+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the type I-G CRISPR effector | |||||||||
 Map data Map data | Main map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CRISPR effector / IMMUNE SYSTEM | |||||||||
| Function / homology | CRISPR-associated protein GSU0053 / CRISPR-associated protein GSU0053 (Cas_GSU0053) / Type I-U CRISPR-associated protein Cas7 Function and homology information Function and homology information | |||||||||
| Biological species |  Thioalkalivibrio sulfidiphilus HL-EbGr7 (bacteria) / synthetic construct (others) Thioalkalivibrio sulfidiphilus HL-EbGr7 (bacteria) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Shangguan Q / Graham S / Sundaramoorthy R / White MF | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2022 Journal: Nucleic Acids Res / Year: 2022Title: Structure and mechanism of the type I-G CRISPR effector. Authors: Qilin Shangguan / Shirley Graham / Ramasubramanian Sundaramoorthy / Malcolm F White /  Abstract: Type I CRISPR systems are the most common CRISPR type found in bacteria. They use a multisubunit effector, guided by crRNA, to detect and bind dsDNA targets, forming an R-loop and recruiting the Cas3 ...Type I CRISPR systems are the most common CRISPR type found in bacteria. They use a multisubunit effector, guided by crRNA, to detect and bind dsDNA targets, forming an R-loop and recruiting the Cas3 enzyme to facilitate target DNA destruction, thus providing immunity against mobile genetic elements. Subtypes have been classified into families A-G, with type I-G being the least well understood. Here, we report the composition, structure and function of the type I-G Cascade CRISPR effector from Thioalkalivibrio sulfidiphilus, revealing key new molecular details. The unique Csb2 subunit processes pre-crRNA, remaining bound to the 3' end of the mature crRNA, and seven Cas7 subunits form the backbone of the effector. Cas3 associates stably with the effector complex via the Cas8g subunit and is important for target DNA recognition. Structural analysis by cryo-Electron Microscopy reveals a strikingly curved backbone conformation with Cas8g spanning the belly of the structure. These biochemical and structural insights shed new light on the diversity of type I systems and open the way to applications in genome engineering. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15540.map.gz emd_15540.map.gz | 11.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15540-v30.xml emd-15540-v30.xml emd-15540.xml emd-15540.xml | 31.5 KB 31.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15540.png emd_15540.png | 30.4 KB | ||
| Filedesc metadata |  emd-15540.cif.gz emd-15540.cif.gz | 6.5 KB | ||
| Others |  emd_15540_additional_1.map.gz emd_15540_additional_1.map.gz emd_15540_additional_2.map.gz emd_15540_additional_2.map.gz emd_15540_additional_3.map.gz emd_15540_additional_3.map.gz emd_15540_additional_4.map.gz emd_15540_additional_4.map.gz emd_15540_additional_5.map.gz emd_15540_additional_5.map.gz emd_15540_additional_6.map.gz emd_15540_additional_6.map.gz emd_15540_half_map_1.map.gz emd_15540_half_map_1.map.gz emd_15540_half_map_2.map.gz emd_15540_half_map_2.map.gz | 128.2 MB 118.4 MB 120.4 MB 128.4 MB 120.4 MB 128.6 MB 129.8 MB 129.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15540 http://ftp.pdbj.org/pub/emdb/structures/EMD-15540 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15540 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15540 | HTTPS FTP |
-Validation report
| Summary document |  emd_15540_validation.pdf.gz emd_15540_validation.pdf.gz | 728 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15540_full_validation.pdf.gz emd_15540_full_validation.pdf.gz | 727.6 KB | Display | |
| Data in XML |  emd_15540_validation.xml.gz emd_15540_validation.xml.gz | 14.5 KB | Display | |
| Data in CIF |  emd_15540_validation.cif.gz emd_15540_validation.cif.gz | 17.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15540 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15540 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15540 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15540 | HTTPS FTP |
-Related structure data
