+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Yeast RQC complex in state D | |||||||||
 Map data Map data | Yeast RQC complex in state D, half map B | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribosome-associated quality control / NEMF / Listerin / CAT tailing / TRANSLATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationRQC complex / ribophagy / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosome-associated ubiquitin-dependent protein catabolic process / pre-mRNA 5'-splice site binding / cytosolic large ribosomal subunit assembly / maturation of 5.8S rRNA / response to cycloheximide / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / SRP-dependent cotranslational protein targeting to membrane ...RQC complex / ribophagy / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosome-associated ubiquitin-dependent protein catabolic process / pre-mRNA 5'-splice site binding / cytosolic large ribosomal subunit assembly / maturation of 5.8S rRNA / response to cycloheximide / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / ribosomal large subunit binding / negative regulation of mRNA splicing, via spliceosome / preribosome, large subunit precursor / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Antigen processing: Ubiquitination & Proteasome degradation / L13a-mediated translational silencing of Ceruloplasmin expression / translational elongation / ribosomal large subunit export from nucleus / 90S preribosome / subtelomeric heterochromatin formation / ribosomal subunit export from nucleus / regulation of translational fidelity / protein-RNA complex assembly / maturation of LSU-rRNA / translation initiation factor activity / proteasomal protein catabolic process / rescue of stalled cytosolic ribosome / cytosolic ribosome / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / cytosolic ribosome assembly / protein catabolic process / assembly of large subunit precursor of preribosome / macroautophagy / translational initiation / RING-type E3 ubiquitin transferase / maintenance of translational fidelity / modification-dependent protein catabolic process / protein tag activity / rRNA processing / ubiquitin-protein transferase activity / ubiquitin protein ligase activity / chromatin organization / ribosome biogenesis / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / ubiquitin-dependent protein catabolic process / cytoplasmic translation / chromosome, telomeric region / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / protein ubiquitination / ribosome / translation / response to antibiotic / mRNA binding / ubiquitin protein ligase binding / nucleolus / RNA binding / zinc ion binding / metal ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
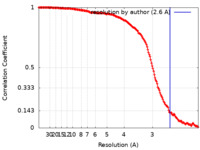
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Tesina P / Buschauer R / Beckmann R | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2023 Journal: Mol Cell / Year: 2023Title: Molecular basis of eIF5A-dependent CAT tailing in eukaryotic ribosome-associated quality control. Authors: Petr Tesina / Shuhei Ebine / Robert Buschauer / Matthias Thoms / Yoshitaka Matsuo / Toshifumi Inada / Roland Beckmann /   Abstract: Ribosome-associated quality control (RQC) is a conserved process degrading potentially toxic truncated nascent peptides whose malfunction underlies neurodegeneration and proteostasis decline in aging. ...Ribosome-associated quality control (RQC) is a conserved process degrading potentially toxic truncated nascent peptides whose malfunction underlies neurodegeneration and proteostasis decline in aging. During RQC, dissociation of stalled ribosomes is followed by elongation of the nascent peptide with alanine and threonine residues, driven by Rqc2 independently of mRNA, the small ribosomal subunit and guanosine triphosphate (GTP)-hydrolyzing factors. The resulting CAT tails (carboxy-terminal tails) and ubiquitination by Ltn1 mark nascent peptides for proteasomal degradation. Here we present ten cryogenic electron microscopy (cryo-EM) structures, revealing the mechanistic basis of individual steps of the CAT tailing cycle covering initiation, decoding, peptidyl transfer, and tRNA translocation. We discovered eIF5A as a crucial eukaryotic RQC factor enabling peptidyl transfer. Moreover, we observed dynamic behavior of RQC factors and tRNAs allowing for processivity of the CAT tailing cycle without additional energy input. Together, these results elucidate key differences as well as common principles between CAT tailing and canonical translation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15426.map.gz emd_15426.map.gz | 20.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15426-v30.xml emd-15426-v30.xml emd-15426.xml emd-15426.xml | 72.9 KB 72.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15426_fsc.xml emd_15426_fsc.xml | 15.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_15426.png emd_15426.png | 62.3 KB | ||
