.4145 A/pixel. Movies were collected in 50 frames. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 60314 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | Initial model was rigid body fitted using UCSF chimera and Coot. Global minimization and B-factor refinement was performed in real space using phenix_real.space.refine. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Correlation coefficient |
| Output model |  PDB-8ayz: |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)