| Related structure data |  8aneMC  8b2xC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15540.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15540.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.008 Å | ||||||||||||||||||||||||||||||||||||
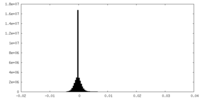
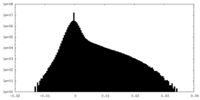
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: First body of Multibody refinement
| File | emd_15540_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | First body of Multibody refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Second body of Multibody refinement
| File | emd_15540_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Second body of Multibody refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Half map of body2 of Multibody
| File | emd_15540_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of body2 of Multibody | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #2
| File | emd_15540_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_15540_additional_5.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Half map of body1 of Multibody
| File | emd_15540_additional_6.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of body1 of Multibody | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map for the main map
| File | emd_15540_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map for the main map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map for the main map
| File | emd_15540_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map for the main map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Type IG cascade core cas7 with CrRNA
| Entire | Name: Type IG cascade core cas7 with CrRNA |
|---|---|
| Components |
|
-Supramolecule #1: Type IG cascade core cas7 with CrRNA
| Supramolecule | Name: Type IG cascade core cas7 with CrRNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Thioalkalivibrio sulfidiphilus HL-EbGr7 (bacteria) Thioalkalivibrio sulfidiphilus HL-EbGr7 (bacteria) |
| Molecular weight | Theoretical: 35 KDa |
-Macromolecule #1: Cas7
| Macromolecule | Name: Cas7 / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Thioalkalivibrio sulfidiphilus HL-EbGr7 (bacteria) Thioalkalivibrio sulfidiphilus HL-EbGr7 (bacteria)Strain: HL-EbGR7 |
| Molecular weight | Theoretical: 34.833082 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKLEPLLSDV PRLLMEADLV PVQGTRFQPT GFPDLGAAHY EGPDGRPMLL VESAQSMANR LETVCWDKDA DDWVVPLRGL PVVKVLDKA GKPLTNSVLE AHRLNSPYIL EGKDKTLFDL LKQELAHMEE GPVDIRKLAE TLLKVDANAV LHGVFLAKKE L AGGRLRLP ...String: MKLEPLLSDV PRLLMEADLV PVQGTRFQPT GFPDLGAAHY EGPDGRPMLL VESAQSMANR LETVCWDKDA DDWVVPLRGL PVVKVLDKA GKPLTNSVLE AHRLNSPYIL EGKDKTLFDL LKQELAHMEE GPVDIRKLAE TLLKVDANAV LHGVFLAKKE L AGGRLRLP RALSAFIEAE DVRVASSGGV KNDHVNPSGD TSRGFGNVPF ARDEYVSPRI KAYFNLDLAQ IRAFGLGEQV DR LLIALAL YKVRRFLVHG LRLRTACDLD CQALRVTRPE GWEVPELSEL EAALPGLIEA VAGEGRFAQP AVTIVTYEK UniProtKB: Type I-U CRISPR-associated protein Cas7 |
-Macromolecule #2: RNA (66-MER)
| Macromolecule | Name: RNA (66-MER) / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 21.082465 KDa |
| Sequence | String: AUUGAAGCAA GCUGUCCCUG AUGGUCGUCA UCUACCUGCC UGGAGUCAUC CGCGGCAUUU AGCCGC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 20 mM Tris-HCl, 250 mM NaCl, pH 7.5 |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 400 / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.0001 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 4 K / Instrument: FEI VITROBOT MARK IV / Details: blot time 3.5 blot force 4. |
| Details | Size exclusion purified monodisperse sample in solution |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77.0 K / Max: 183.0 K |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5000 pixel / Digitization - Dimensions - Height: 4000 pixel / Number grids imaged: 1 / Number real images: 15710 / Average exposure time: 3.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 2.8000000000000003 µm / Calibrated defocus min: 1.6 µm / Calibrated magnification: 105000 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.6 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 188 / Target criteria: correlation coefficient |
|---|---|
| Output model |  PDB-8ane: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)