| Filedesc metadata |  emd-15426.cif.gz emd-15426.cif.gz | 15.4 KB | ||
| Others |  emd_15426_half_map_1.map.gz emd_15426_half_map_1.map.gz emd_15426_half_map_2.map.gz emd_15426_half_map_2.map.gz | 322.6 MB 322.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15426 http://ftp.pdbj.org/pub/emdb/structures/EMD-15426 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15426 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15426 | HTTPS FTP |
-Related structure data
| Related structure data |  8agwMC  8aafC  8agtC  8aguC  8agvC  8agxC  8agzC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15426.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15426.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Yeast RQC complex in state D, half map B | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.059 Å | ||||||||||||||||||||||||||||||||||||
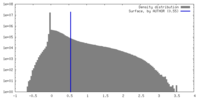
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Yeast RQC complex in state D, half map A
| File | emd_15426_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Yeast RQC complex in state D, half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
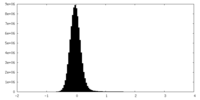
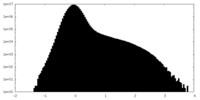
| Density Histograms |
-Half map: Yeast RQC complex in state D
| File | emd_15426_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Yeast RQC complex in state D | ||||||||||||
| Projections & Slices |
| ||||||||||||
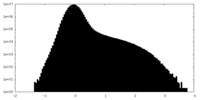
| Density Histograms |
- Sample components
Sample components
+Entire : Yeast RQC complex in state D
+Supramolecule #1: Yeast RQC complex in state D
+Macromolecule #1: 60S ribosomal protein L15-A
+Macromolecule #2: 60S ribosomal protein L16-A
+Macromolecule #3: 60S ribosomal protein L17-A
+Macromolecule #4: 60S ribosomal protein L18-A
+Macromolecule #5: 60S ribosomal protein L19-A
+Macromolecule #6: 60S ribosomal protein L20-A
+Macromolecule #7: 60S ribosomal protein L21-A
+Macromolecule #8: 60S ribosomal protein L22-A
+Macromolecule #9: 60S ribosomal protein L23-A
+Macromolecule #10: 60S ribosomal protein L24-A
+Macromolecule #11: 60S ribosomal protein L25
+Macromolecule #12: 60S ribosomal protein L26-A
+Macromolecule #13: 60S ribosomal protein L27-A
+Macromolecule #14: 60S ribosomal protein L28
+Macromolecule #15: 60S ribosomal protein L29
+Macromolecule #16: 60S ribosomal protein L30
+Macromolecule #17: 60S ribosomal protein L31-A
+Macromolecule #18: 60S ribosomal protein L32
+Macromolecule #19: 60S ribosomal protein L33-A
+Macromolecule #20: 60S ribosomal protein L34-A
+Macromolecule #21: 60S ribosomal protein L35-A
+Macromolecule #22: 60S ribosomal protein L36-A
+Macromolecule #23: 60S ribosomal protein L37-A
+Macromolecule #24: 60S ribosomal protein L38
+Macromolecule #25: 60S ribosomal protein L39
+Macromolecule #26: Ubiquitin-60S ribosomal protein L40
+Macromolecule #27: 60S ribosomal protein L42-A
+Macromolecule #28: 60S ribosomal protein L43-A
+Macromolecule #29: 60S ribosomal protein L41-A
+Macromolecule #33: 60S ribosomal protein L2-A
+Macromolecule #34: 60S ribosomal protein L3
+Macromolecule #35: 60S ribosomal protein L4-A
+Macromolecule #36: 60S ribosomal protein L5
+Macromolecule #37: 60S ribosomal protein L6-B
+Macromolecule #38: 60S ribosomal protein L7-A
+Macromolecule #39: 60S ribosomal protein L8-A
+Macromolecule #40: 60S ribosomal protein L9-A
+Macromolecule #41: 60S ribosomal protein L10
+Macromolecule #42: 60S ribosomal protein L11-A
+Macromolecule #43: 60S ribosomal protein L13-A
+Macromolecule #44: 60S ribosomal protein L14-A
+Macromolecule #45: RQC2 isoform 1
+Macromolecule #46: E3 ubiquitin-protein ligase listerin
+Macromolecule #47: Eukaryotic translation initiation factor 6
+Macromolecule #49: 60S ribosomal protein L12-A
+Macromolecule #50: 60S acidic ribosomal protein P0
+Macromolecule #51: CAT-tailed nascent peptide
+Macromolecule #52: 60S ribosomal protein L1-A
+Macromolecule #30: 25S rRNA
+Macromolecule #31: 5S rRNA
+Macromolecule #32: 5.8S rRNA
+Macromolecule #48: Ala tRNA
+Macromolecule #53: MAGNESIUM ION
+Macromolecule #54: ZINC ION
+Macromolecule #55: SPERMIDINE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 46.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.4 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)